![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-157_CAGAGAGG-GTAAGGAG_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1066943 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
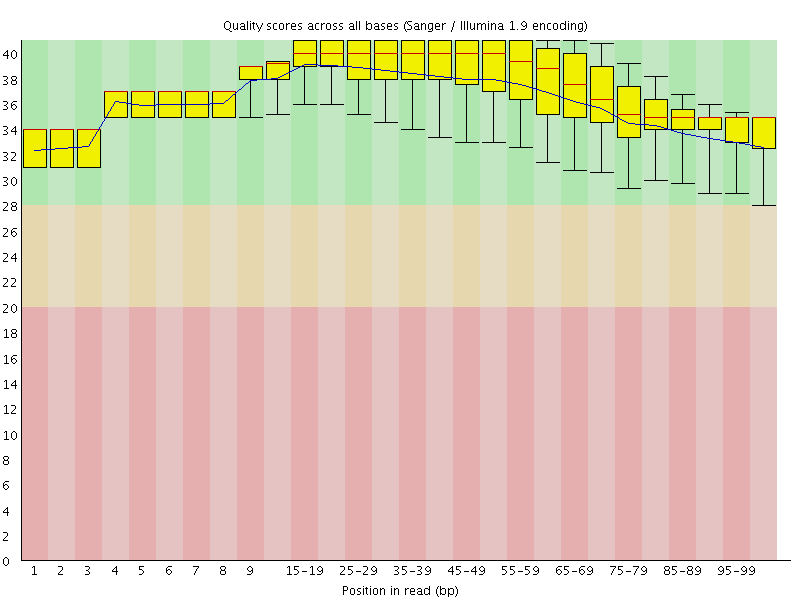
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
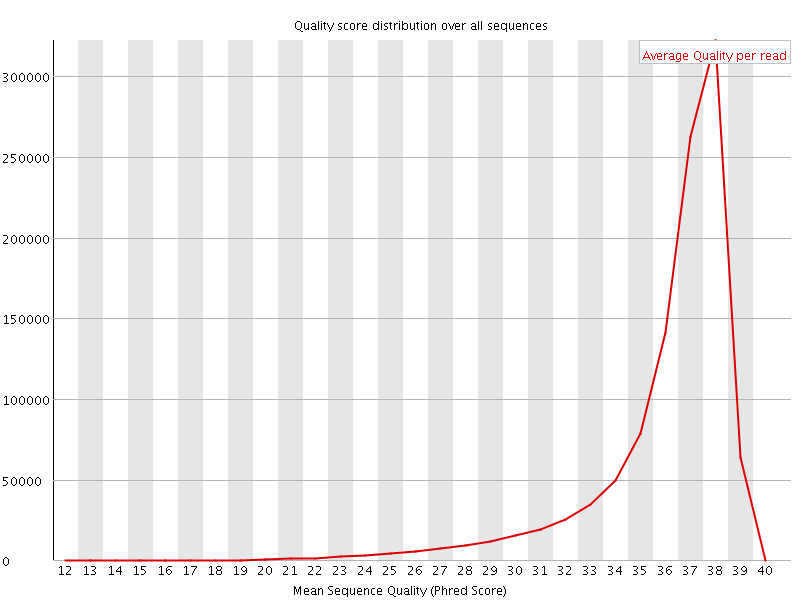
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
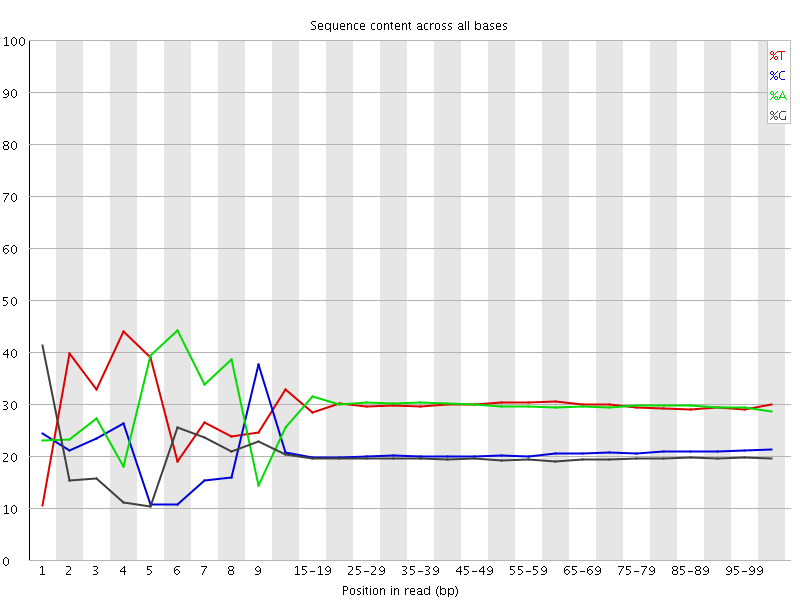
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
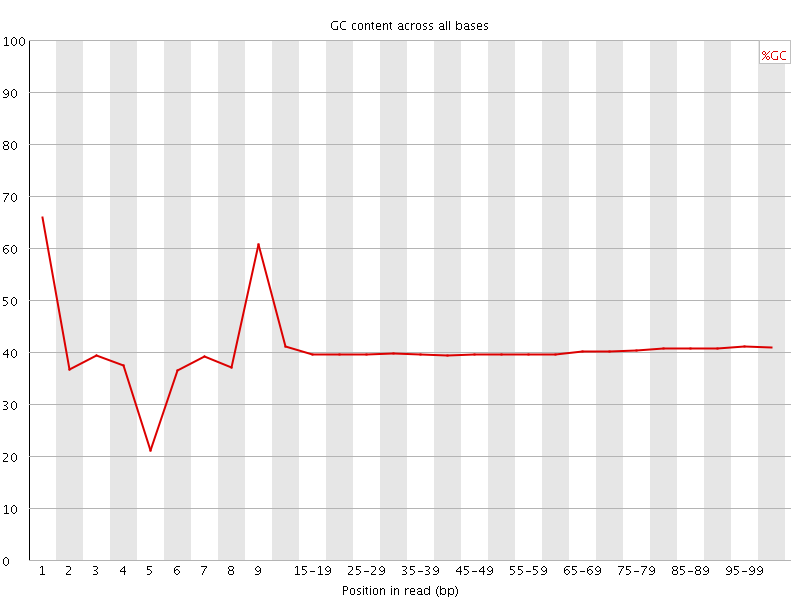
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
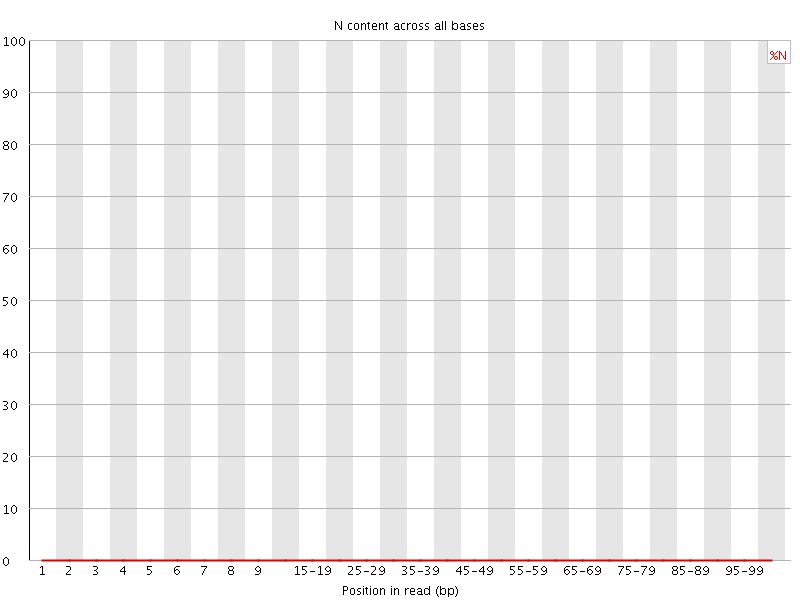
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
