![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-150_CTCTCTAC-ACTGCATA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3195394 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
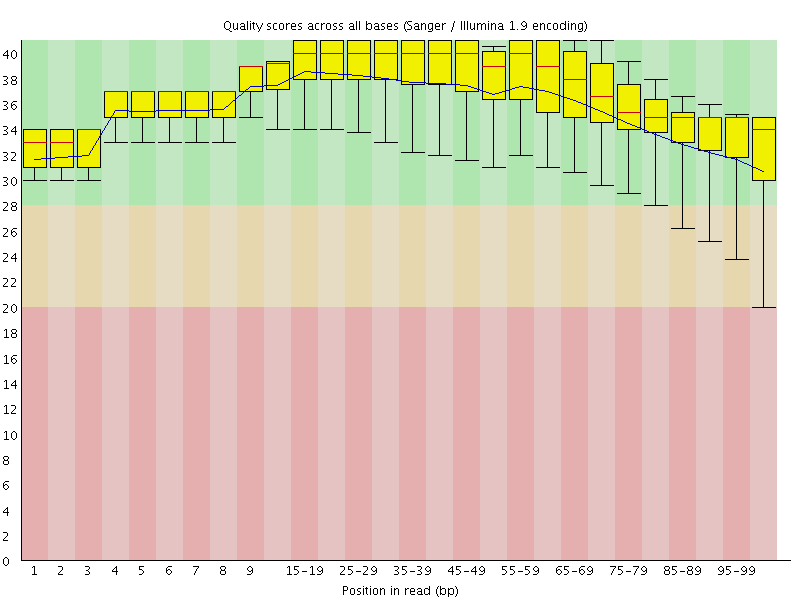
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
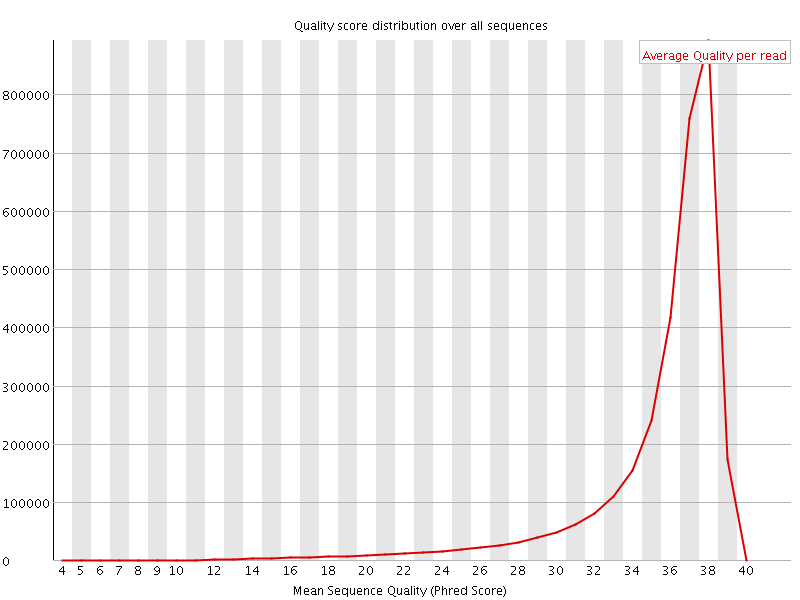
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
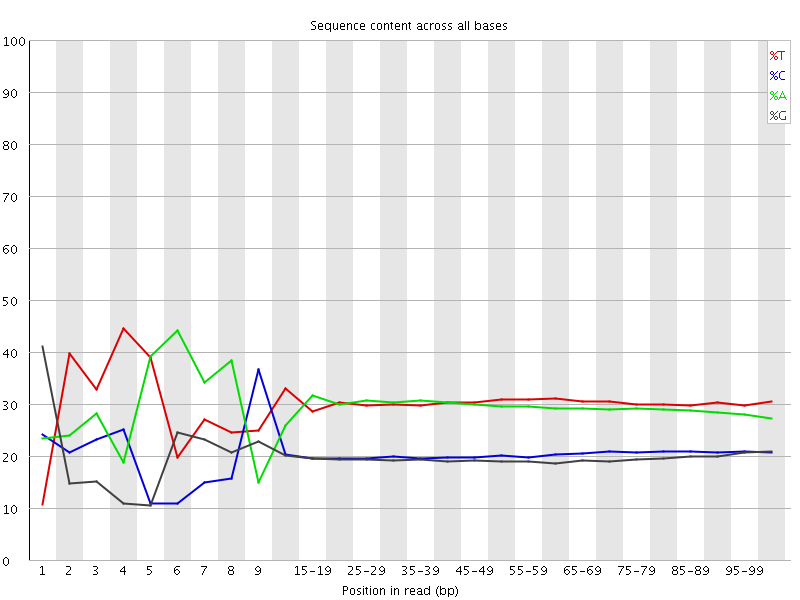
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
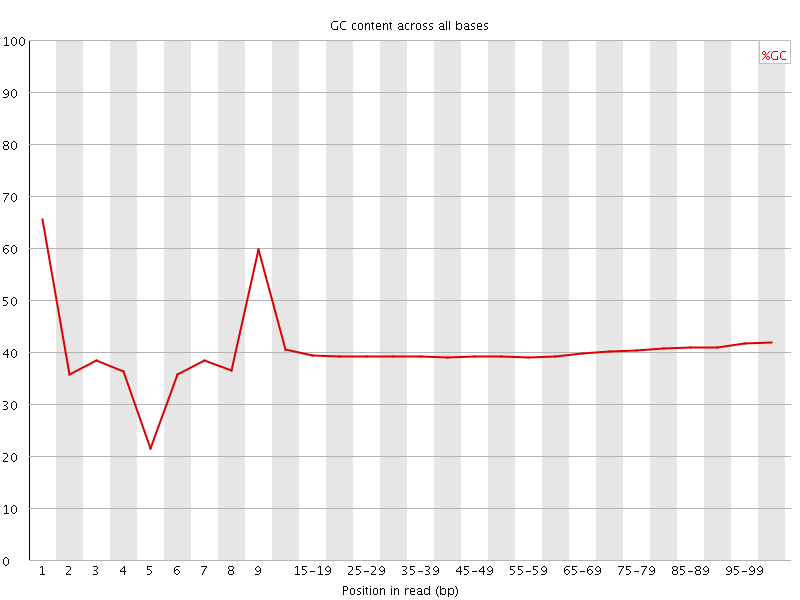
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
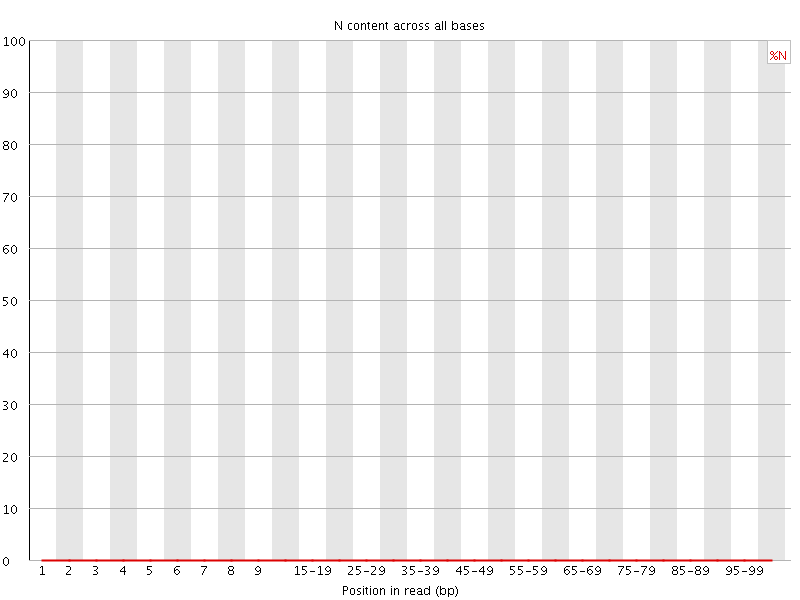
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
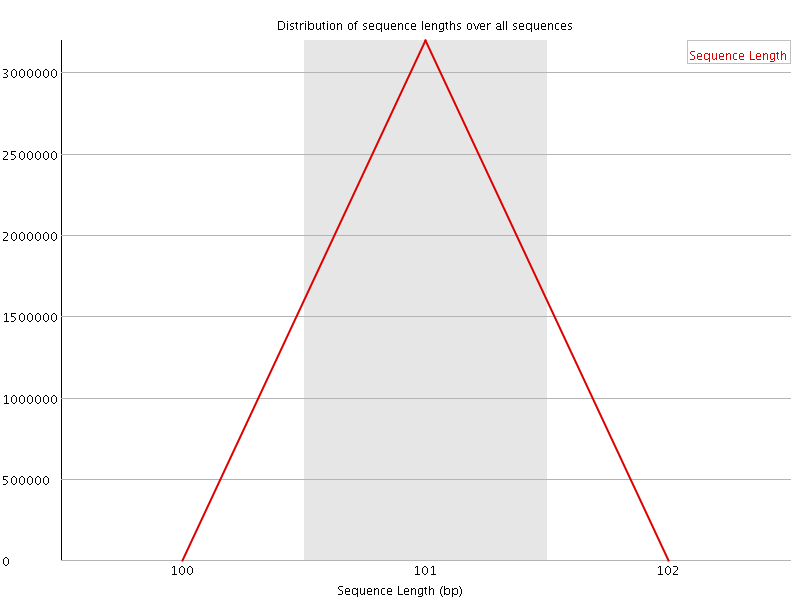
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
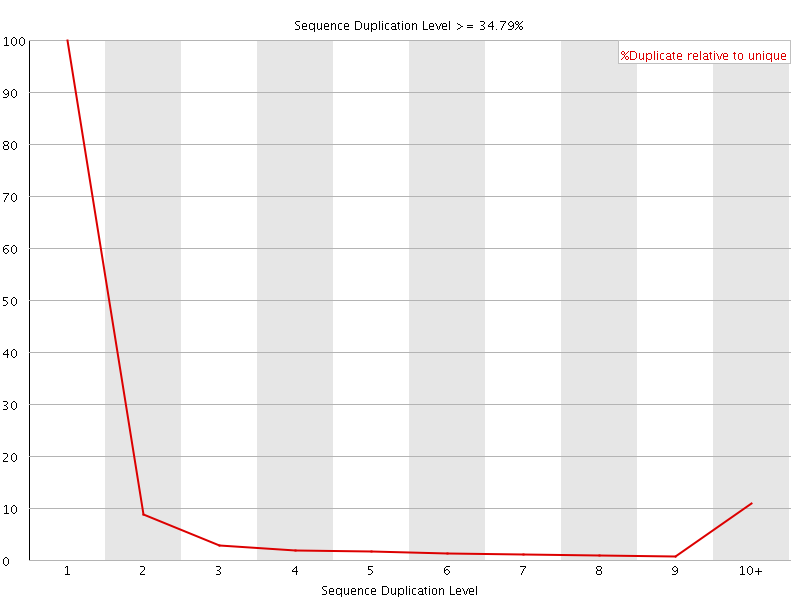
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 3305 | 0.10343012473579159 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
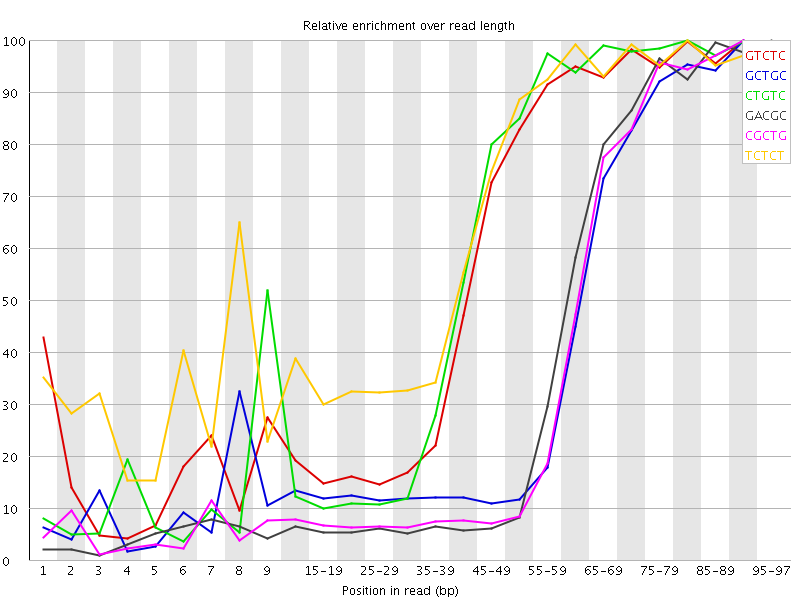
