![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-150_CTCTCTAC-ACTGCATA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3195394 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
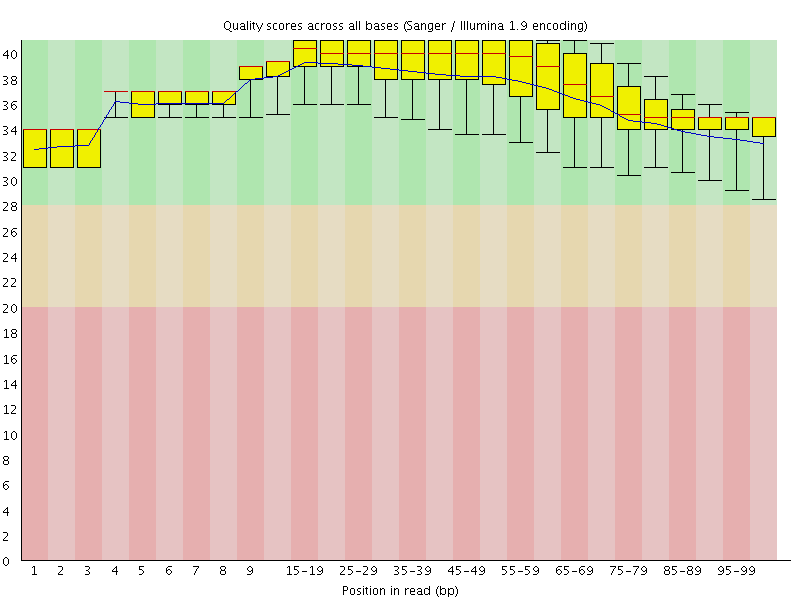
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
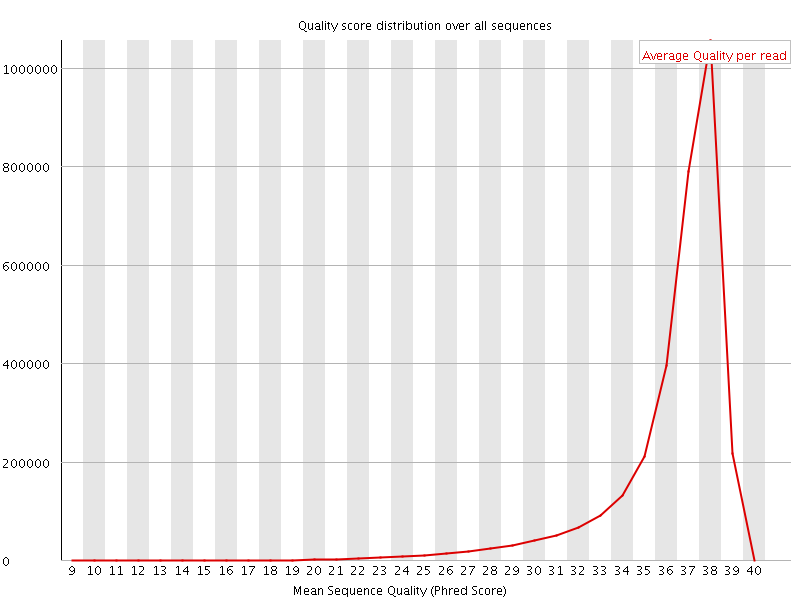
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
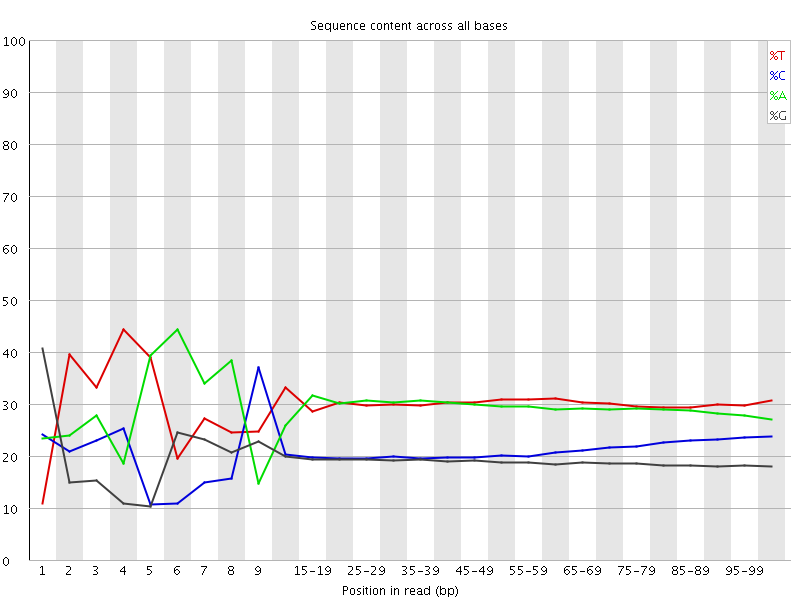
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
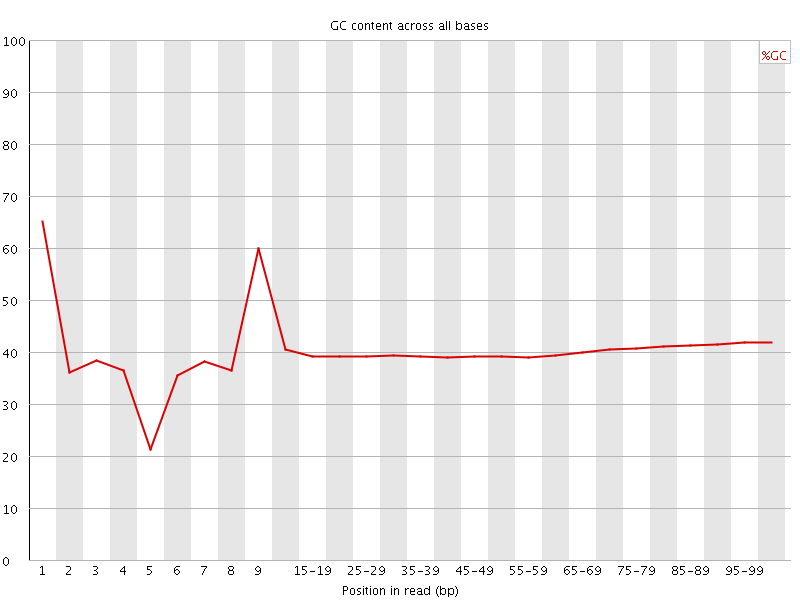
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
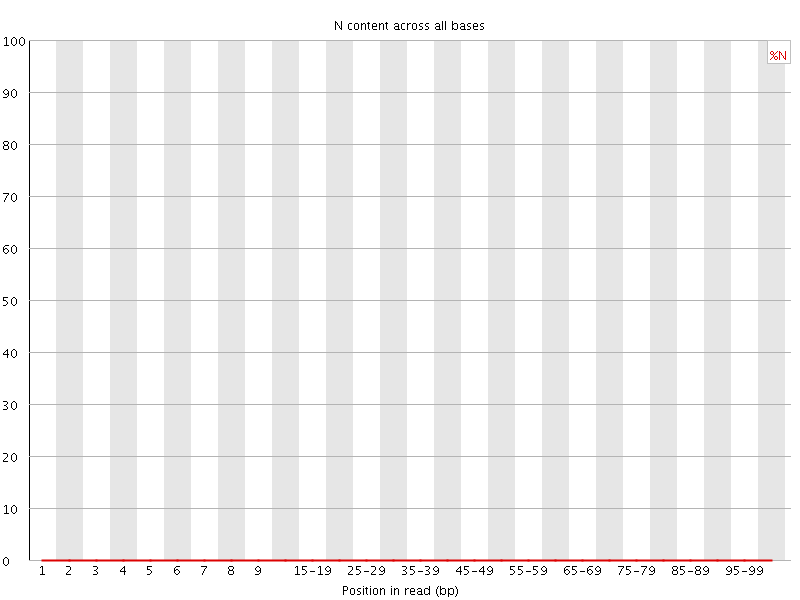
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
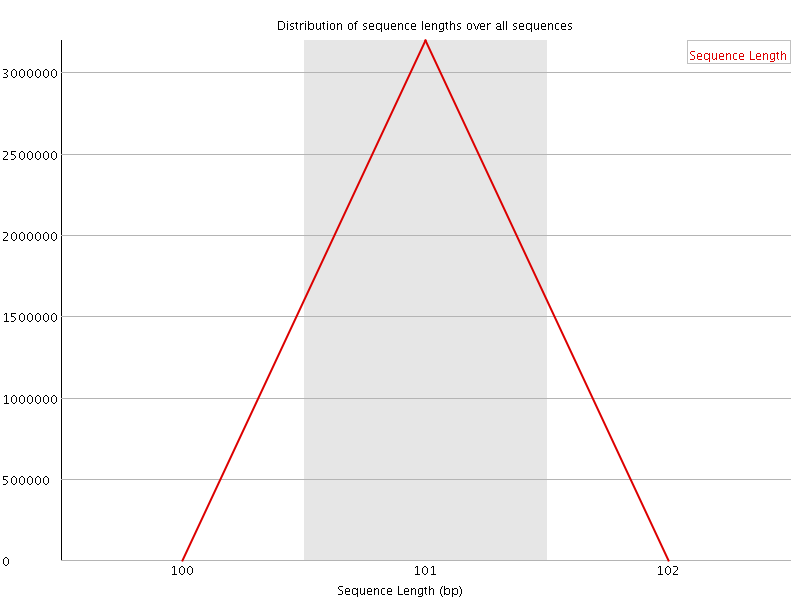
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
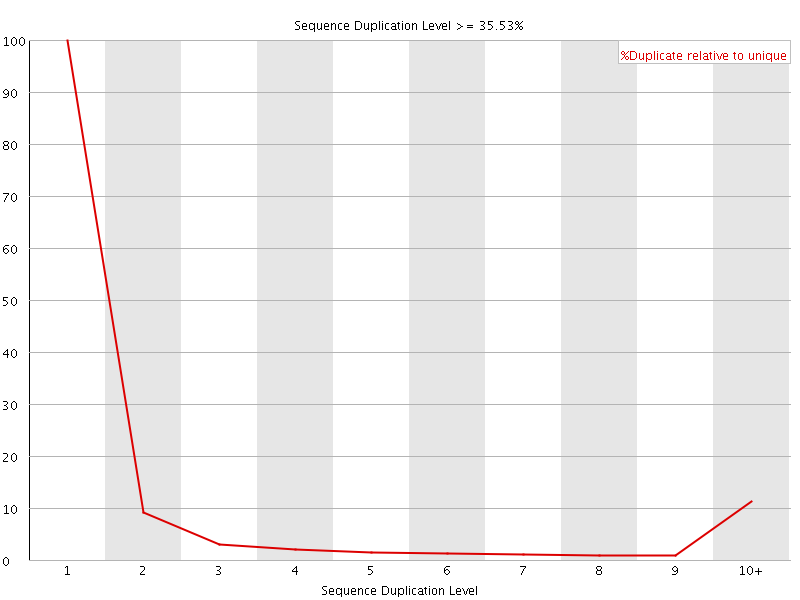
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 3424 | 0.10715423512718618 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
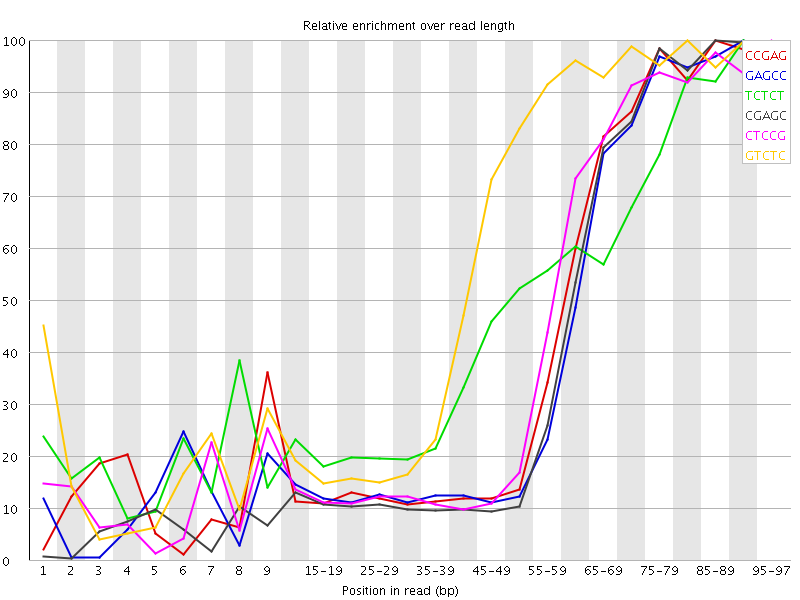
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 604790 | 4.062657 | 9.363495 | 85-89 |
| GAGCC | 582020 | 3.9097002 | 9.353536 | 90-94 |
