![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-151_CTCTCTAC-AAGGAGTA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1766755 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
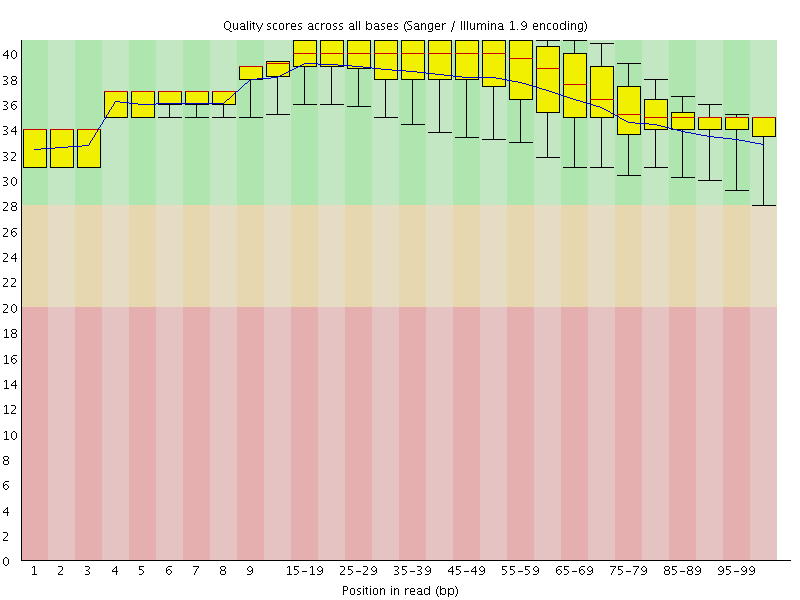
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
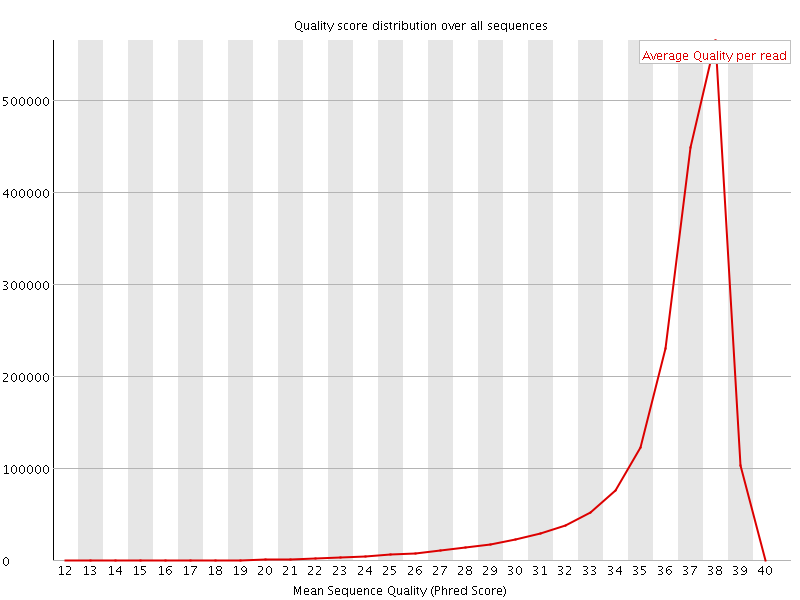
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
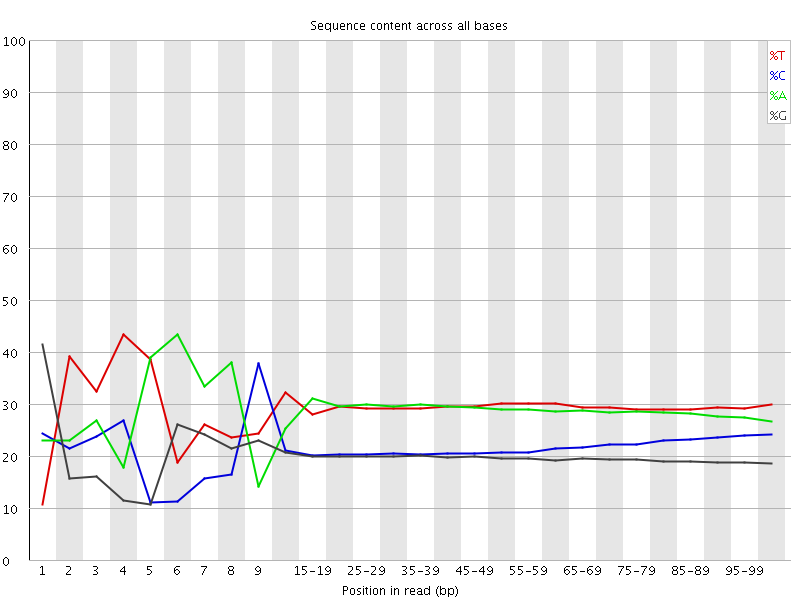
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
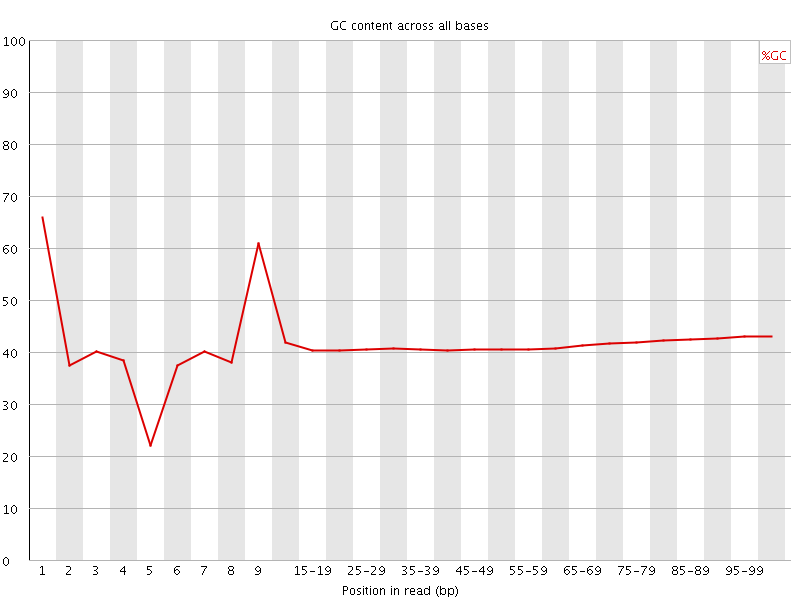
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
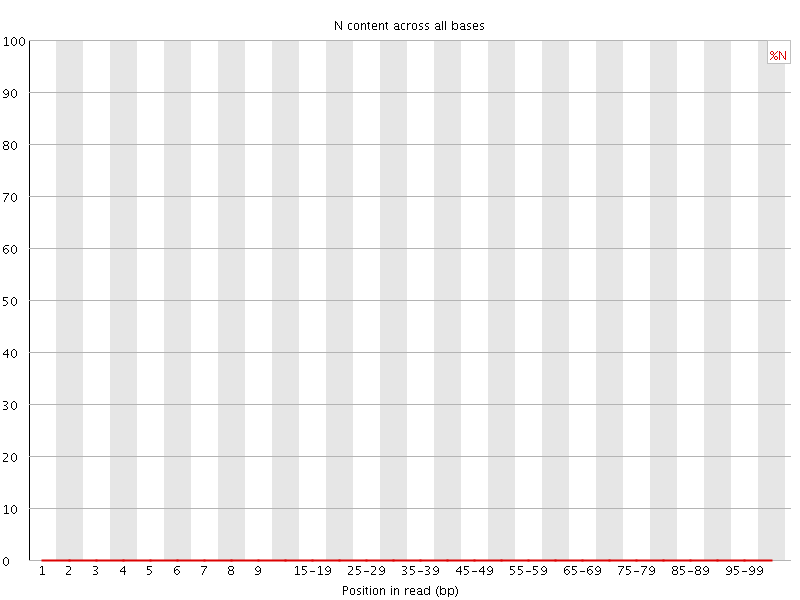
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
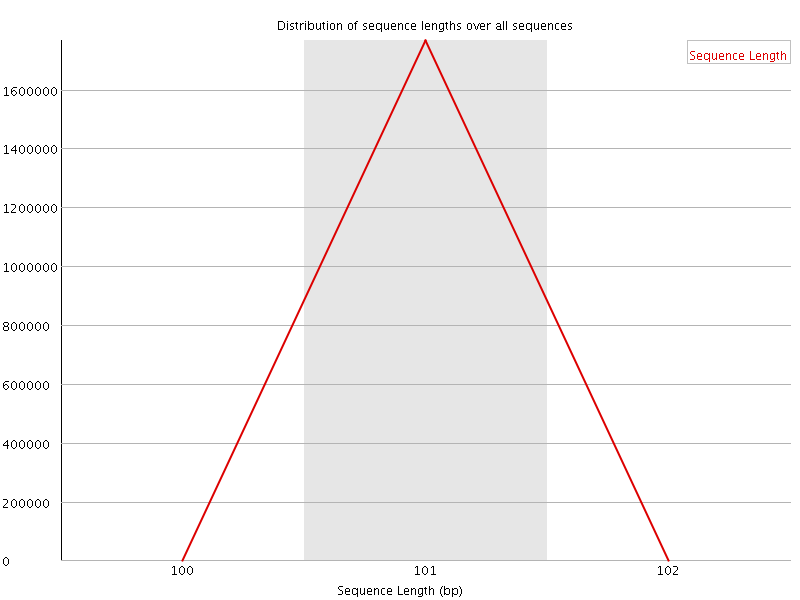
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
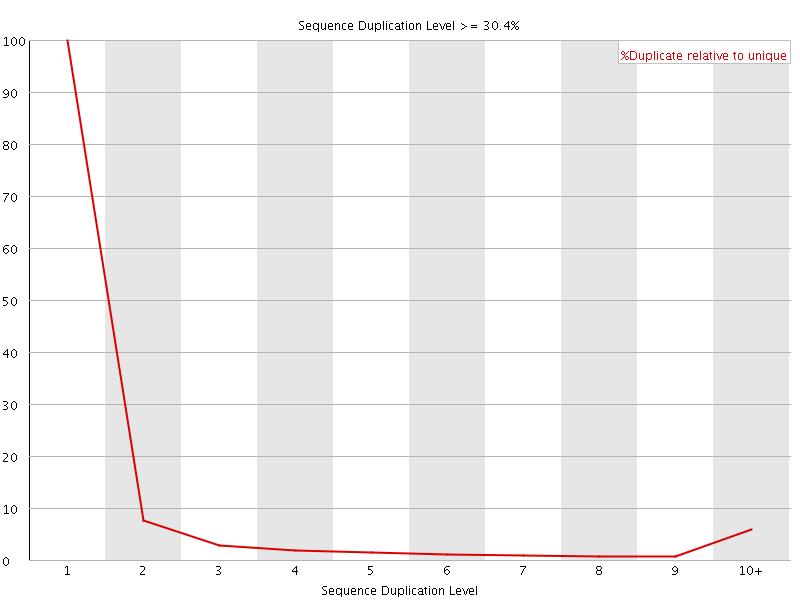
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
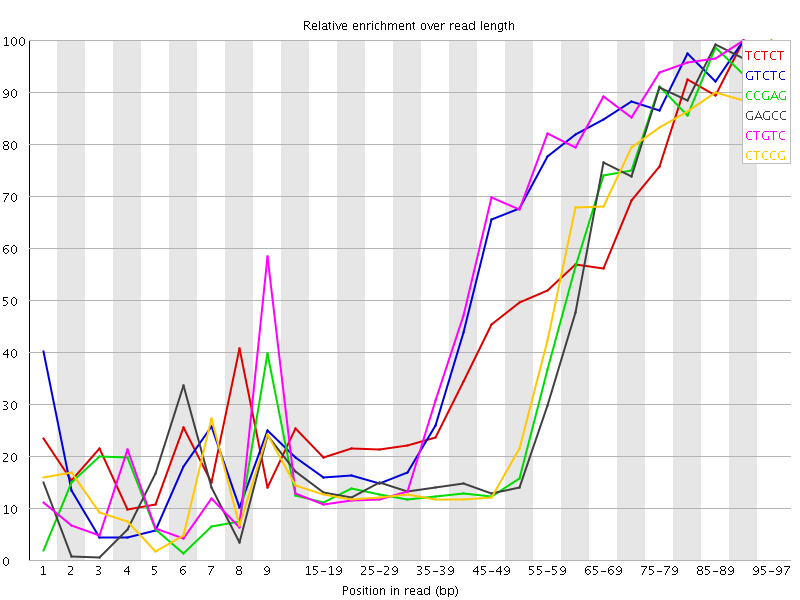
