![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-152_CTCTCTAC-CTAAGCCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2104103 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
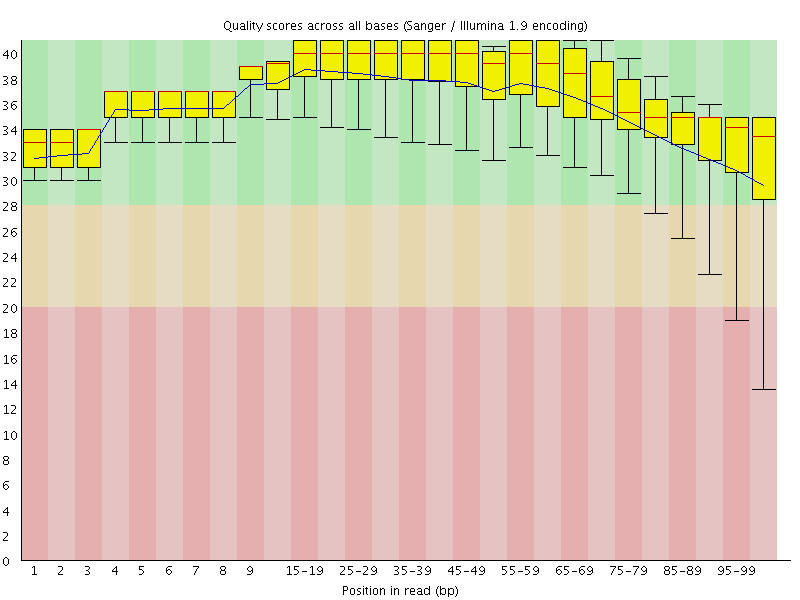
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
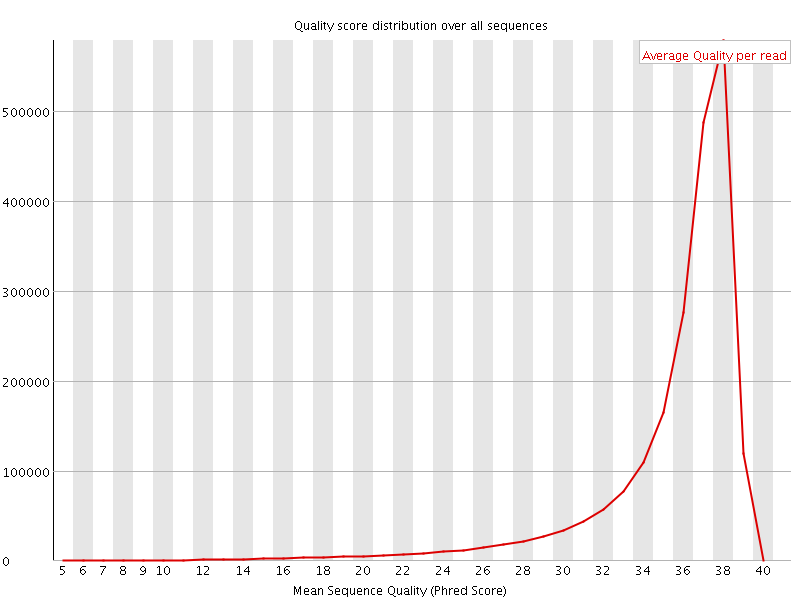
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
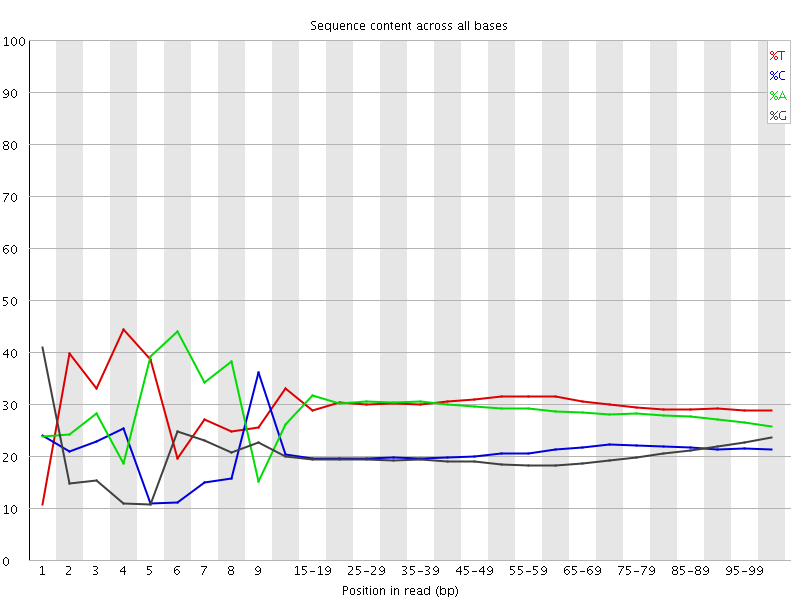
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
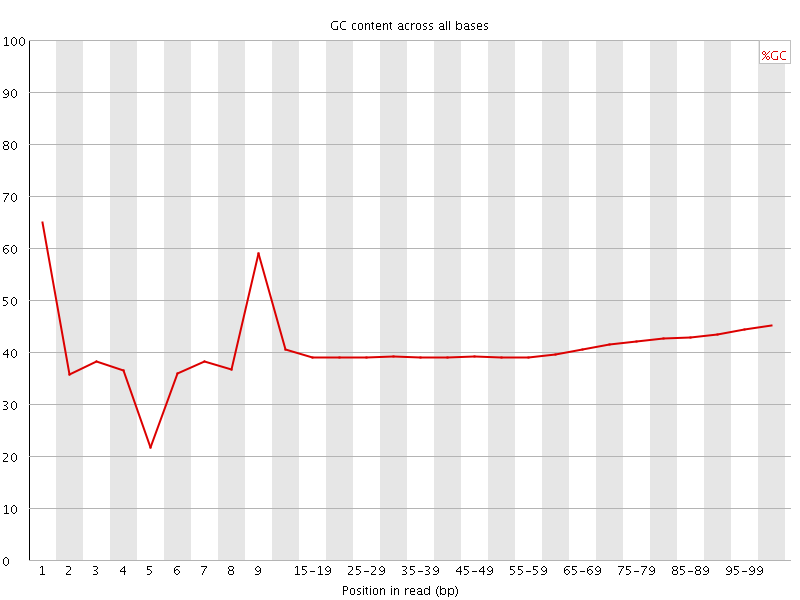
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
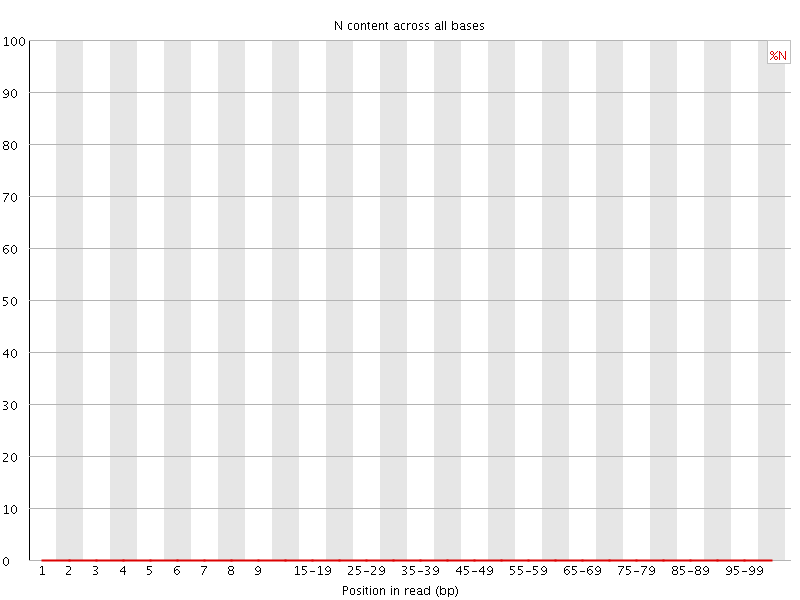
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
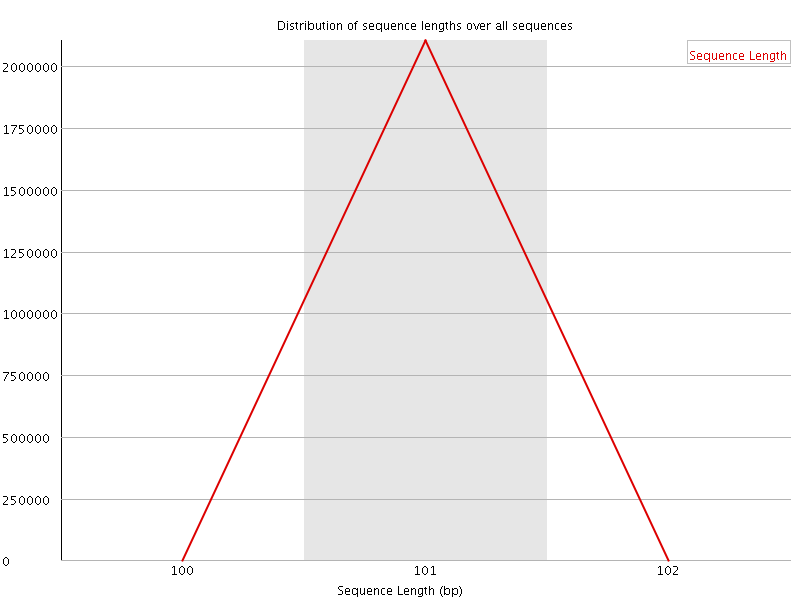
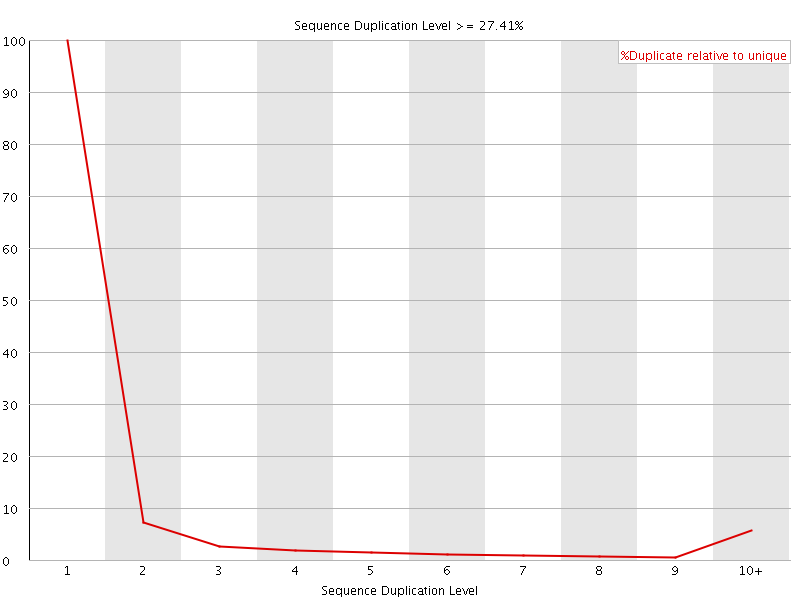
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
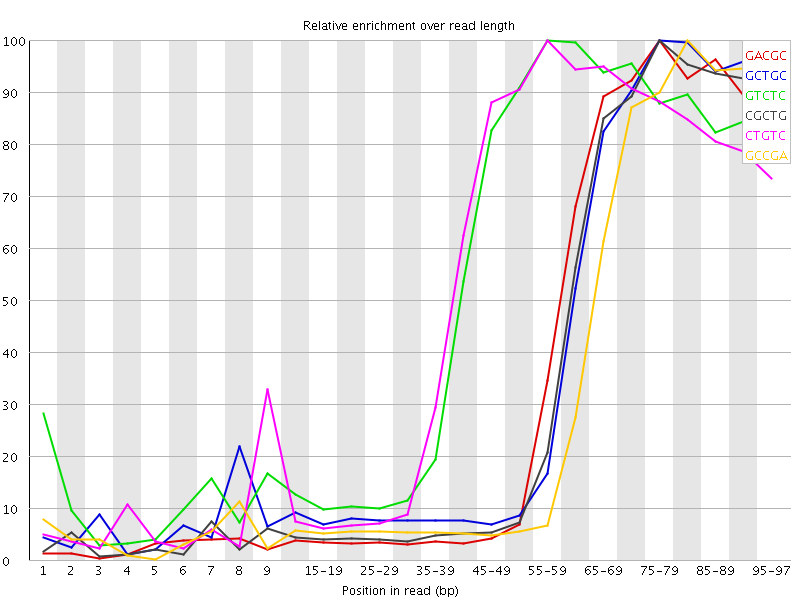
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GACGC | 579500 | 5.6835413 | 14.54592 | 75-79 |
| GCTGC | 592995 | 5.5805364 | 14.092371 | 75-79 |
| GTCTC | 885630 | 5.494222 | 9.664403 | 55-59 |
| CGCTG | 570895 | 5.3725586 | 14.163572 | 75-79 |