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 894115 | 3.814438 | 6.360658 | 90-94 |
| GCTGC | 565435 | 3.721699 | 9.196754 | 90-94 |
| CTGTC | 841815 | 3.591318 | 5.9712787 | 80-84 |
| GACGC | 524540 | 3.5527878 | 8.994789 | 95-97 |
| CGCTG | 517515 | 3.4062886 | 8.848803 | 90-94 |
| TCTCT | 1206200 | 3.3353024 | 4.9578743 | 80-84 |
| GCCGA | 489885 | 3.3180642 | 9.281693 | 95-97 |
| CTGCC | 514050 | 3.2665806 | 8.816745 | 90-94 |
| CGACG | 475750 | 3.2223258 | 9.361632 | 95-97 |
| CTCTT | 1129320 | 3.122719 | 4.6916018 | 70-74 |
| TGCCG | 469935 | 3.0931165 | 8.831407 | 95-97 |
| ACACA | 1012745 | 3.0514927 | 7.741217 | 6 |
| CCGAC | 466425 | 3.050015 | 8.860569 | 95-97 |
| TACAC | 944295 | 2.7649531 | 9.19673 | 5 |
| AGAAG | 848910 | 2.7441945 | 5.195032 | 5 |
| CTGAC | 606490 | 2.6625206 | 6.1919947 | 95-97 |
| TGACG | 577565 | 2.6262777 | 6.331249 | 95-97 |
| CACAT | 892590 | 2.6135578 | 5.4654336 | 7 |
| GACGA | 528810 | 2.4744098 | 6.7503843 | 95-97 |
| ACGCT | 534200 | 2.345164 | 5.8832073 | 85-89 |
| CAAAG | 750050 | 2.3408465 | 5.268361 | 4 |