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TCTCT | 793805 | 3.8087513 | 7.7670283 | 90-94 |
| GTCTC | 487700 | 3.4872313 | 6.236754 | 90-94 |
| CCGAG | 315450 | 3.4435008 | 8.227113 | 95-97 |
| GAGCC | 310240 | 3.3866277 | 8.066558 | 95-97 |
| CTGTC | 462775 | 3.3090084 | 5.924708 | 90-94 |
| CTCCG | 333250 | 3.2540262 | 7.77897 | 95-97 |
| CGAGC | 297625 | 3.2489204 | 8.031271 | 95-97 |
| TGTCT | 562265 | 2.9440567 | 5.00781 | 90-94 |
| AGCCC | 289425 | 2.8951383 | 7.1291747 | 90-94 |
| TCTCC | 440605 | 2.8869622 | 6.0279403 | 95-97 |
| GCCCA | 287240 | 2.8732817 | 7.3484144 | 90-94 |
| AGAAG | 453470 | 2.7857187 | 5.4208517 | 5 |
| CATCT | 563615 | 2.7703443 | 7.019712 | 95-97 |
| ATCTC | 560220 | 2.7536564 | 6.9069004 | 95-97 |
| ACATC | 545055 | 2.7445676 | 7.0268073 | 95-97 |
| ACACA | 528825 | 2.727897 | 7.2505617 | 6 |
| TCCGA | 348235 | 2.5508378 | 5.8130536 | 95-97 |
| CCCAC | 274705 | 2.5180454 | 6.7826476 | 90-94 |
| CCACG | 250200 | 2.5027683 | 7.0923123 | 90-94 |