| TCTCT | 1457565 | 3.810166 | 7.7710447 | 90-94 |
| CGAGC | 562400 | 3.7779036 | 9.157632 | 85-89 |
| CTCCG | 625500 | 3.7158415 | 8.40715 | 95-97 |
| GTCTC | 897225 | 3.709359 | 6.1677084 | 80-84 |
| CTGTC | 845980 | 3.4974988 | 5.7964196 | 75-79 |
| AGCCC | 550560 | 3.360169 | 8.465642 | 90-94 |
| GCCCA | 547600 | 3.3421035 | 8.570996 | 90-94 |
| TGTCT | 1047775 | 3.0146239 | 4.823765 | 90-94 |
| TCTCC | 802155 | 3.0130522 | 5.969633 | 95-97 |
| CCACG | 479760 | 2.9280634 | 8.203217 | 95-97 |
| AGAAG | 846730 | 2.9076288 | 5.2013164 | 5 |
| CCCAC | 522035 | 2.894723 | 7.6253514 | 90-94 |
| ACACA | 1012090 | 2.8688972 | 7.0257854 | 6 |
| ACATC | 1030335 | 2.842809 | 7.424113 | 95-97 |
| TCCGA | 666395 | 2.830452 | 6.171331 | 85-89 |
| CATCT | 1050975 | 2.8225067 | 7.339623 | 95-97 |
| ATCTC | 1050260 | 2.8205864 | 7.158183 | 95-97 |
| CGAGA | 552350 | 2.652859 | 6.9515014 | 95-97 |
| GAGAC | 547570 | 2.6299012 | 7.0042877 | 95-97 |
| TACAC | 950135 | 2.6215281 | 8.568338 | 5 |
| ACGAG | 518680 | 2.4911466 | 6.843599 | 95-97 |
| CACAT | 895560 | 2.4709496 | 5.04736 | 7 |
| CAAAG | 751185 | 2.343645 | 5.2687383 | 4 |
