![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-158_CAGAGAGG-ACTGCATA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1269630 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
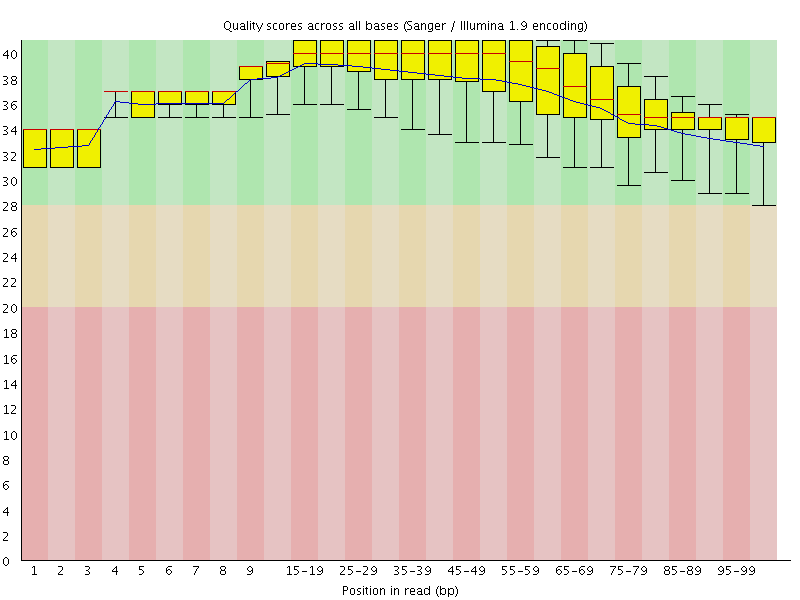
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
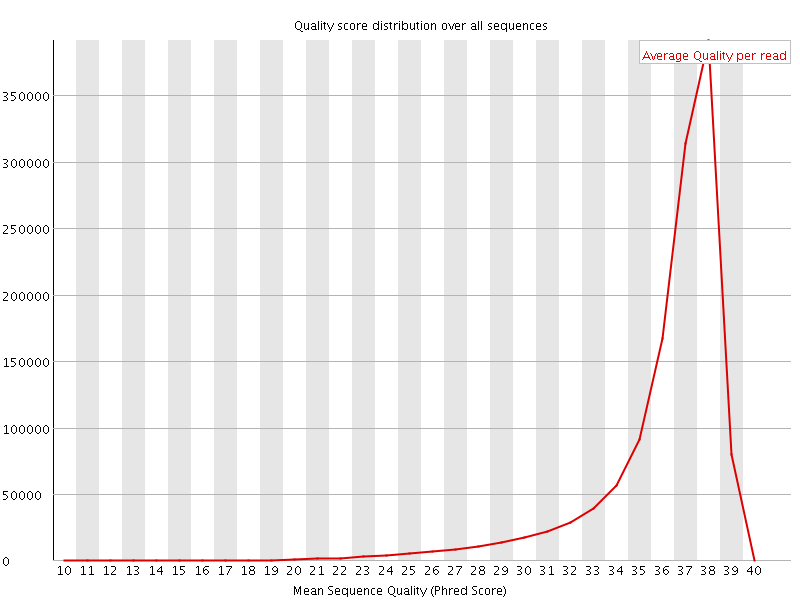
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
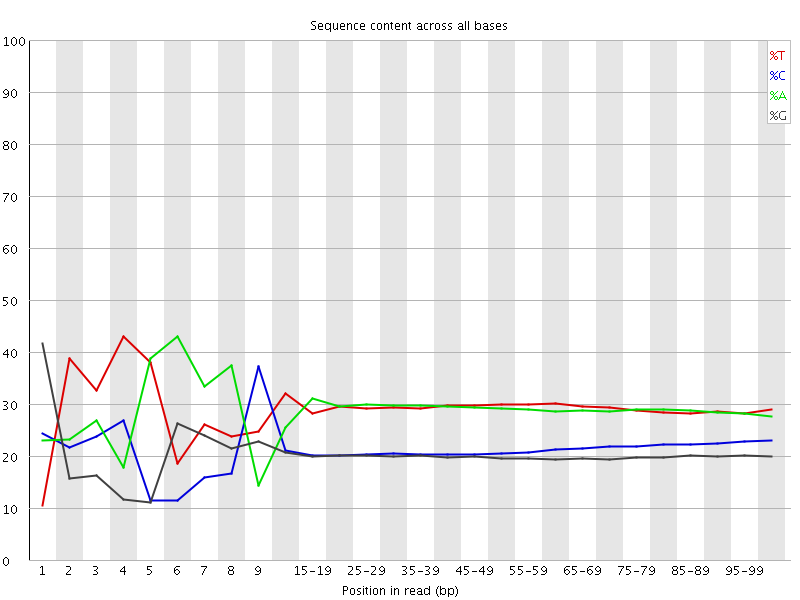
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
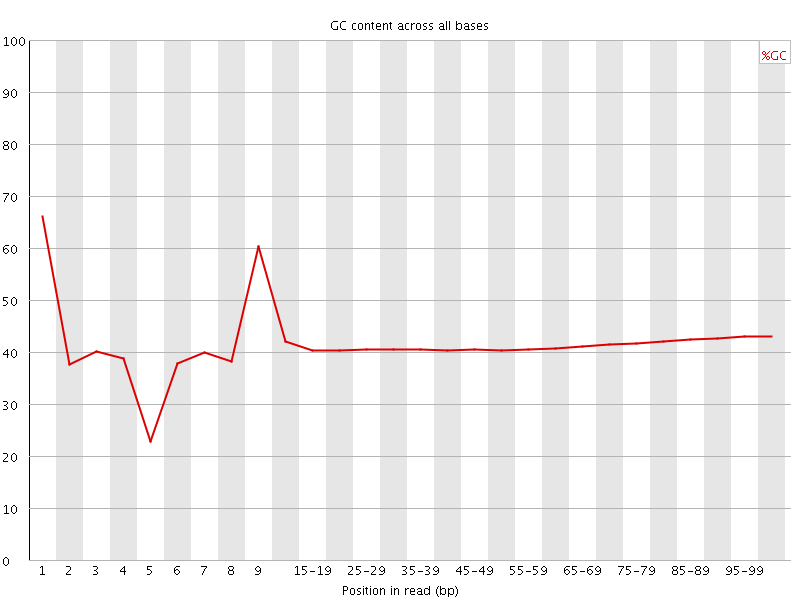
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
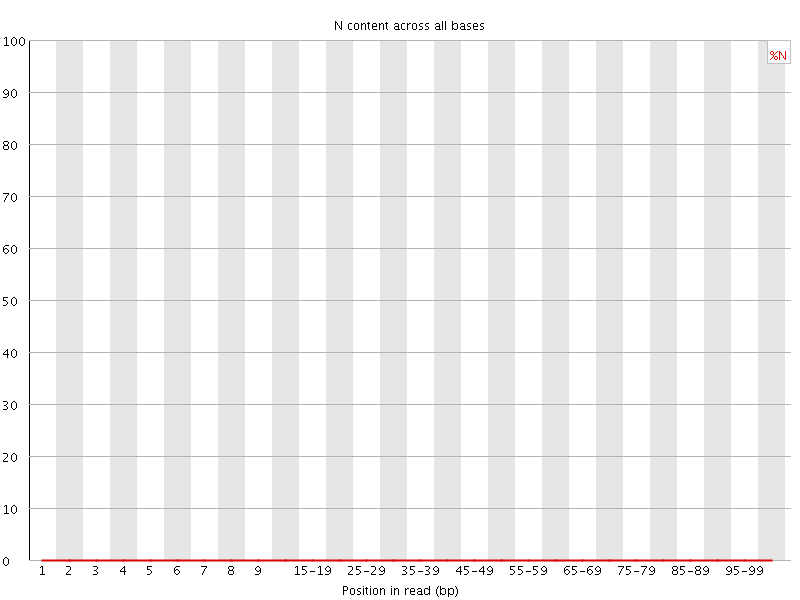
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
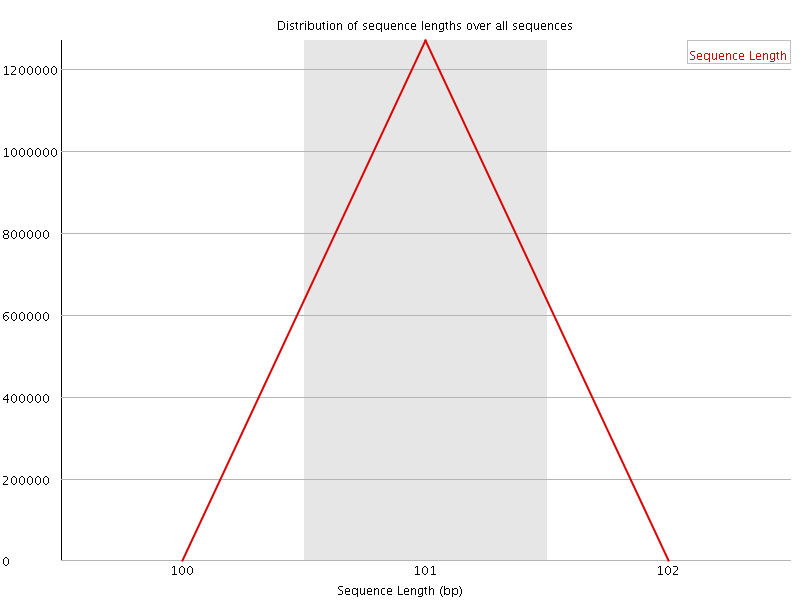
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
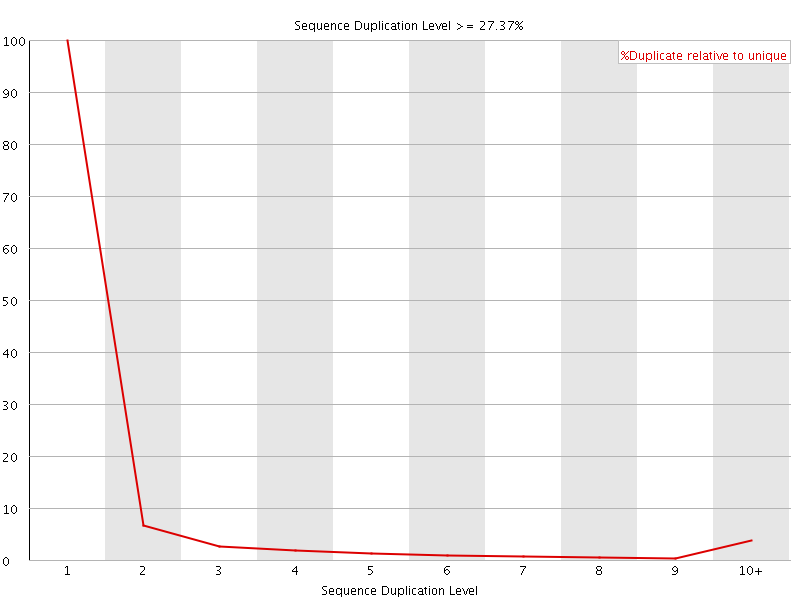
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
