![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-158_CAGAGAGG-ACTGCATA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1269630 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
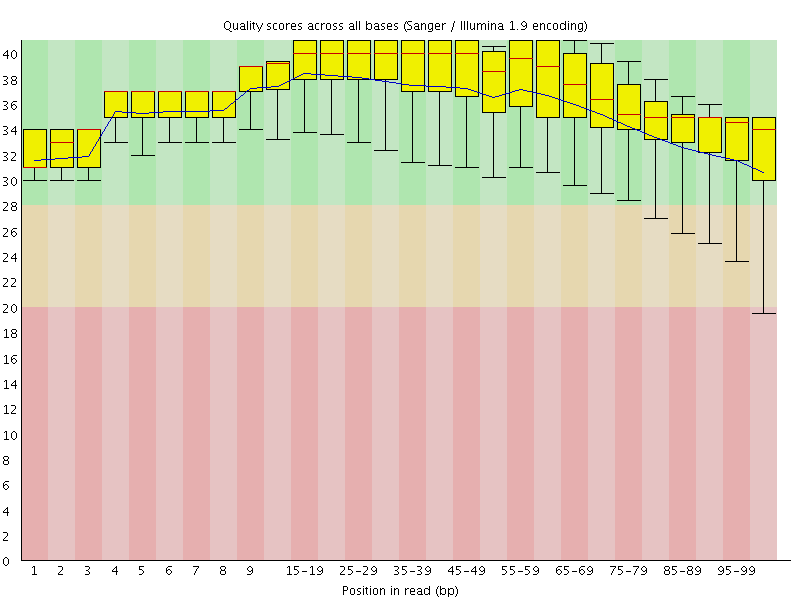
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
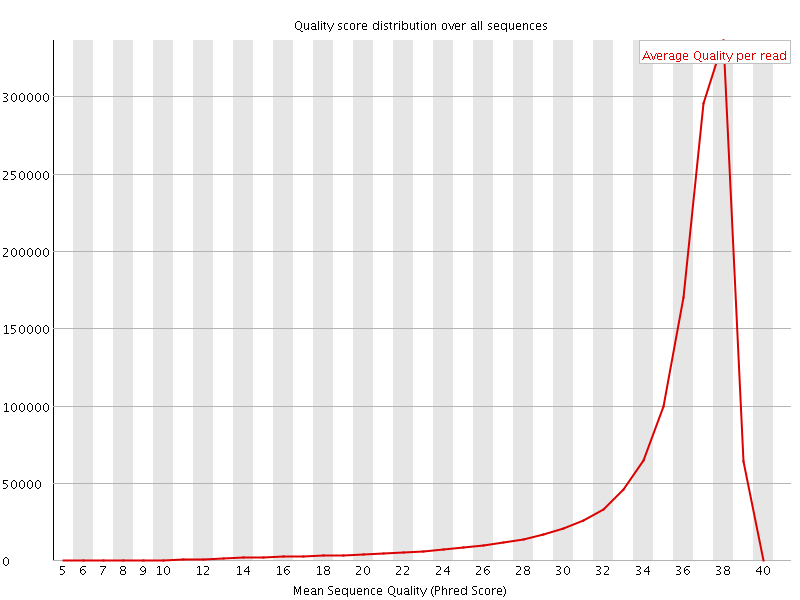
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
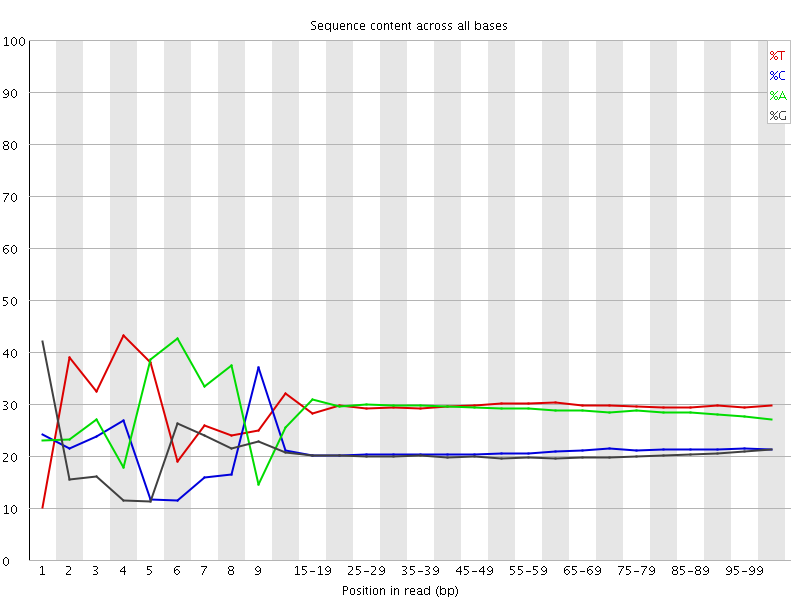
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
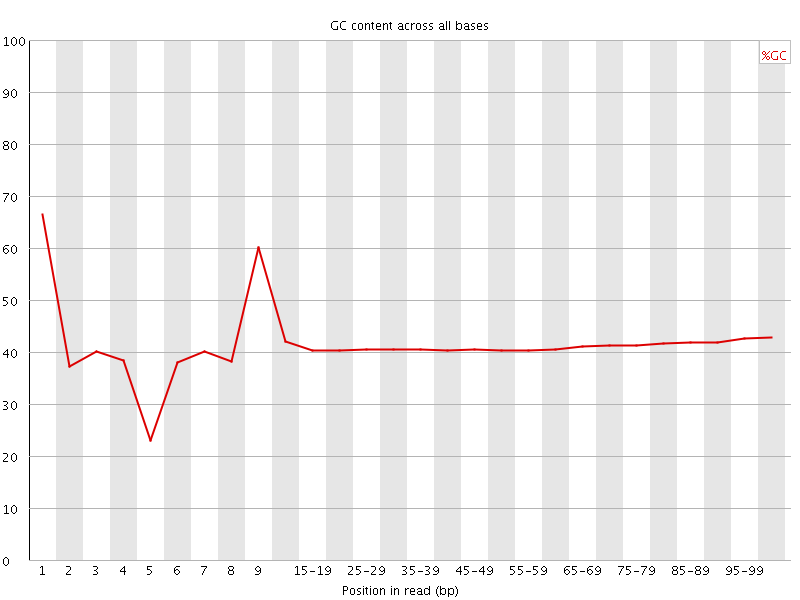
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
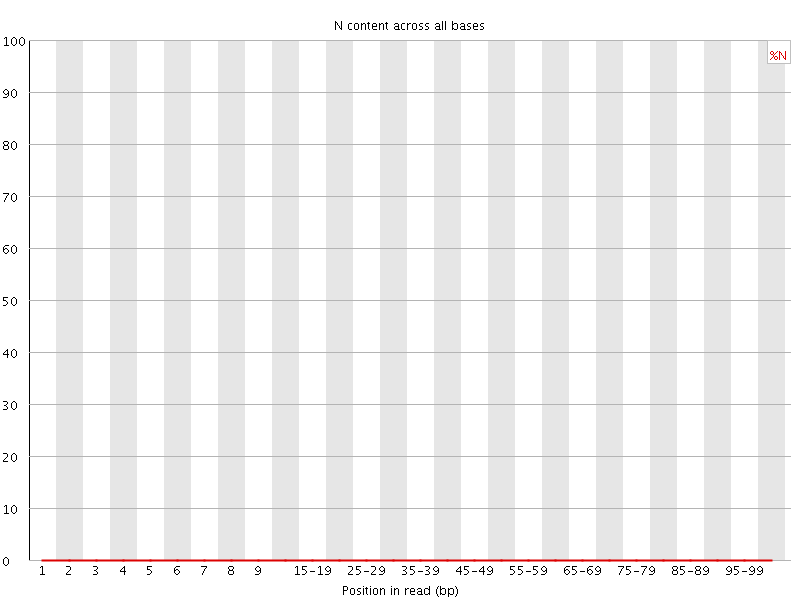
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
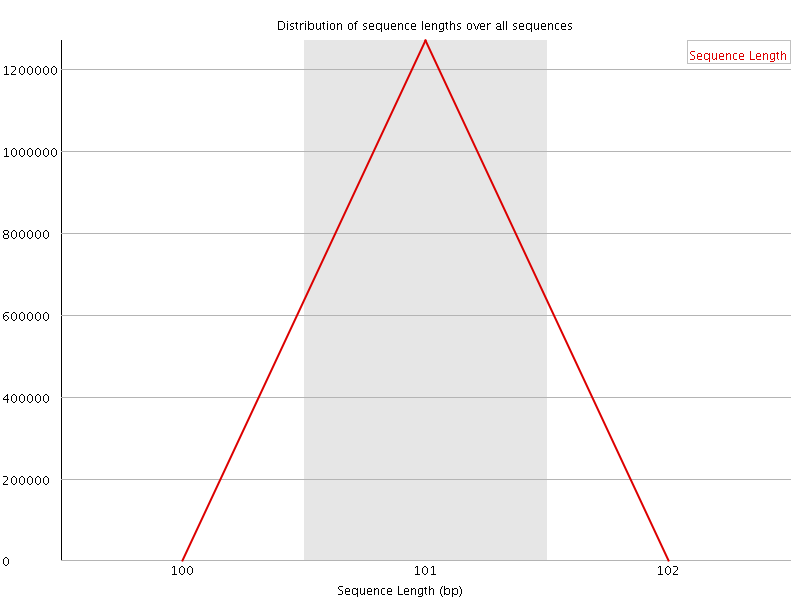