| CTCTC | 611300 | 2.2961633 | 5.749654 | 90-94 |
| CACGA | 506000 | 2.2080107 | 6.117632 | 95-97 |
| ATACA | 1079795 | 2.1301234 | 5.2223296 | 6 |
| CTTTG | 738920 | 2.1259966 | 9.688256 | 9 |
| AGACC | 481770 | 2.1022794 | 6.0054936 | 85-89 |
| CCTCT | 546650 | 2.0533252 | 5.5165205 | 90-94 |
| GACCT | 481700 | 2.0459769 | 5.801018 | 85-89 |
| TATAC | 954360 | 1.8325213 | 5.7535186 | 5 |
| ACCTC | 474790 | 1.8322153 | 5.289895 | 90-94 |
| AGAGA | 531680 | 1.8257627 | 5.0364857 | 8 |
| GAGAG | 338065 | 1.787101 | 5.971907 | 7 |
| GCTTT | 616425 | 1.7735578 | 5.4314804 | 1 |
| TGGCT | 369275 | 1.6803373 | 5.5068007 | 6 |
| GCCAA | 384400 | 1.67739 | 5.357758 | 1 |
| TTGAG | 490055 | 1.5943573 | 6.0446973 | 9 |
| ATGGA | 464290 | 1.551875 | 5.4128017 | 4 |
| GAGTC | 327660 | 1.5317808 | 8.762807 | 9 |
| GACTC | 344190 | 1.4619159 | 5.6900125 | 7 |
| AAGAC | 459055 | 1.4322197 | 6.186948 | 5 |
| GTCTT | 487945 | 1.4038993 | 6.222554 | 1 |
| CATGA | 455555 | 1.3834362 | 5.159298 | 9 |
| TATGA | 646805 | 1.3669709 | 6.6666584 | 4 |
| GTATG | 417200 | 1.357329 | 5.7165923 | 3 |