| TACAC | 483880 | 2.436527 | 8.957539 | 5 |
| GAGAC | 294265 | 2.4097266 | 6.574091 | 95-97 |
| CGAGA | 289235 | 2.3685362 | 6.372911 | 95-97 |
| ACGAG | 274725 | 2.2497144 | 6.109524 | 95-97 |
| CTCTC | 340890 | 2.2336028 | 5.174962 | 90-94 |
| CAAAG | 384765 | 2.1659474 | 5.082056 | 4 |
| CACGA | 263010 | 1.973627 | 5.378656 | 90-94 |
| CTTTG | 376245 | 1.9700438 | 8.222872 | 9 |
| AGAGA | 318690 | 1.9577495 | 5.2004433 | 8 |
| AGACC | 257855 | 1.9349438 | 5.605768 | 95-97 |
| GACCT | 257280 | 1.8845881 | 5.398681 | 95-97 |
| GAGAG | 207380 | 1.8532431 | 6.2826223 | 7 |
| TATAC | 493085 | 1.8181617 | 6.008785 | 5 |
| TGGAG | 200720 | 1.7509501 | 5.100795 | 5 |
| GCTTT | 319660 | 1.673761 | 5.0524983 | 1 |
| TTGAG | 263055 | 1.5398217 | 5.545694 | 9 |
| AAGAC | 241685 | 1.360511 | 5.846275 | 5 |
| GTCTT | 258020 | 1.3510098 | 6.045225 | 1 |
| GAGTC | 168845 | 1.3496928 | 6.379461 | 9 |
| GTATG | 222360 | 1.301609 | 5.7727437 | 3 |