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
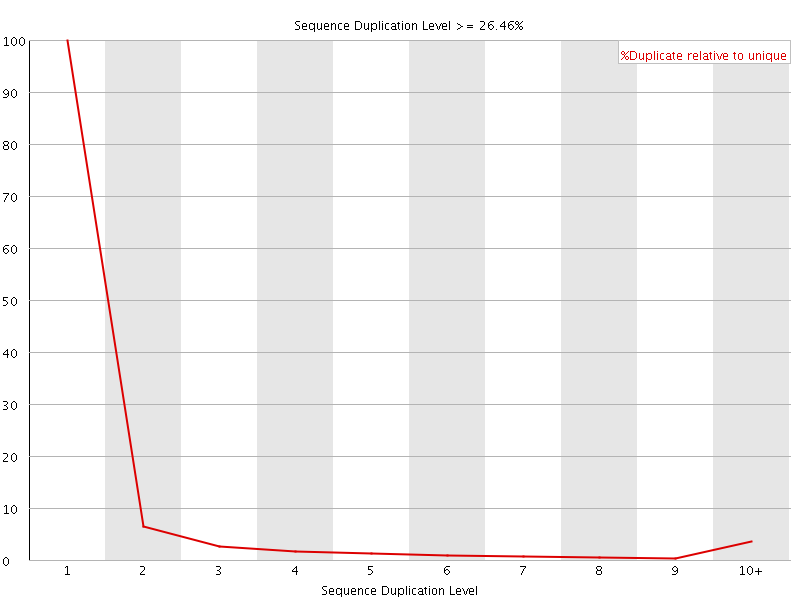
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
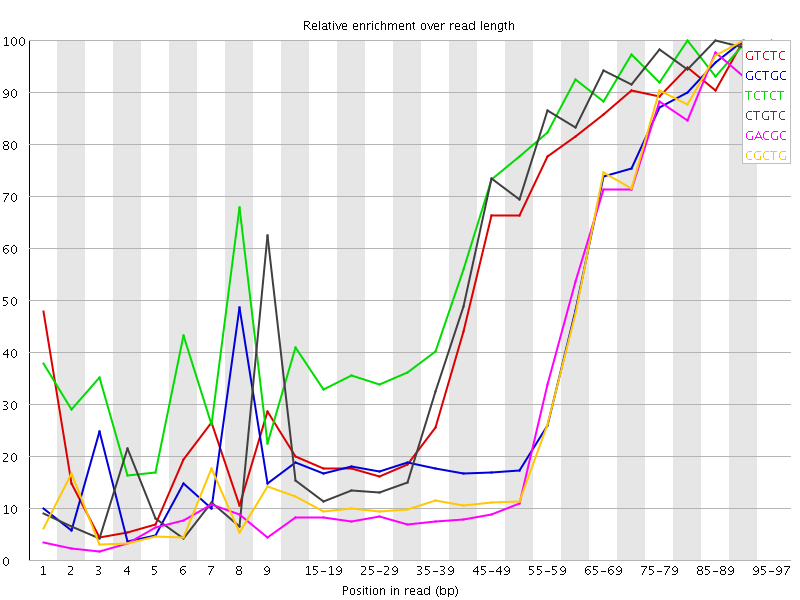
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 327165 | 3.366853 | 5.9812093 | 90-94 |
| GCTGC | 217430 | 3.268638 | 7.538723 | 90-94 |
| TCTCT | 456800 | 3.218054 | 4.8402014 | 80-84 |
| CTGTC | 306555 | 3.1547554 | 5.434252 | 85-89 |
| GACGC | 185870 | 2.866546 | 7.5125184 | 95-97 |
| CGCTG | 189205 | 2.8443298 | 7.2091775 | 90-94 |
| CTGCC | 190630 | 2.7835333 | 7.258702 | 90-94 |
| ACACA | 359165 | 2.7319198 | 6.403599 | 6 |
| GCCGA | 176825 | 2.7270508 | 7.3793917 | 90-94 |
