![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-154_CAGAGAGG-CTCTCTAT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1786736 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
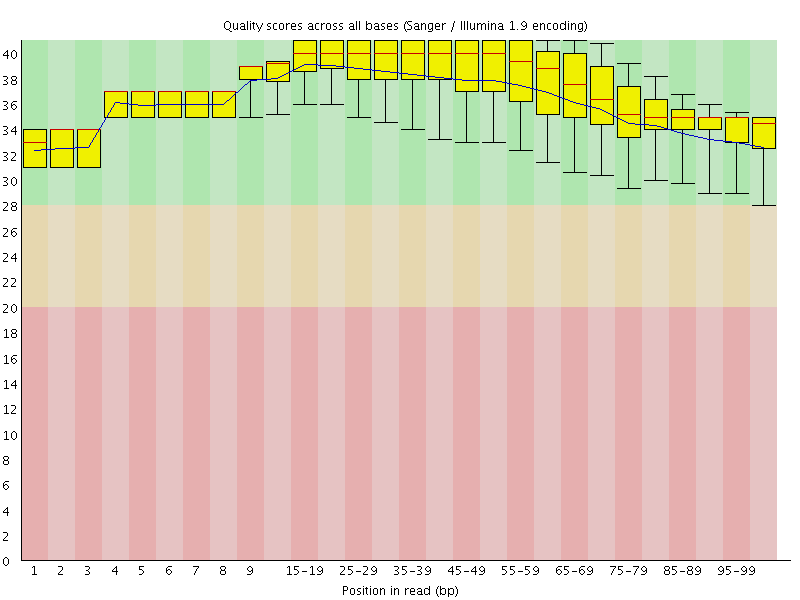
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
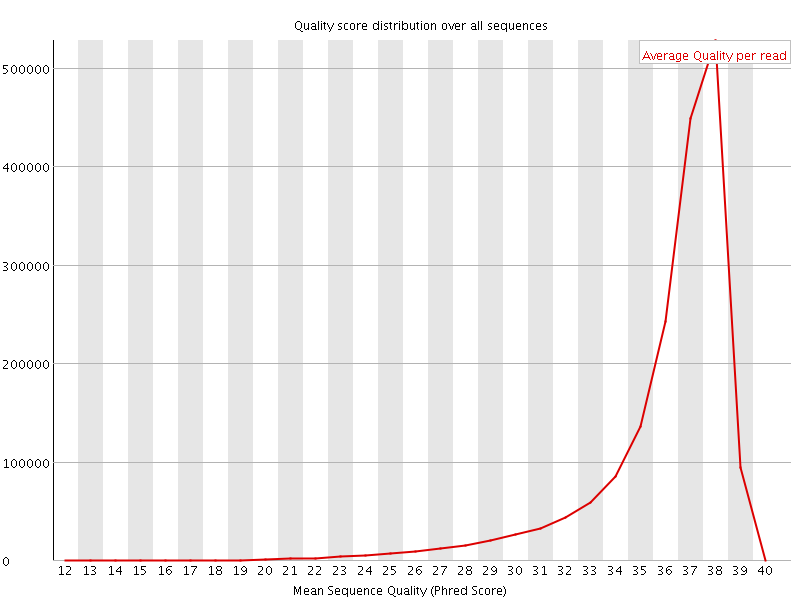
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
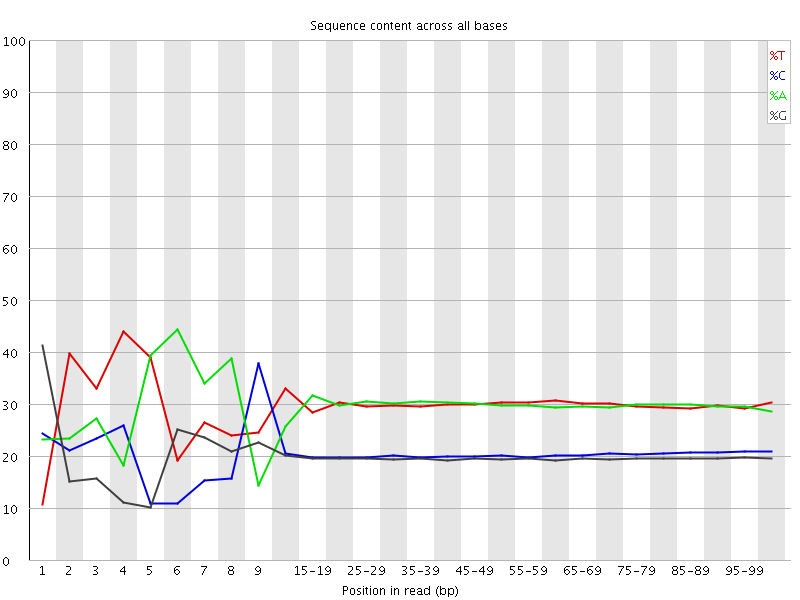
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
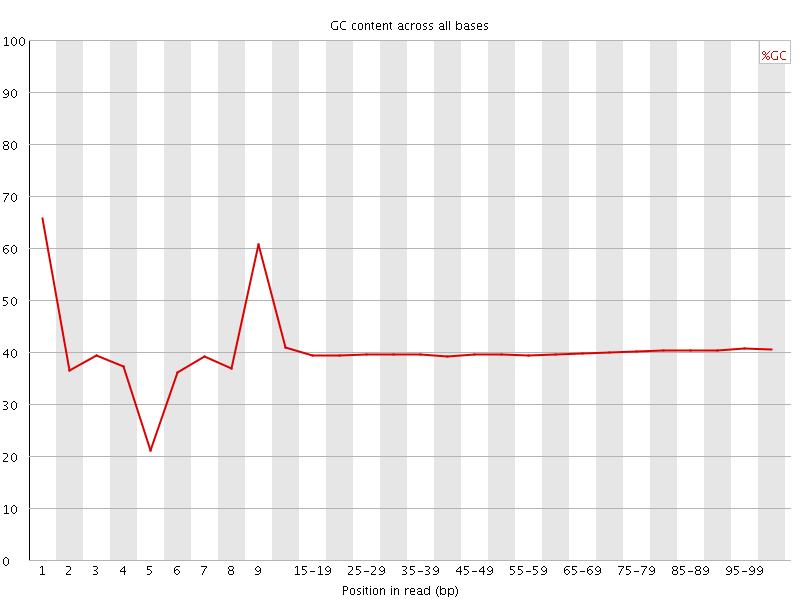
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
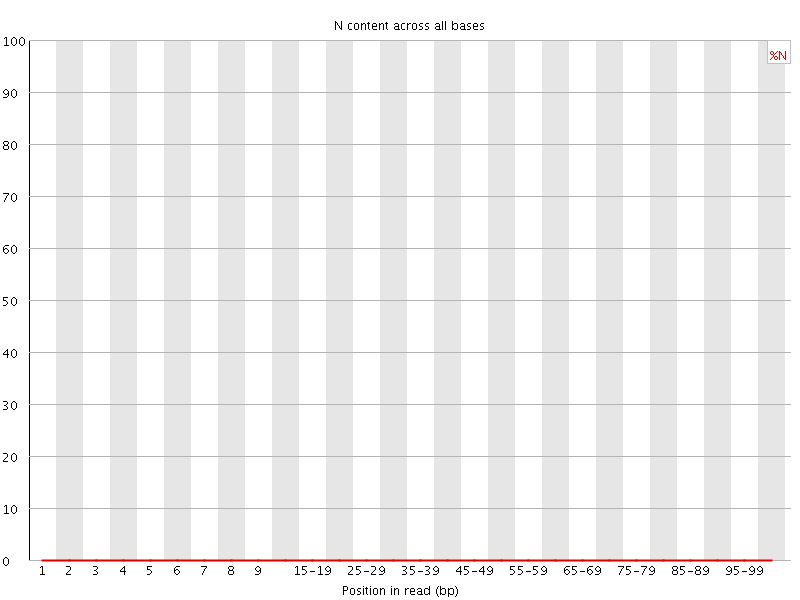
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
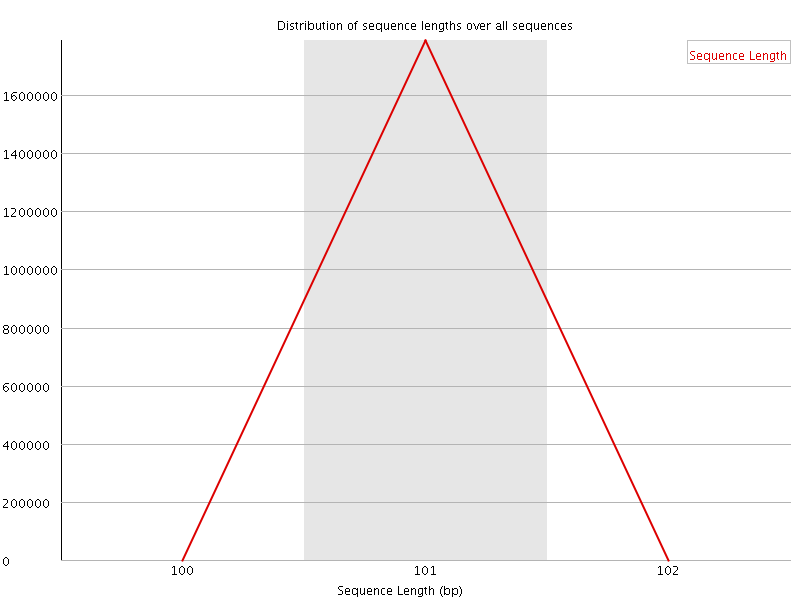
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
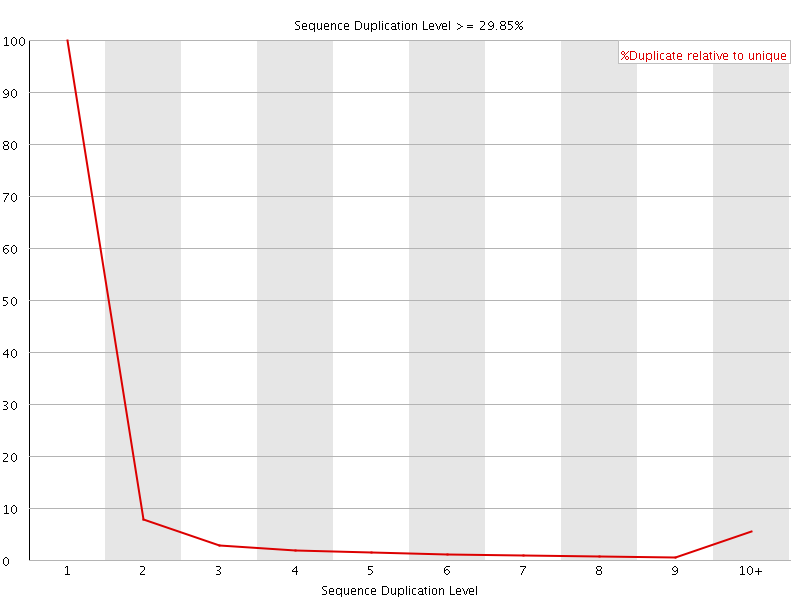
