![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-149_CTCTCTAC-GTAAGGAG_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2298827 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 42 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
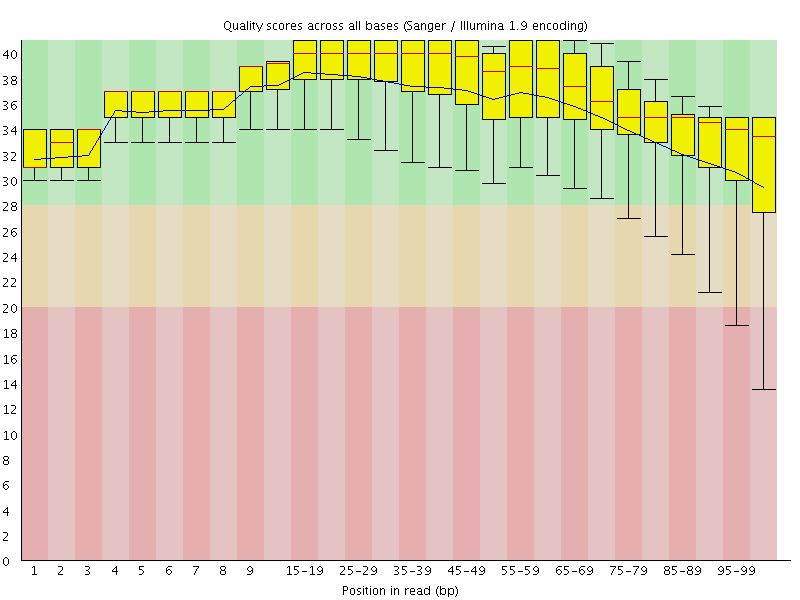
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
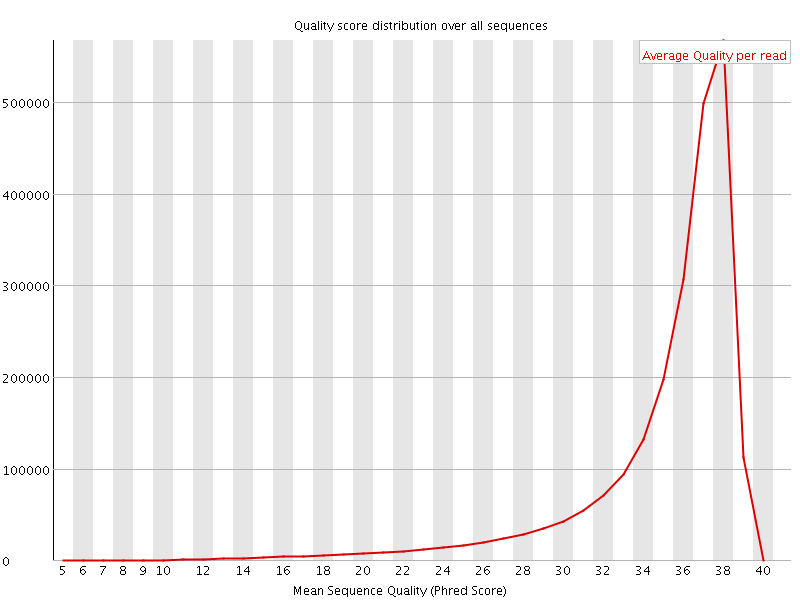
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
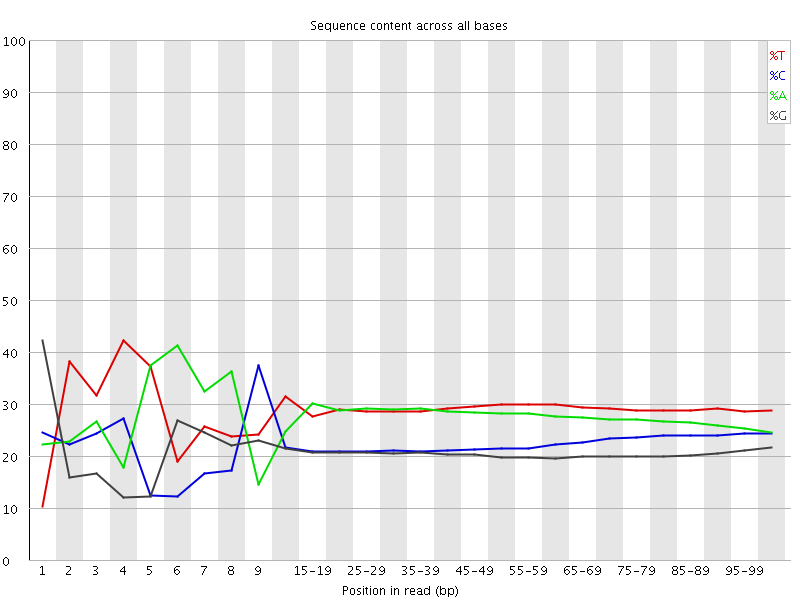
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
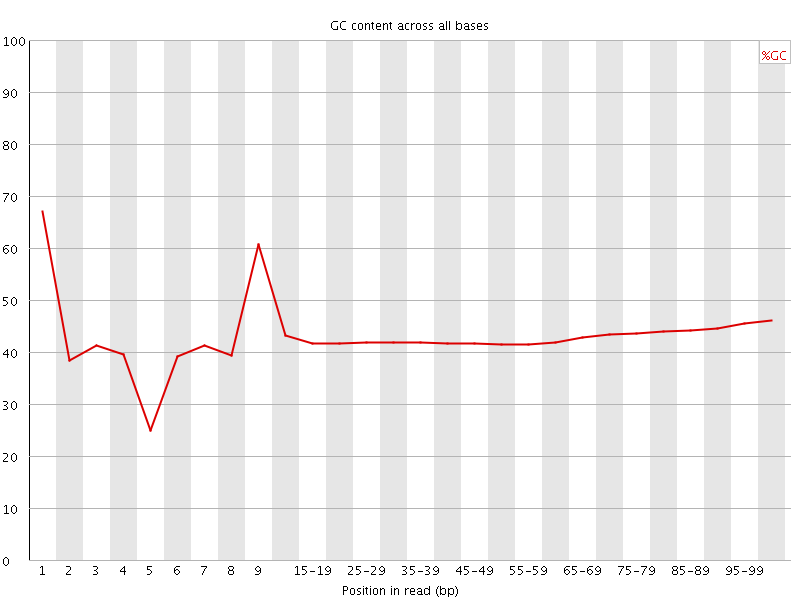
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
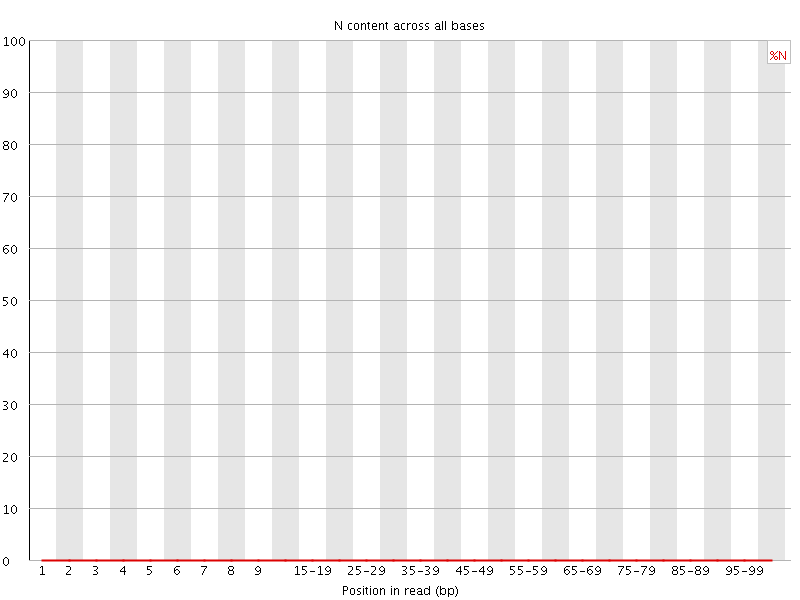
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
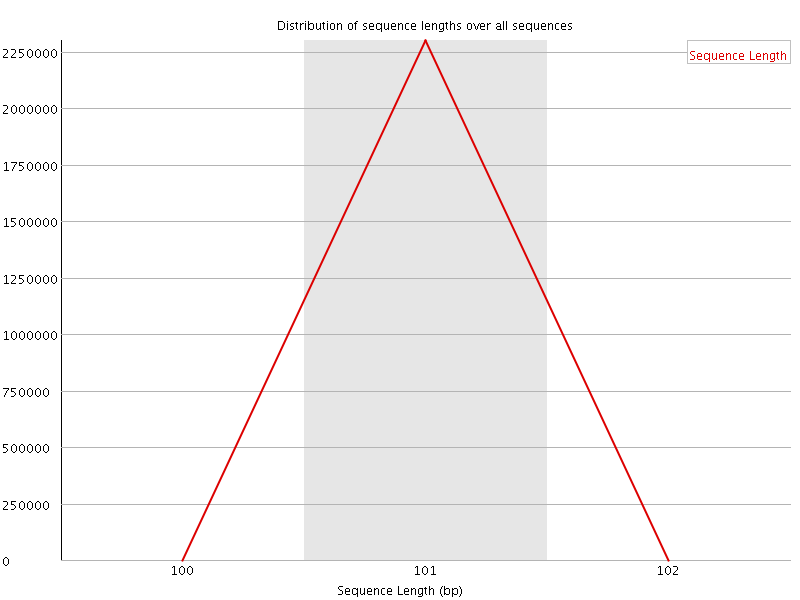
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
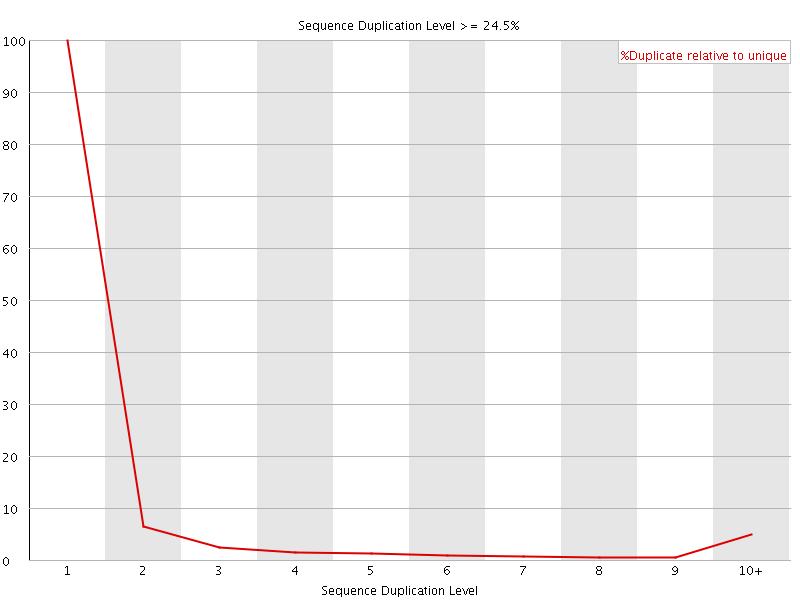
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
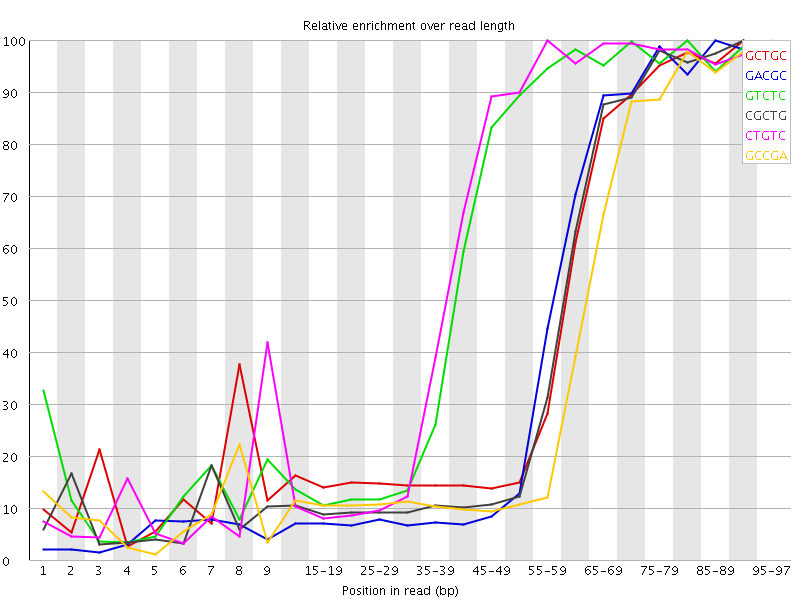