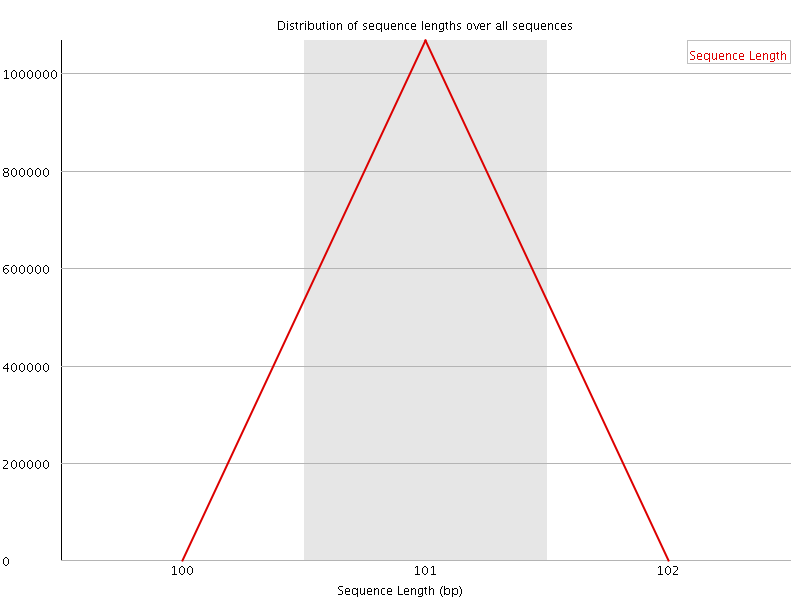
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
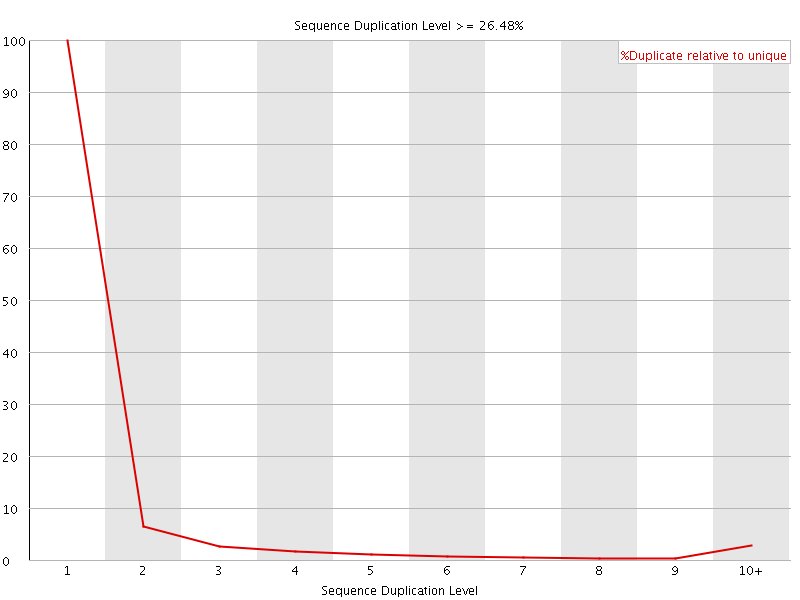
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
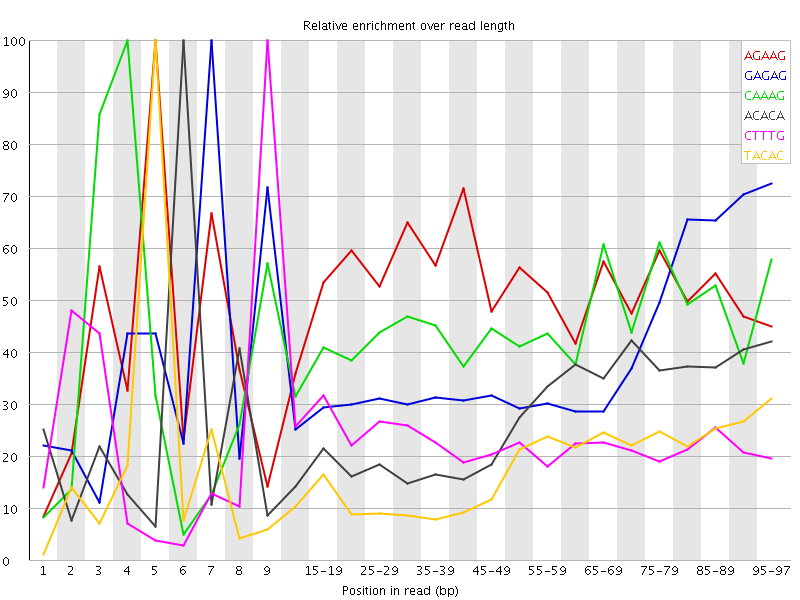
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 289390 | 2.710789 | 5.2093215 | 5 |
| GAGAG | 165345 | 2.3251562 | 5.9317923 | 7 |
| CAAAG | 250385 | 2.2609246 | 5.0873256 | 4 |
| ACACA | 248935 | 2.1668525 | 7.8203177 | 6 |
| CTTTG | 242535 | 2.153357 | 9.285331 | 9 |
| TACAC | 216930 | 1.8776629 | 10.348959 | 5 |
| GCTTT | 205490 | 1.8244513 | 5.342811 | 1 |
| CACAT | 209575 | 1.8140007 | 5.5437856 | 7 |
| TGGCT | 122740 | 1.6452125 | 5.277127 | 6 |
