![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-152_CTCTCTAC-CTAAGCCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2104103 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
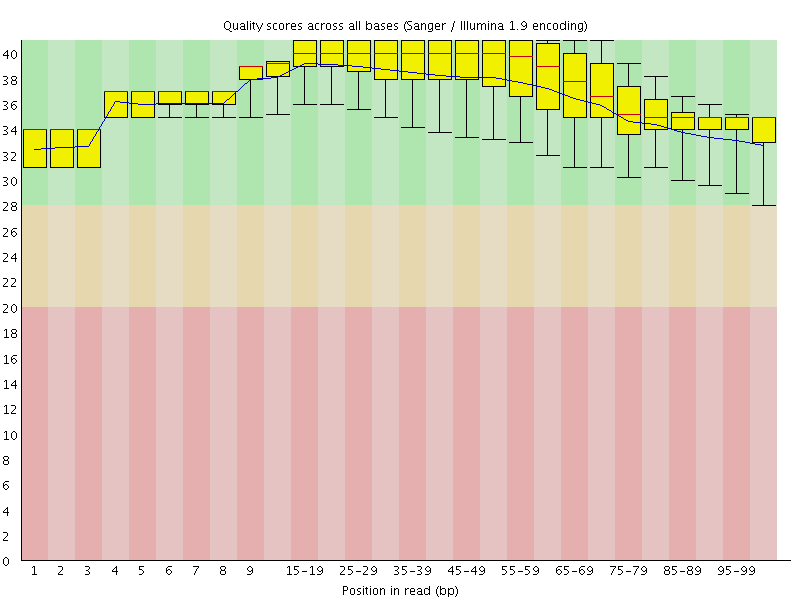
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
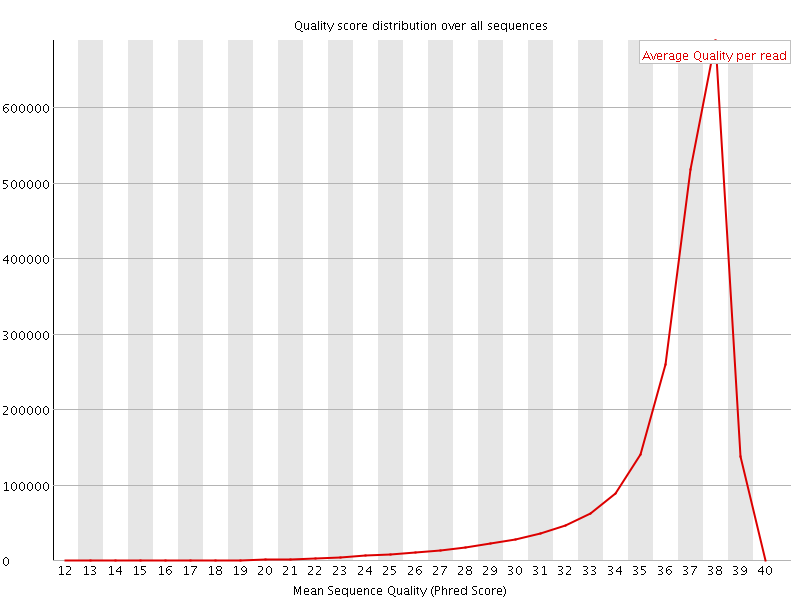
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
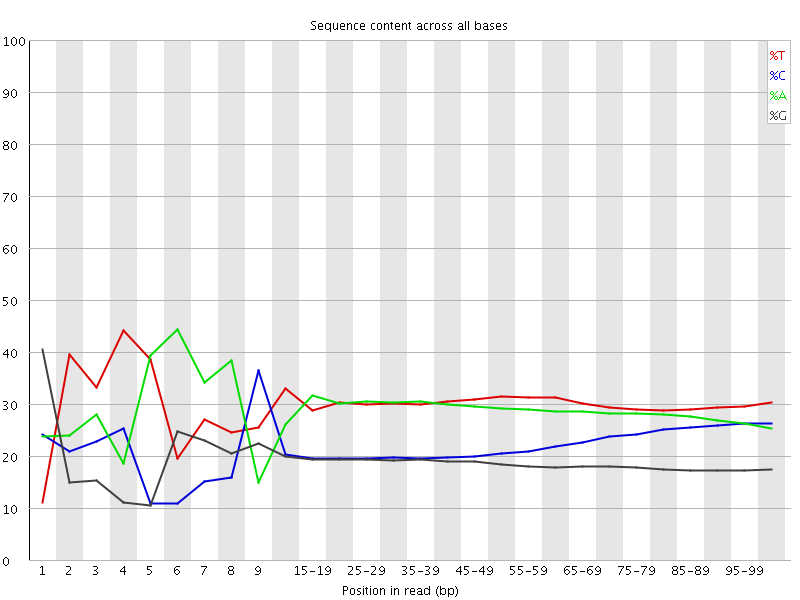
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
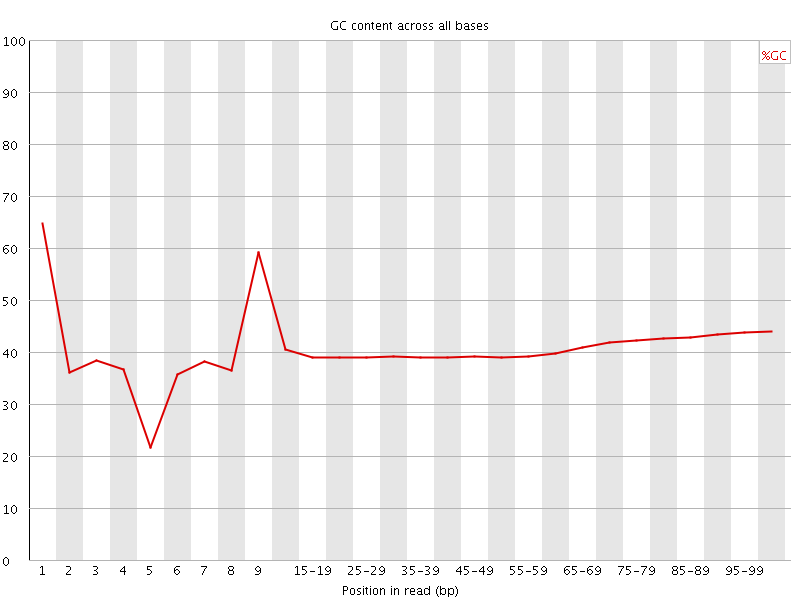
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
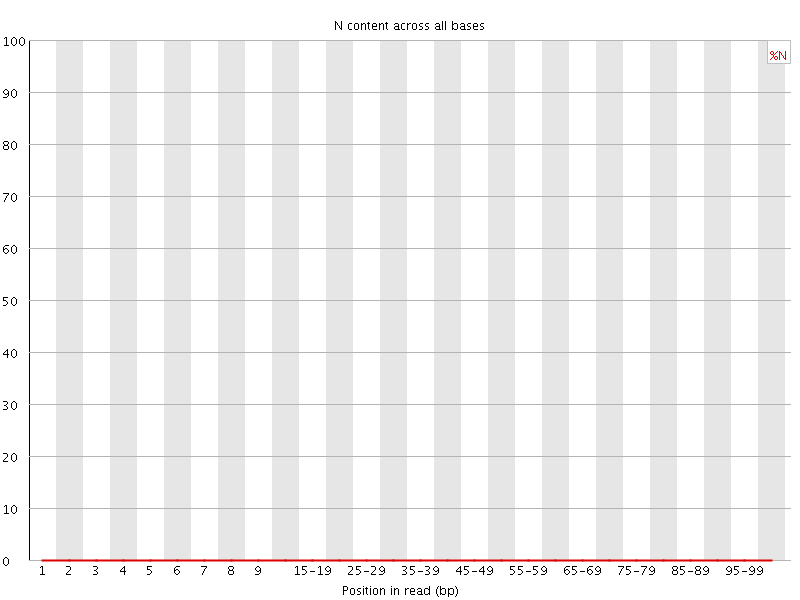
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
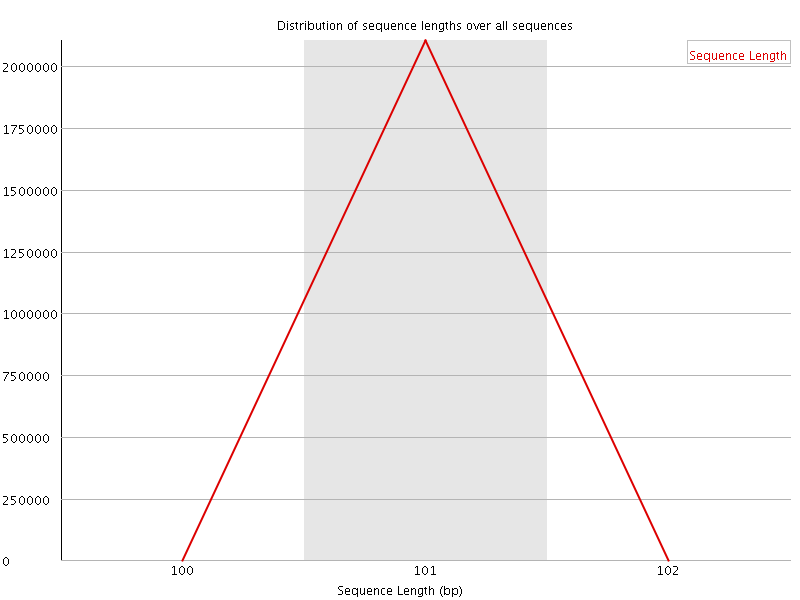
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
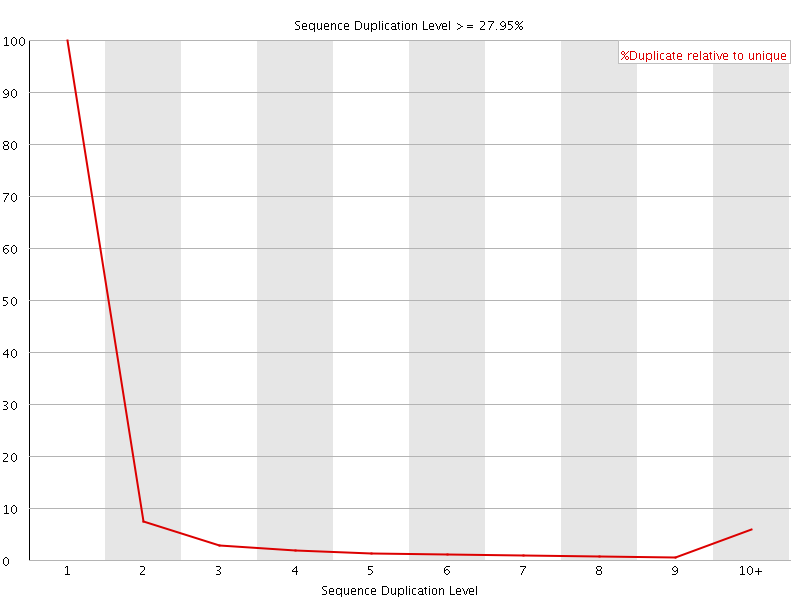
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
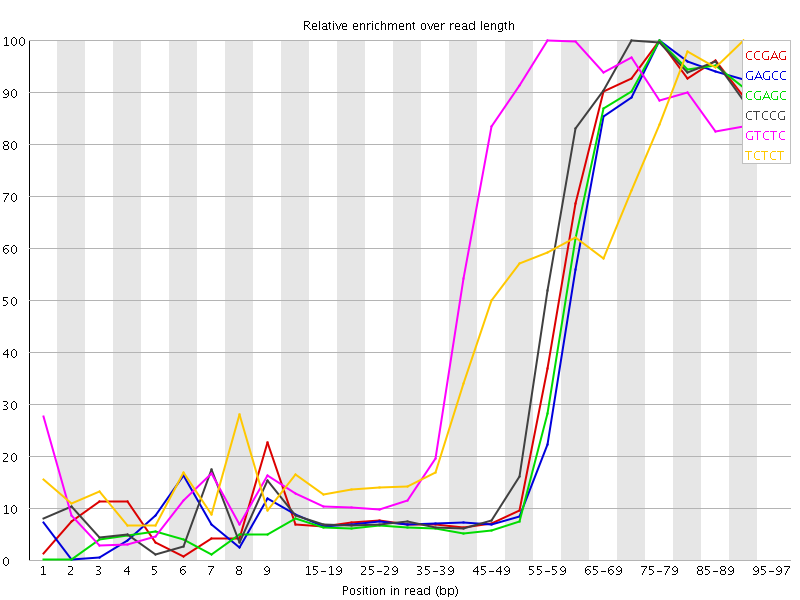
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 623920 | 6.1859303 | 14.984848 | 75-79 |
| GAGCC | 604600 | 5.99438 | 15.152137 | 75-79 |
| CGAGC | 596315 | 5.912237 | 14.973311 | 75-79 |
| CTCCG | 651145 | 5.268516 | 12.066894 | 70-74 |