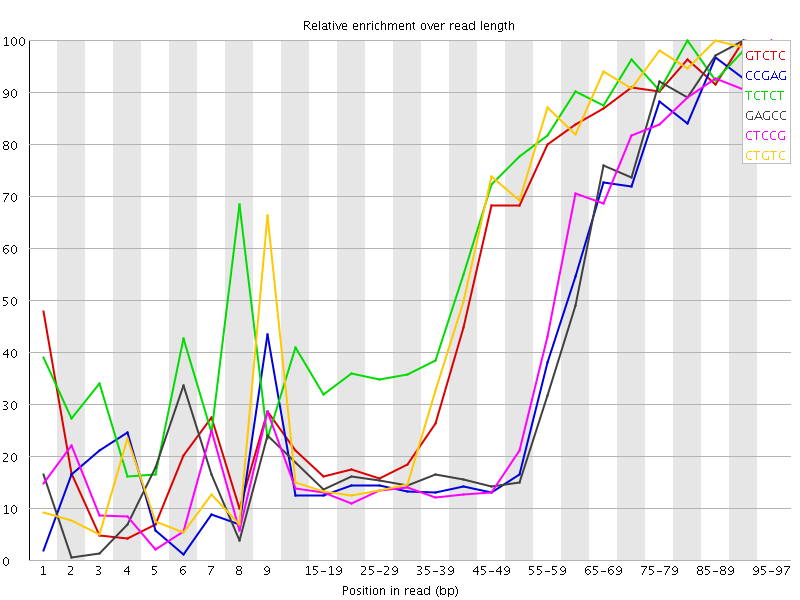
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 326205 | 3.343627 | 5.8529315 | 90-94 |
| CCGAG | 216415 | 3.2731214 | 7.832232 | 95-97 |
| TCTCT | 457045 | 3.2081847 | 4.8745365 | 80-84 |
| GAGCC | 211465 | 3.1982563 | 7.4363723 | 90-94 |
| CTCCG | 223415 | 3.1605952 | 7.378597 | 95-97 |
| CTGTC | 307840 | 3.155384 | 5.4234996 | 85-89 |
| CGAGC | 202185 | 3.0579028 | 7.3848853 | 95-97 |
| TCTCC | 297365 | 2.8808432 | 5.746641 | 95-97 |
| AGCCC | 198355 | 2.83544 | 6.9065223 | 90-94 |
| GCCCA | 198110 | 2.8319378 | 7.129716 | 90-94 |
| ATCTC | 377520 | 2.677695 | 6.3946676 | 95-97 |
| GAGAG | 224065 | 2.62505 | 5.934455 | 95-97 |
| ACACA | 361255 | 2.6162333 | 6.102784 | 6 |
| AGAAG | 313150 | 2.5386918 | 5.2121468 | 5 |
| CCCAC | 187385 | 2.5317142 | 6.6259966 | 90-94 |