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 1953 | 0.10930545978812761 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
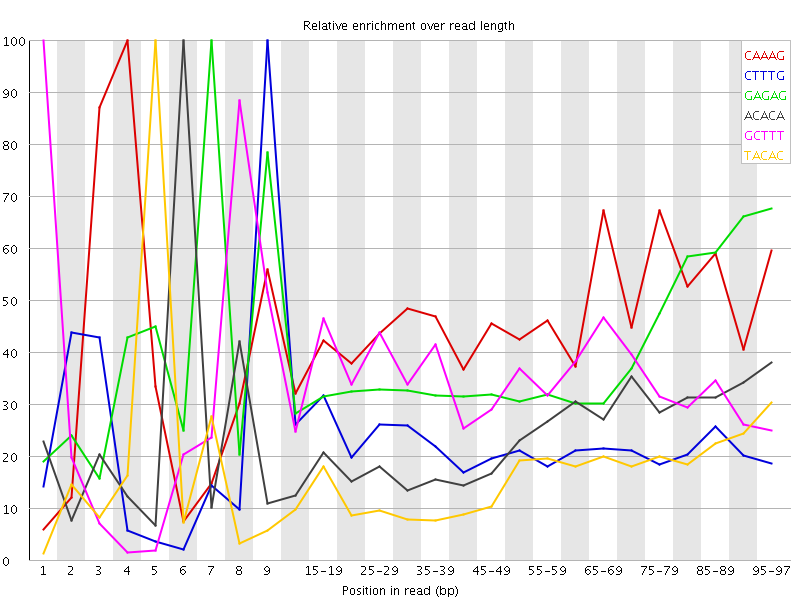
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CAAAG | 451010 | 2.4338334 | 5.2538 | 4 |
| CTTTG | 441950 | 2.3403478 | 10.393288 | 9 |
| GAGAG | 248235 | 2.0871117 | 5.323914 | 7 |
| ACACA | 397715 | 2.0795975 | 8.650107 | 6 |
| GCTTT | 369760 | 1.9580654 | 5.639155 | 1 |
| TACAC | 345195 | 1.7936568 | 10.941323 | 5 |
| GCCAA | 223935 | 1.7677058 | 5.481576 | 1 |
| TGGCT | 216960 | 1.7454207 | 5.527076 | 6 |
| CACAT | 327885 | 1.7037128 | 6.2529044 | 7 |
| GACTC | 207930 | 1.6310703 | 6.248823 | 7 |
| TTGAG | 294705 | 1.620782 | 6.082296 | 9 |
| GAGTC | 197140 | 1.5959808 | 8.961155 | 9 |
