![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-155_CAGAGAGG-TATCCTCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1297177 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
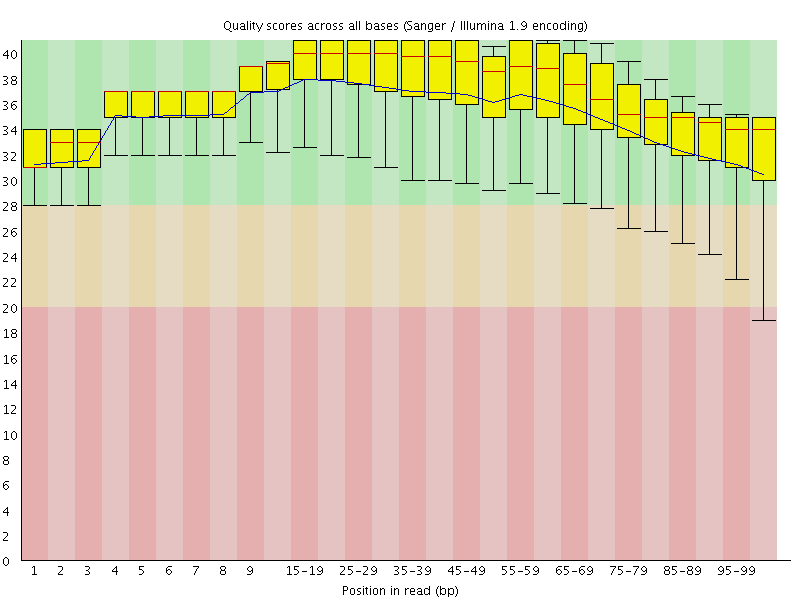
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
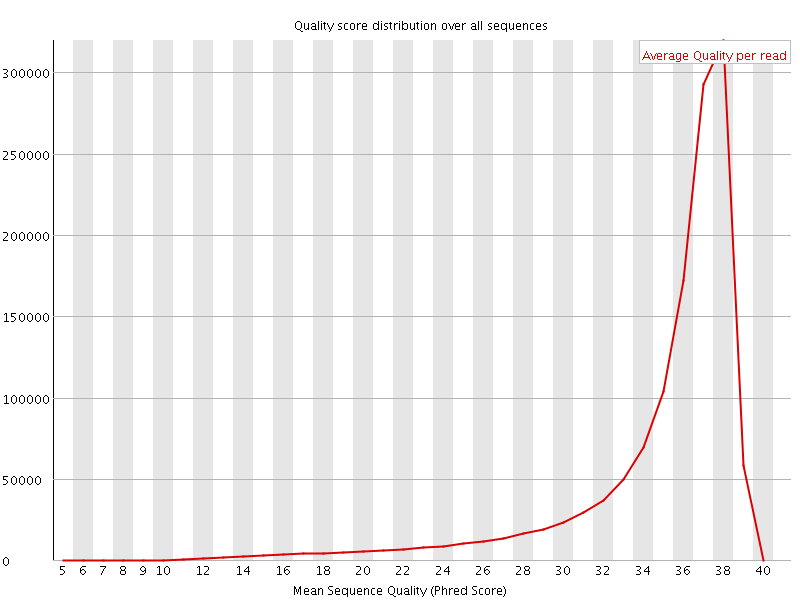
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
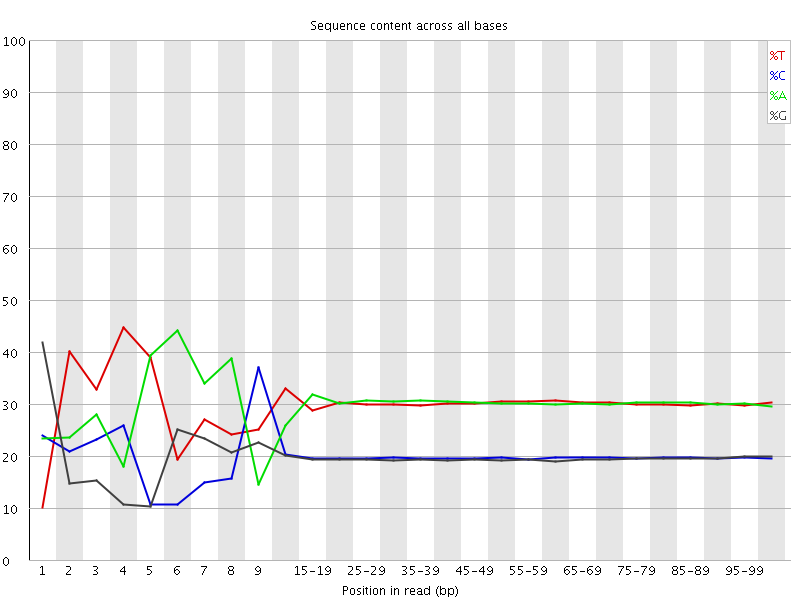
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
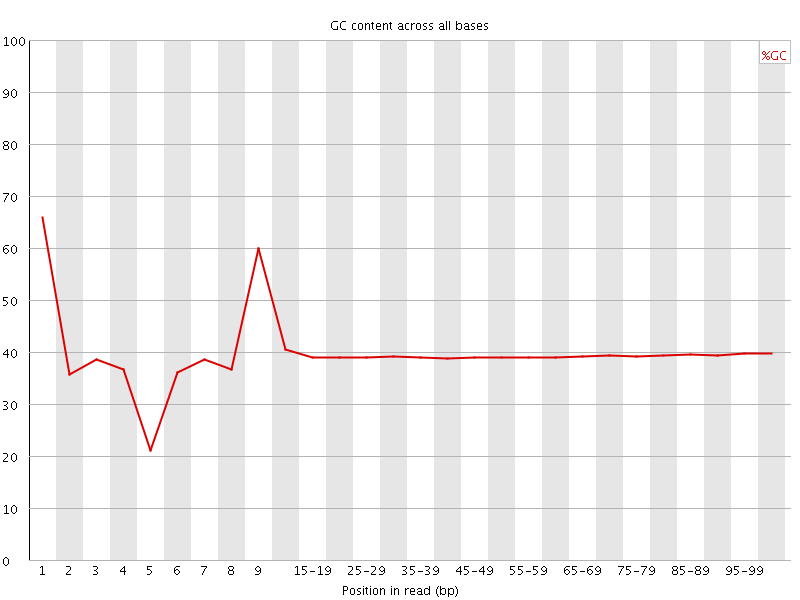
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
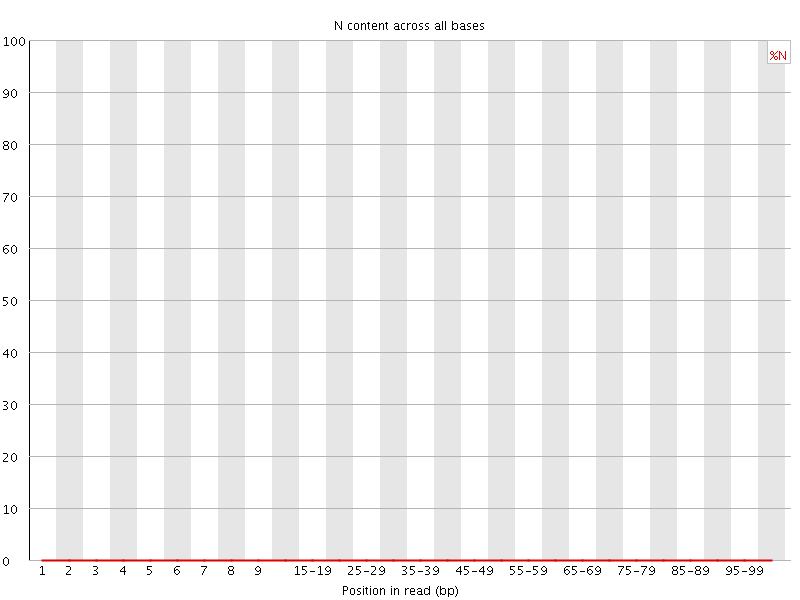
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
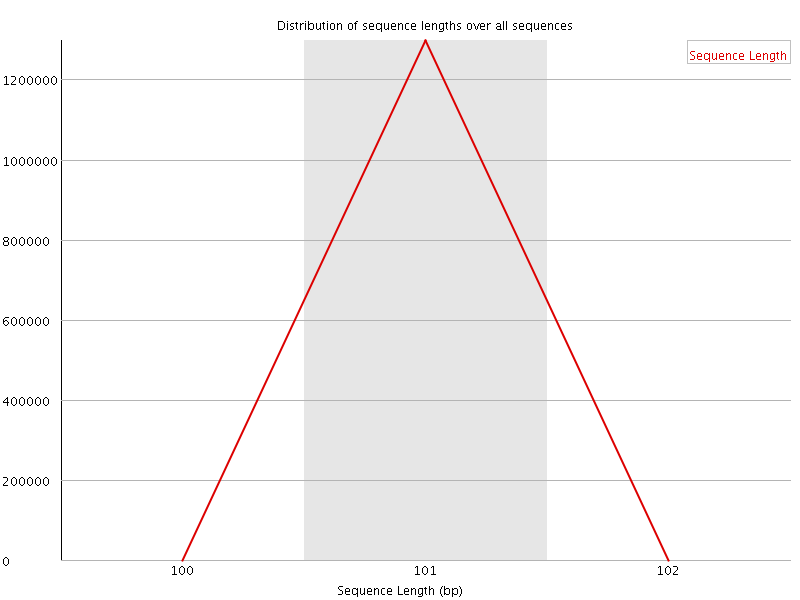
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