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 657435 | 4.7369924 | 10.61352 | 90-94 |
| GACGC | 600220 | 4.5541315 | 10.69582 | 85-89 |
| GTCTC | 884495 | 4.496643 | 7.4131417 | 80-84 |
| CGCTG | 614055 | 4.424428 | 10.382183 | 90-94 |
| CTGTC | 869900 | 4.4224443 | 7.192576 | 55-59 |
| GCCGA | 555945 | 4.2181973 | 10.823422 | 95-97 |
| CGACG | 534520 | 4.055637 | 10.808702 | 95-97 |
| TGCCG | 550030 | 3.9631116 | 10.197876 | 95-97 |
| CTGCC | 584620 | 3.885049 | 9.530383 | 90-94 |
| TCTCT | 1079080 | 3.8707032 | 5.988625 | 60-64 |
| TGTCT | 979010 | 3.8075924 | 6.0266075 | 80-84 |
| ACACA | 894635 | 3.7472935 | 6.5859094 | 70-74 |
| CTCTT | 1021355 | 3.6636412 | 5.754466 | 60-64 |
| CCGAC | 515235 | 3.6055658 | 9.83764 | 95-97 |
| TGACG | 619685 | 3.5969658 | 8.221879 | 85-89 |
| CTGAC | 639510 | 3.4236202 | 7.6038456 | 95-97 |
| TACAC | 844870 | 3.3605976 | 7.515155 | 5 |
| ATCTG | 817615 | 3.3485548 | 6.5785427 | 95-97 |