| GTCTC | 884830 | 5.217813 | 9.153941 | 55-59 |
| TCTCT | 1412165 | 5.162688 | 10.665321 | 90-94 |
| CTGTC | 858905 | 5.0649343 | 9.0664015 | 55-59 |
| AGCCC | 577070 | 4.8668537 | 12.558548 | 75-79 |
| GCCCA | 566025 | 4.7737036 | 12.633945 | 80-84 |
| CCACG | 519730 | 4.3832636 | 12.596324 | 80-84 |
| TGTCT | 979310 | 4.2088833 | 7.1006317 | 55-59 |
| ACATC | 1052615 | 4.180982 | 11.049544 | 95-97 |
| CGAGA | 544835 | 4.1036315 | 11.577029 | 85-89 |
| TCCGA | 666855 | 4.098918 | 9.4032135 | 75-79 |
| GAGAC | 541840 | 4.0810738 | 11.7362585 | 85-89 |
| CATCT | 1058510 | 4.033615 | 10.764542 | 95-97 |
| ACGAG | 533015 | 4.0146046 | 11.298284 | 80-84 |
| ATCTC | 1041840 | 3.9700923 | 10.686325 | 95-97 |
| CCCAC | 548630 | 3.9358904 | 10.753148 | 80-84 |
| TCTCC | 776840 | 3.8967626 | 8.154678 | 70-74 |
| CTCTT | 1031235 | 3.7700582 | 6.3337245 | 60-64 |
| ACACA | 895330 | 3.706816 | 6.886357 | 70-74 |
| TACAC | 854745 | 3.3950434 | 7.343129 | 5 |
| CACGA | 529160 | 3.3902676 | 9.673475 | 80-84 |
| CACAT | 836300 | 3.3217797 | 6.28204 | 65-69 |
| AGACC | 498195 | 3.1918788 | 9.849699 | 85-89 |
| GACCT | 494750 | 3.0410504 | 9.574783 | 85-89 |