| CTGTC | 856085 | 5.3109326 | 9.549472 | 55-59 |
| GCCGA | 525010 | 5.1491218 | 14.786341 | 80-84 |
| CTGCC | 551790 | 4.970977 | 13.507689 | 80-84 |
| CGACG | 504370 | 4.9466915 | 14.508411 | 85-89 |
| TGCCG | 516540 | 4.8610363 | 13.905847 | 80-84 |
| CCGAC | 504745 | 4.7389336 | 14.03912 | 80-84 |
| TCTCT | 1095165 | 4.4788084 | 7.321777 | 60-64 |
| CTCTT | 1029220 | 4.2091184 | 7.090901 | 60-64 |
| TGACG | 618500 | 4.1772575 | 10.115682 | 75-79 |
| TGTCT | 975010 | 4.1653256 | 7.0621104 | 55-59 |
| CTGAC | 642755 | 4.15566 | 9.622014 | 70-74 |
| ACACA | 893505 | 4.1362042 | 7.689527 | 70-74 |
| GAAGG | 560560 | 4.1216526 | 11.719851 | 85-89 |
| TACAC | 851140 | 3.7806396 | 7.9549875 | 5 |
| ACGCT | 579015 | 3.7435565 | 9.666696 | 75-79 |
| GACGA | 530975 | 3.737372 | 10.740689 | 85-89 |
| ATCTG | 836260 | 3.7232475 | 7.673766 | 70-74 |
| CACAT | 832180 | 3.696422 | 6.9906044 | 65-69 |
| ACATC | 809310 | 3.594837 | 7.1193867 | 70-74 |
| CATCT | 830785 | 3.5408885 | 7.008643 | 70-74 |
| CGAAG | 499875 | 3.5184686 | 10.752296 | 85-89 |
| TCTGA | 774305 | 3.4474075 | 7.15982 | 70-74 |