| CACAT | 823720 | 3.2764704 | 5.975305 | 90-94 |
| GACGA | 535120 | 3.2708583 | 8.5281315 | 85-89 |
| ACATC | 801240 | 3.1870525 | 5.9618006 | 90-94 |
| CATCT | 822550 | 3.1070175 | 5.795266 | 80-84 |
| ACGCT | 575765 | 3.082361 | 7.371388 | 75-79 |
| TCTGA | 749320 | 3.0688517 | 6.0649323 | 85-89 |
| ATACA | 932830 | 2.9891286 | 5.036208 | 90-94 |
| GACTC | 528745 | 2.8306391 | 7.6993713 | 85-89 |
| TATAC | 866375 | 2.6363482 | 5.2606378 | 5 |
| ACGAC | 465095 | 2.6219542 | 7.6635404 | 95-97 |
| AGAAG | 526460 | 2.5923364 | 5.0565076 | 5 |
| CGACT | 466745 | 2.4987218 | 7.0258646 | 85-89 |
| ACTCC | 487985 | 2.4094486 | 6.4282765 | 90-94 |
| CTCCT | 510830 | 2.3952034 | 6.390567 | 90-94 |
| TCCTT | 584515 | 2.0966787 | 5.147401 | 90-94 |
| GGTGG | 222360 | 1.8834786 | 7.5746217 | 95-97 |
| CGCCG | 199950 | 1.88322 | 5.204811 | 95-97 |
| GTGTA | 397660 | 1.7658244 | 9.3763895 | 1 |
| CTTTG | 452670 | 1.7605364 | 7.702789 | 9 |
| ACGTG | 297865 | 1.7289594 | 7.086358 | 95-97 |