| ATACA | 948515 | 2.8620696 | 5.0076537 | 65-69 |
| AGAAG | 498250 | 2.8508728 | 5.897927 | 5 |
| CTCTC | 558330 | 2.800679 | 8.132406 | 90-94 |
| GAAGA | 462320 | 2.64529 | 5.2570896 | 6 |
| CCTCT | 519710 | 2.6069543 | 7.949503 | 90-94 |
| TATAC | 888745 | 2.5727878 | 5.252138 | 5 |
| ACCTC | 478825 | 2.5035617 | 7.9403143 | 85-89 |
| CAAAG | 422105 | 2.054446 | 5.040587 | 4 |
| AGAGA | 355150 | 2.0320876 | 6.011669 | 8 |
| GAGAG | 222950 | 1.9740912 | 6.9504375 | 7 |
| TCTCG | 329820 | 1.9449376 | 8.123297 | 90-94 |
| CTCTA | 474105 | 1.8066504 | 6.001337 | 90-94 |
| CTTTG | 417650 | 1.7949783 | 8.287201 | 9 |
| TGGAG | 210790 | 1.7906078 | 5.32131 | 5 |
| CTACA | 448835 | 1.7827705 | 6.146838 | 95-97 |
| AAGAG | 303140 | 1.7344984 | 5.4318643 | 7 |
| GTATG | 324725 | 1.7101253 | 7.5660534 | 95-97 |
| CTCGT | 288645 | 1.70213 | 7.977472 | 90-94 |
| TCTAC | 433390 | 1.6514995 | 5.8263288 | 95-97 |
| AGGAG | 177390 | 1.5706842 | 5.5423193 | 7 |
| TTGGA | 292570 | 1.540785 | 5.0965743 | 4 |
| ATGGA | 278910 | 1.5310364 | 5.102102 | 4 |
| TTGAG | 288440 | 1.519035 | 5.339147 | 9 |
| TGCCG | 151770 | 1.4436198 | 12.143484 | 95-97 |
| GCCGT | 149170 | 1.4188889 | 11.662295 | 95-97 |