| TGAGT | 414725 | 1.3492768 | 6.655161 | 8 |
| ATGAG | 401165 | 1.3408817 | 6.13883 | 7 |
| CCATG | 313735 | 1.3325608 | 5.255487 | 9 |
| GACTT | 446025 | 1.3184112 | 7.3493333 | 7 |
| GTGTT | 403485 | 1.2777373 | 6.2914224 | 1 |
| GATTG | 391910 | 1.2750498 | 6.6677804 | 7 |
| TCATA | 652375 | 1.2526627 | 6.23664 | 2 |
| GTGTA | 383960 | 1.249185 | 10.813184 | 1 |
| ACTCA | 426005 | 1.175395 | 5.1904993 | 8 |
| TGGAC | 248765 | 1.1629537 | 8.372946 | 5 |
| GTTCT | 399790 | 1.1502626 | 5.000366 | 1 |
| ATGTG | 348140 | 1.1326474 | 5.164493 | 7 |
| AACAC | 398615 | 1.1299247 | 5.076927 | 5 |
| ACATG | 362050 | 1.0994788 | 5.686418 | 8 |
| GGACT | 232320 | 1.086075 | 8.051084 | 6 |
| GTCCA | 249315 | 1.0589428 | 7.9646544 | 1 |
| GCCGT | 161740 | 1.0575389 | 6.532071 | 95-97 |
| GAGTA | 309155 | 1.033341 | 5.4686637 | 1 |
| CCGTC | 171765 | 1.0203861 | 5.3413634 | 95-97 |
| AGACT | 332980 | 1.0111988 | 5.8719416 | 6 |
| TGCCG | 151055 | 0.9876749 | 6.8068385 | 95-97 |
| ACACT | 347375 | 0.95844626 | 5.0259557 | 6 |
| GGATA | 269310 | 0.9001604 | 5.0618367 | 1 |
| GTATA | 362240 | 0.76556545 | 7.1923037 | 1 |