| AAGGC | 467375 | 3.2897108 | 10.329687 | 85-89 |
| GGCTT | 498420 | 3.2300286 | 9.694152 | 90-94 |
| AGGCT | 471765 | 3.1862311 | 10.029633 | 90-94 |
| ATACA | 948020 | 3.0220976 | 5.266828 | 65-69 |
| TCTTA | 1024690 | 3.0074754 | 5.029163 | 60-64 |
| TATAC | 885285 | 2.7079048 | 5.3786316 | 5 |
| ACGAA | 547205 | 2.6461349 | 7.45975 | 85-89 |
| AGAAG | 502025 | 2.535971 | 5.4177775 | 5 |
| AGGTG | 328325 | 2.3163946 | 9.461555 | 95-97 |
| GGTGT | 327405 | 2.2164261 | 8.406656 | 95-97 |
| TTAGG | 448200 | 2.0845363 | 6.889147 | 95-97 |
| CAAAG | 420950 | 2.0356002 | 5.252059 | 4 |
| GGTGG | 195070 | 2.003224 | 10.609071 | 95-97 |
| CTTAG | 441960 | 1.967721 | 6.4592676 | 90-94 |
| GCTTA | 440200 | 1.9598851 | 6.4130597 | 90-94 |
| CTTTG | 419960 | 1.7941048 | 8.1835575 | 9 |
| AGAGA | 352285 | 1.7795618 | 5.271915 | 8 |
| GTGTA | 378340 | 1.7596238 | 8.898079 | 1 |
| GGGGG | 109880 | 1.7117037 | 5.2486205 | 95-97 |
| CTCGG | 179385 | 1.68815 | 9.799767 | 95-97 |
| AGATC | 360345 | 1.6720133 | 5.2121706 | 95-97 |
| GATCT | 369270 | 1.6440862 | 5.194872 | 95-97 |