| GTCTT | 313880 | 1.348995 | 5.7351 | 1 |
| ATGCC | 219440 | 1.3488189 | 8.224699 | 95-97 |
| GAGTC | 181355 | 1.3104597 | 6.957913 | 9 |
| AAGAC | 266015 | 1.2947336 | 6.0623913 | 5 |
| ATGAG | 231460 | 1.2705666 | 5.147348 | 7 |
| TGAGT | 237365 | 1.2500544 | 5.4387283 | 8 |
| GATTG | 235685 | 1.2412068 | 6.082184 | 7 |
| TATGA | 361490 | 1.2302105 | 5.9516153 | 4 |
| GTGTT | 241970 | 1.2225443 | 6.2374535 | 1 |
| CCGTC | 149370 | 1.2085761 | 9.052041 | 95-97 |
| TCGTA | 266700 | 1.1947551 | 6.0008106 | 95-97 |
| TATGC | 259490 | 1.1624559 | 6.1007276 | 95-97 |
| CGTAT | 246785 | 1.1055404 | 6.130413 | 95-97 |
| GACTT | 246145 | 1.1026734 | 6.005603 | 7 |
| GTGTA | 205945 | 1.0845847 | 10.000018 | 1 |
| TCATA | 364205 | 1.0543206 | 5.3579016 | 2 |
| TGGAC | 143795 | 1.0390536 | 7.490442 | 5 |
| GAGTA | 179690 | 0.9863824 | 5.203721 | 1 |
| GGACT | 133925 | 0.9677335 | 6.7617173 | 6 |
| AGACT | 199220 | 0.9302465 | 5.508239 | 6 |
| CGTCT | 157065 | 0.92620707 | 6.0139847 | 95-97 |
| GTCCA | 145490 | 0.8942747 | 6.7894936 | 1 |
| GGATA | 162600 | 0.89256924 | 5.2649407 | 1 |
| GTATA | 215390 | 0.7330079 | 7.13864 | 1 |