| ATACA | 1074270 | 2.1730762 | 5.355983 | 6 |
| CTTTG | 738420 | 2.1148999 | 9.593412 | 9 |
| TGCAG | 448210 | 2.0380807 | 6.1776004 | 95-97 |
| TATAC | 948760 | 1.8650295 | 5.764201 | 5 |
| GCTTT | 617975 | 1.7699349 | 5.4628625 | 1 |
| GCCAA | 382450 | 1.727731 | 5.535478 | 1 |
| GGTGG | 244395 | 1.7258055 | 5.8990707 | 95-97 |
| GCAGT | 366600 | 1.6669873 | 5.7570853 | 95-97 |
| GTGTG | 364195 | 1.66691 | 5.885853 | 95-97 |
| GAGAG | 342290 | 1.6589626 | 5.516354 | 7 |
| TTGGC | 371195 | 1.6402493 | 5.0093336 | 5 |
| TGGCT | 370385 | 1.6366699 | 5.5264225 | 6 |
| GTGTA | 531710 | 1.6231693 | 10.471047 | 1 |
| CAGTG | 353065 | 1.6054416 | 5.7931085 | 95-97 |
| GACTC | 342545 | 1.5037892 | 5.760945 | 7 |
| TTGAG | 490515 | 1.4974121 | 5.504219 | 9 |
| GAGTC | 328190 | 1.4923309 | 8.622116 | 9 |
| AAGAC | 459305 | 1.4334549 | 6.099492 | 5 |
| CTCGG | 216845 | 1.4272758 | 6.0550313 | 95-97 |
| CATGA | 455455 | 1.3813258 | 5.149204 | 9 |
| CCATG | 309945 | 1.3606737 | 5.382 | 9 |