| ATGGA | 271730 | 1.503859 | 5.3666677 | 4 |
| ATACA | 419905 | 1.4915663 | 5.440594 | 6 |
| CCATG | 188355 | 1.4775175 | 5.822853 | 9 |
| AAGAC | 272420 | 1.4700891 | 6.3291545 | 5 |
| GTCTT | 275975 | 1.4614267 | 6.3376307 | 1 |
| CATGA | 270880 | 1.4526105 | 5.6186523 | 9 |
| TGTGT | 263690 | 1.4411138 | 5.228016 | 8 |
| TATGA | 386990 | 1.4097956 | 6.879693 | 4 |
| GACTT | 262195 | 1.3972179 | 7.870001 | 7 |
| TGAGT | 252840 | 1.3905381 | 6.1169605 | 8 |
| TCATA | 387335 | 1.3672433 | 7.062569 | 2 |
| GTGTT | 245645 | 1.3424946 | 6.1696715 | 1 |
| ATGAG | 242520 | 1.3421996 | 5.594751 | 7 |
| ACTCA | 257090 | 1.335857 | 5.8296614 | 8 |
| GATTG | 237815 | 1.3079054 | 6.7302566 | 7 |
| AACAC | 243225 | 1.2717903 | 5.3822327 | 5 |
| GTGTA | 229590 | 1.2626706 | 11.785161 | 1 |