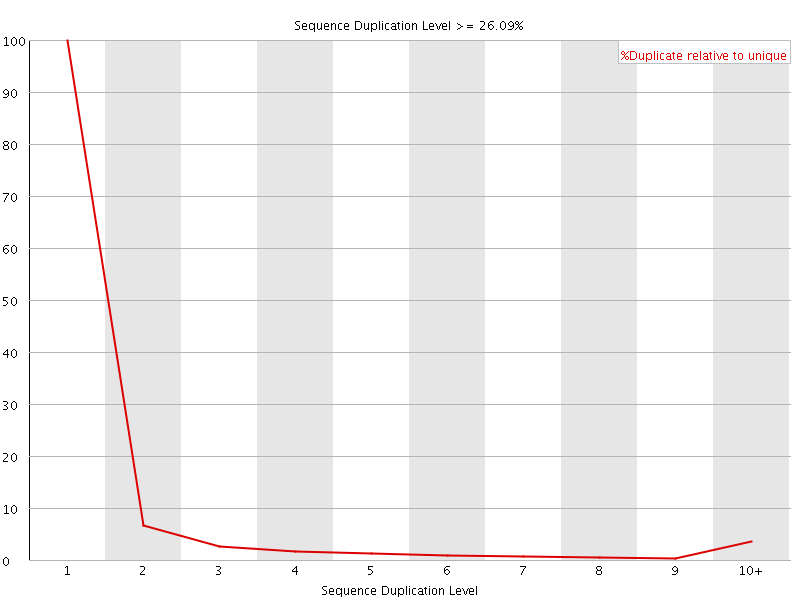
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
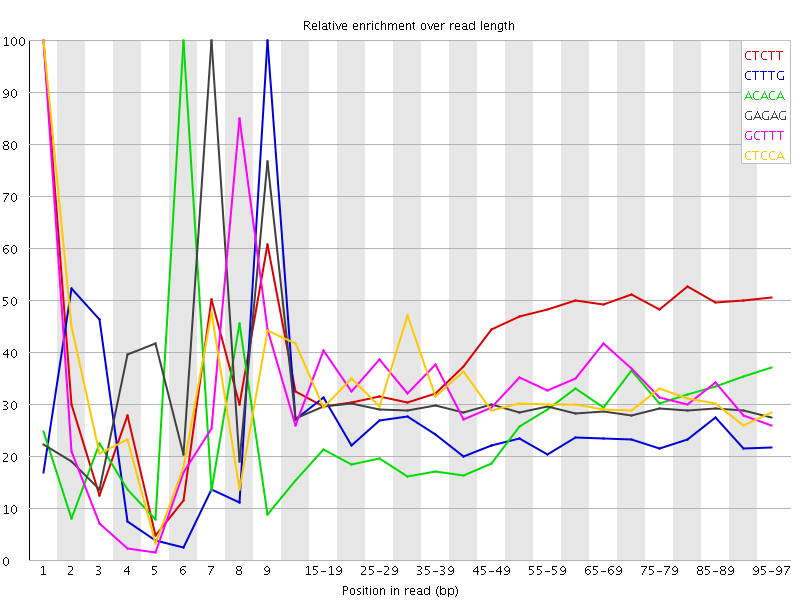
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CTCTT | 291600 | 2.1109967 | 5.0428004 | 1 |
| CTTTG | 279725 | 2.0450711 | 8.356706 | 9 |
| ACACA | 248350 | 1.8228959 | 7.0620813 | 6 |
| GAGAG | 153950 | 1.764094 | 5.8903975 | 7 |
| GCTTT | 235935 | 1.7249222 | 5.170731 | 1 |
| CTCCA | 151940 | 1.6826108 | 5.1785226 | 1 |
| TACAC | 204215 | 1.4920595 | 8.785785 | 5 |
| GACTC | 125745 | 1.4063046 | 5.1198516 | 7 |
| CCATG | 123045 | 1.3761083 | 5.212072 | 9 |
| GAGTC | 120635 | 1.3625079 | 6.4467463 | 9 |
| AAGAC | 176945 | 1.3116353 | 5.5696993 | 5 |
| GTCTT | 176115 | 1.2875777 | 6.5077443 | 1 |
| GTGTT | 165700 | 1.2234232 | 6.0850596 | 1 |
| GACTT | 165530 | 1.2157747 | 6.0373 | 7 |
| GATTG | 163680 | 1.214085 | 5.528713 | 7 |
| GTTCT | 165585 | 1.2105931 | 5.6211314 | 1 |
| TATGA | 247310 | 1.1984087 | 5.5167327 | 4 |
| TCATA | 248810 | 1.1938617 | 5.7211103 | 2 |
| GTGTA | 154295 | 1.1444725 | 9.930699 | 1 |
| TGGAC | 98090 | 1.1078742 | 7.3084807 | 5 |
| GTCCA | 97470 | 1.090083 | 7.7687097 | 1 |
| TATAC | 219065 | 1.0511367 | 6.158594 | 5 |
| ACACT | 140545 | 1.0268663 | 5.201372 | 6 |
| GGACT | 90350 | 1.0204551 | 6.857463 | 6 |
| AGACT | 129890 | 0.95840997 | 5.1705503 | 6 |
| GTATG | 127285 | 0.9441276 | 5.037136 | 3 |
| ATACT | 158845 | 0.7621839 | 5.242528 | 6 |
| GTATA | 154450 | 0.7484301 | 7.517251 | 1 |