| CGTGT | 278820 | 1.5368944 | 6.4647384 | 95-97 |
| GAGAG | 229695 | 1.522261 | 5.190442 | 7 |
| CTCGG | 211260 | 1.5221843 | 7.074675 | 95-97 |
| CGGTG | 184470 | 1.4411286 | 6.73347 | 95-97 |
| TTACG | 350815 | 1.4367679 | 5.2065496 | 95-97 |
| AAGAC | 285825 | 1.298073 | 5.64173 | 5 |
| TACGT | 307570 | 1.2596576 | 5.009889 | 95-97 |
| TATGA | 380290 | 1.2546968 | 5.9198685 | 4 |
| GAGTC | 201975 | 1.1723653 | 6.2969503 | 9 |
| TCATA | 378820 | 1.152736 | 5.85546 | 2 |
| GTGTT | 263080 | 1.1093746 | 5.635382 | 1 |
| GATTG | 248985 | 1.1056272 | 5.4309382 | 7 |
| GTCTT | 284115 | 1.1049877 | 5.6389804 | 1 |
| GACTT | 267885 | 1.0971271 | 5.7954617 | 7 |
| TGGAC | 169195 | 0.9820936 | 6.636341 | 5 |
| GTCCA | 170295 | 0.9116752 | 6.8089843 | 1 |
| GAATA | 261665 | 0.90910655 | 5.1098685 | 1 |
| GGACT | 151590 | 0.8799052 | 6.099708 | 6 |
| TATAA | 354755 | 0.8696427 | 5.1772923 | 2 |
| GTATA | 218770 | 0.7217914 | 7.0626755 | 1 |