| TTGAG | 167940 | 1.5555198 | 5.4204364 | 9 |
| ATACA | 259245 | 1.5505791 | 5.170594 | 6 |
| GACTC | 113950 | 1.4806801 | 5.884362 | 7 |
| GAGTC | 107240 | 1.4455665 | 7.2872176 | 9 |
| CCATG | 108780 | 1.4135004 | 5.644955 | 9 |
| AAGAC | 154815 | 1.3979473 | 6.247515 | 5 |
| GTCTT | 157425 | 1.3977042 | 6.264134 | 1 |
| CATGA | 153470 | 1.3780211 | 5.010872 | 9 |
| GACTT | 148095 | 1.3222921 | 7.0260477 | 7 |
| TATGA | 212735 | 1.3125366 | 6.311906 | 4 |
| TGAGT | 139460 | 1.291728 | 5.0701513 | 8 |
| TATAC | 216490 | 1.287585 | 6.5170665 | 5 |
| TCATA | 214375 | 1.2750059 | 6.35921 | 2 |
| GTGTT | 137665 | 1.2679425 | 6.283861 | 1 |
| ACTCA | 145915 | 1.2629843 | 5.3003793 | 8 |
| GATTG | 135085 | 1.2512052 | 6.3006396 | 7 |
| AACAC | 137935 | 1.200654 | 5.148833 | 5 |
| GTGTA | 129310 | 1.197715 | 10.828669 | 1 |
| GTATG | 124460 | 1.1527927 | 5.6629415 | 3 |
| TGGAC | 84635 | 1.1408572 | 7.947315 | 5 |
| ACACC | 89810 | 1.1313126 | 5.5761476 | 6 |
| ACATG | 125305 | 1.125125 | 5.6900167 | 8 |
| GTCCA | 85635 | 1.1127516 | 7.8320336 | 1 |
| GGACT | 78490 | 1.0580243 | 7.810067 | 6 |
| ACACT | 121500 | 1.0516573 | 5.7746015 | 6 |
| AGACT | 112520 | 1.0103273 | 5.833682 | 6 |
| GAGTA | 106080 | 0.9880986 | 5.343281 | 1 |
| CATAC | 105830 | 0.91602385 | 5.472442 | 3 |
| GTATA | 125205 | 0.7724923 | 7.356775 | 1 |
| ATACT | 129675 | 0.7712485 | 5.3289995 | 6 |