| TCCGA | 236005 | 2.4443824 | 5.6532383 | 95-97 |
| CCACG | 170135 | 2.4320416 | 6.819186 | 90-94 |
| GAGGA | 206940 | 2.4244213 | 6.1029835 | 95-97 |
| TACAC | 323625 | 2.3194447 | 7.6867485 | 5 |
| GAGAC | 202520 | 2.2425082 | 5.8058424 | 95-97 |
| CGAGA | 200020 | 2.2148256 | 5.734254 | 95-97 |
| AGAGG | 183865 | 2.154084 | 5.690184 | 95-97 |
| CAGAG | 193680 | 2.1446228 | 5.4659085 | 90-94 |
| ACGAG | 187375 | 2.0748074 | 5.5696 | 95-97 |
| CAAAG | 268890 | 2.0603204 | 5.0117297 | 4 |
| GACCA | 194085 | 2.0312374 | 5.0679083 | 95-97 |
| CTTTG | 264265 | 1.9626254 | 7.6771593 | 9 |
| GGATC | 172615 | 1.8915765 | 5.302924 | 95-97 |
| CACGA | 178130 | 1.8642571 | 5.1626363 | 95-97 |
| AGACC | 174290 | 1.8240685 | 5.265821 | 95-97 |
| TATAC | 336300 | 1.7463827 | 5.5694623 | 5 |
| GCTTT | 224120 | 1.6644795 | 5.0636106 | 1 |
| GTCTT | 180600 | 1.3412681 | 5.8055053 | 1 |