| TATGA | 645590 | 1.314488 | 6.3201222 | 4 |
| GACTT | 441950 | 1.3025417 | 7.282145 | 7 |
| GTCTT | 453905 | 1.3000238 | 6.4812436 | 1 |
| TCATA | 648555 | 1.2749 | 6.457421 | 2 |
| TGAGT | 416185 | 1.2705022 | 6.311129 | 8 |
| ATGAG | 403145 | 1.2664336 | 5.7356515 | 7 |
| ACTCA | 424395 | 1.2426544 | 5.2865934 | 8 |
| AACAC | 401860 | 1.2108408 | 5.1038756 | 5 |
| GATTG | 394455 | 1.2041663 | 6.1881647 | 7 |
| GTGTT | 401195 | 1.1901793 | 5.8598294 | 1 |
| CGGTG | 169180 | 1.1533949 | 5.365685 | 95-97 |
| GTTCT | 398435 | 1.141153 | 5.103025 | 1 |
| TGGAC | 248425 | 1.1296271 | 7.909766 | 5 |
| ACATG | 361640 | 1.096799 | 5.6611056 | 8 |
| GTCCA | 248255 | 1.0898515 | 8.315927 | 1 |
| GGACT | 231715 | 1.0536442 | 7.770712 | 6 |
| ACACT | 348810 | 1.0213369 | 5.1046276 | 6 |
| AGACT | 333185 | 1.0104995 | 5.8153033 | 6 |
| GAGTA | 306805 | 0.9637926 | 5.1521277 | 1 |
| GTATG | 307295 | 0.93808997 | 5.3930902 | 3 |
| CATAC | 307060 | 0.89909035 | 5.2822022 | 3 |
| GTATA | 360360 | 0.7337302 | 7.206177 | 1 |