| CGACG | 171910 | 2.6512504 | 7.612253 | 95-97 |
| TGCCG | 169310 | 2.545247 | 7.206261 | 90-94 |
| AGAAG | 310610 | 2.5042267 | 5.021754 | 5 |
| CCGAC | 166380 | 2.4923472 | 7.2388597 | 95-97 |
| TACAC | 322785 | 2.393233 | 7.955894 | 5 |
| CTGAC | 218665 | 2.3085475 | 5.56605 | 95-97 |
| TGACG | 206820 | 2.2479892 | 5.737486 | 95-97 |
| GACGA | 191015 | 2.1299598 | 5.8229523 | 95-97 |
| TGCAG | 176630 | 1.919845 | 5.0838037 | 90-94 |
| CTTTG | 262465 | 1.9036223 | 7.4568305 | 9 |
| TATAC | 334660 | 1.7487495 | 5.4981365 | 5 |
| GAGAG | 150940 | 1.7328079 | 5.5953374 | 7 |
| GCTTT | 222260 | 1.6120212 | 5.0278316 | 1 |
| GTGTA | 181815 | 1.392783 | 9.409283 | 1 |
| CCATG | 125725 | 1.3273369 | 5.6473417 | 9 |
| AAGAC | 167565 | 1.3121979 | 5.360129 | 5 |
| GAGTC | 117775 | 1.2801322 | 6.478323 | 9 |
| GTCTT | 170170 | 1.2342196 | 6.2557626 | 1 |
| TATGA | 227710 | 1.2250338 | 5.517002 | 4 |
| TCATA | 225955 | 1.1807169 | 5.4955583 | 2 |
| GACTT | 157890 | 1.1748064 | 5.723069 | 7 |
| GATTG | 152900 | 1.1712815 | 5.45002 | 7 |
| GTGTT | 153480 | 1.1460494 | 5.6762247 | 1 |
| GTTCT | 156645 | 1.1361245 | 5.2635655 | 1 |
| TGGAC | 94540 | 1.027584 | 6.3008885 | 5 |
| GTCCA | 97190 | 1.0260798 | 7.451806 | 1 |
| AGACT | 123650 | 0.9438608 | 5.1936417 | 6 |
| GGACT | 85645 | 0.9309014 | 5.808764 | 6 |
| GAGTA | 116995 | 0.9194398 | 5.169574 | 1 |
| GTATA | 136785 | 0.73587567 | 6.8011036 | 1 |