| GAGAG | 223045 | 1.6399921 | 5.961911 | 7 |
| TAGGT | 350770 | 1.6313983 | 6.0657873 | 95-97 |
| TGTAG | 337235 | 1.5684484 | 5.3138614 | 95-97 |
| GTAGA | 301800 | 1.4628423 | 5.5054574 | 90-94 |
| TCTCG | 227185 | 1.4093977 | 6.5033097 | 95-97 |
| CGGTG | 142805 | 1.4038649 | 8.951217 | 95-97 |
| AAGAC | 266015 | 1.2863764 | 5.86429 | 5 |
| GAGTC | 181780 | 1.2277153 | 6.681698 | 9 |
| GGTCG | 119390 | 1.1736805 | 5.23055 | 95-97 |
| TATGA | 360490 | 1.1518621 | 5.621709 | 4 |
| GTCTT | 262995 | 1.1235371 | 6.1444974 | 1 |
| GATTG | 237735 | 1.1056831 | 5.304871 | 7 |
| TCATA | 360235 | 1.1018848 | 5.501836 | 2 |
| CGCCG | 80575 | 1.1011316 | 5.170021 | 95-97 |
| TGAGT | 235600 | 1.0957534 | 5.099708 | 8 |
| GACTT | 245390 | 1.0925403 | 5.9074035 | 7 |
| GTGTT | 244490 | 1.0910835 | 5.489944 | 1 |
| TCGGT | 162475 | 1.052925 | 5.8693705 | 95-97 |
| TGGAC | 145560 | 0.98309076 | 6.431102 | 5 |
| AGACT | 201105 | 0.9331342 | 5.2361794 | 6 |
| GTCCA | 143390 | 0.927072 | 7.1695614 | 1 |
| GGACT | 134820 | 0.91055447 | 6.029791 | 6 |
| GTATA | 215875 | 0.68977845 | 6.9579067 | 1 |