| CCATG | 174025 | 1.2747414 | 5.4303193 | 9 |
| TATGA | 316615 | 1.2740269 | 6.5182495 | 4 |
| GATTG | 216985 | 1.2701459 | 6.3432064 | 7 |
| GTGTT | 217915 | 1.2451699 | 5.8905077 | 1 |
| AGAAC | 218305 | 1.2288985 | 5.073868 | 5 |
| GACTT | 224895 | 1.2063333 | 6.2105384 | 7 |
| TCATA | 320505 | 1.1818042 | 6.206022 | 2 |
| GTTCT | 221100 | 1.1576942 | 5.1108937 | 1 |
| GTGTA | 194675 | 1.1395518 | 10.74614 | 1 |
| TGGAC | 136710 | 1.092816 | 7.5111766 | 5 |
| ACATG | 198090 | 1.0885102 | 5.528349 | 8 |
| GAGTA | 169615 | 1.0171161 | 5.277532 | 1 |
| GTCCA | 138835 | 1.0169729 | 7.0575876 | 1 |
| GGACT | 125460 | 1.0028871 | 6.6740174 | 6 |
| GCCGT | 93685 | 0.9982913 | 5.680025 | 95-97 |
| AGACT | 178590 | 0.98135716 | 5.3605003 | 6 |
| TGTAT | 246265 | 0.9673138 | 5.129193 | 2 |
| TGCCG | 87715 | 0.93467605 | 5.7523603 | 95-97 |
| GGATA | 152180 | 0.9125651 | 5.120515 | 1 |
| GTATA | 188665 | 0.7591689 | 7.0944533 | 1 |