| CCATG | 126765 | 1.3129473 | 5.5943384 | 9 |
| AAGAC | 169050 | 1.2953147 | 5.516989 | 5 |
| GAGTC | 118050 | 1.2936338 | 6.2165127 | 9 |
| TATGA | 226300 | 1.2433532 | 5.6342473 | 4 |
| GATTG | 152775 | 1.213021 | 5.643722 | 7 |
| GTGTT | 153035 | 1.2025027 | 5.9633017 | 1 |
| TCATA | 227005 | 1.1788212 | 5.3939776 | 2 |
| GACTT | 156155 | 1.1718566 | 5.6980453 | 7 |
| GTTCT | 156425 | 1.1617267 | 5.6146293 | 1 |
| GTGTA | 134405 | 1.0671648 | 9.383152 | 1 |
| TGGAC | 95865 | 1.0505227 | 6.8541036 | 5 |
| GCCGT | 67380 | 1.0085211 | 5.12629 | 95-97 |
| GTCCA | 96730 | 1.0018649 | 6.4338994 | 1 |
| TGCCG | 65870 | 0.98591995 | 5.3996606 | 95-97 |
| AGACT | 124480 | 0.94392806 | 5.1583714 | 6 |
| GGACT | 85570 | 0.9377065 | 6.1633797 | 6 |
| GAGTA | 116665 | 0.93600327 | 5.174467 | 1 |
| GTATA | 136810 | 0.75167114 | 6.9458947 | 1 |