| GTTCT | 234180 | 1.240101 | 5.3566837 | 1 |
| TGGAC | 148625 | 1.2032193 | 8.792373 | 5 |
| TATAC | 337095 | 1.1899025 | 6.5377398 | 5 |
| ACCAT | 228600 | 1.1878213 | 5.1696053 | 8 |
| GTCCA | 150815 | 1.1830418 | 8.570205 | 1 |
| ACACC | 154635 | 1.1827649 | 5.5812607 | 6 |
| GTATG | 213340 | 1.1733009 | 6.3542795 | 3 |
| ATGTG | 211450 | 1.1629065 | 5.215682 | 7 |
| ACATG | 216680 | 1.1619598 | 6.4116597 | 8 |
| GGACT | 138150 | 1.118417 | 8.819848 | 6 |
| ACACT | 212870 | 1.1060871 | 5.771718 | 6 |
| TGTAT | 291605 | 1.0556475 | 5.420953 | 2 |
| AGACT | 194680 | 1.0439833 | 6.1204567 | 6 |
| GAGTA | 181830 | 1.0063176 | 5.209182 | 1 |
| CATAC | 190030 | 0.9874089 | 6.192441 | 3 |
| ATACT | 225645 | 0.7964981 | 5.4441223 | 6 |
| GTATA | 218130 | 0.79464257 | 7.1493087 | 1 |