![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-149_CTCTCTAC-GTAAGGAG_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2298827 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 42 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
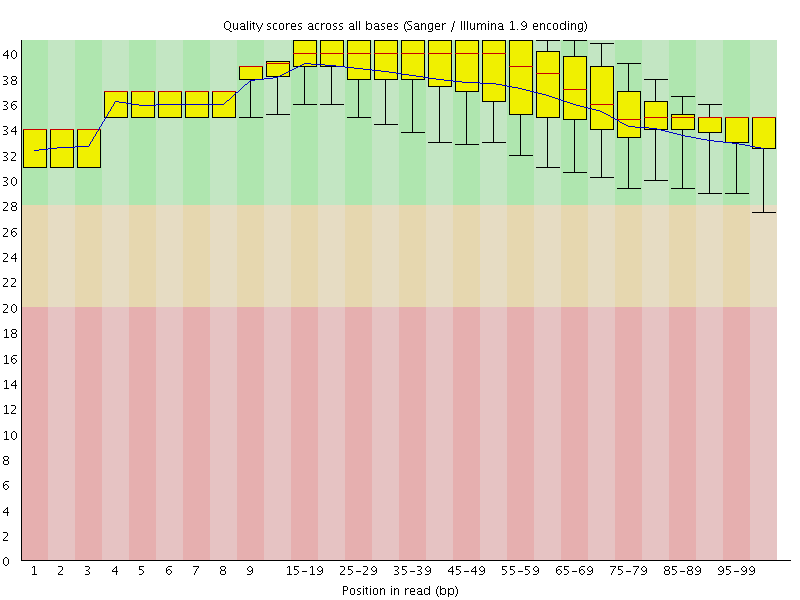
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
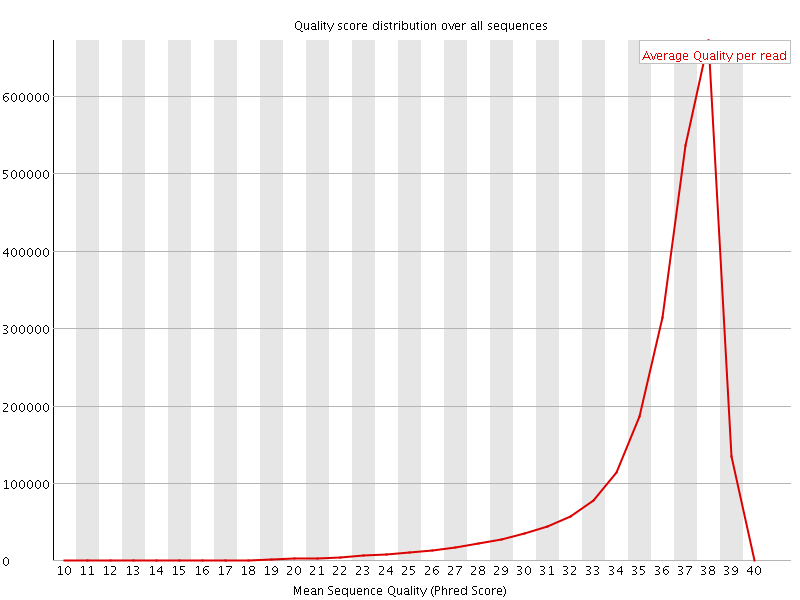
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
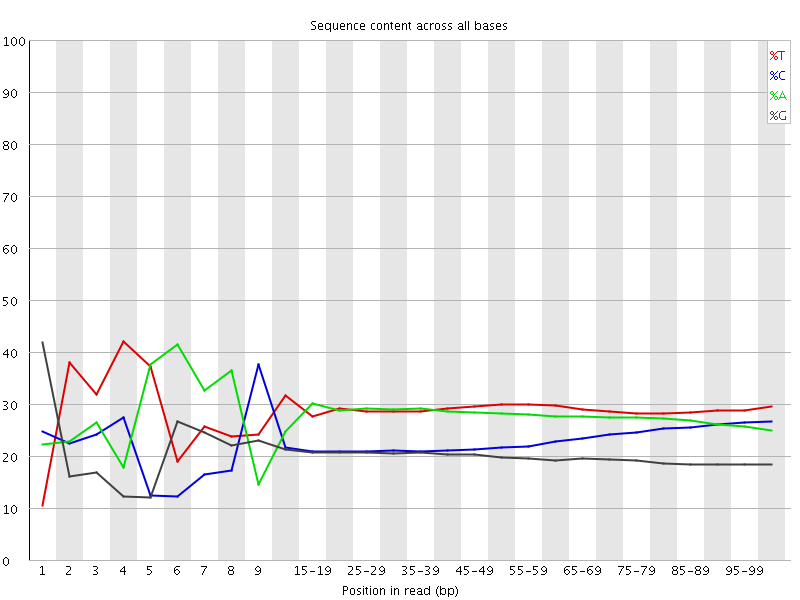
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
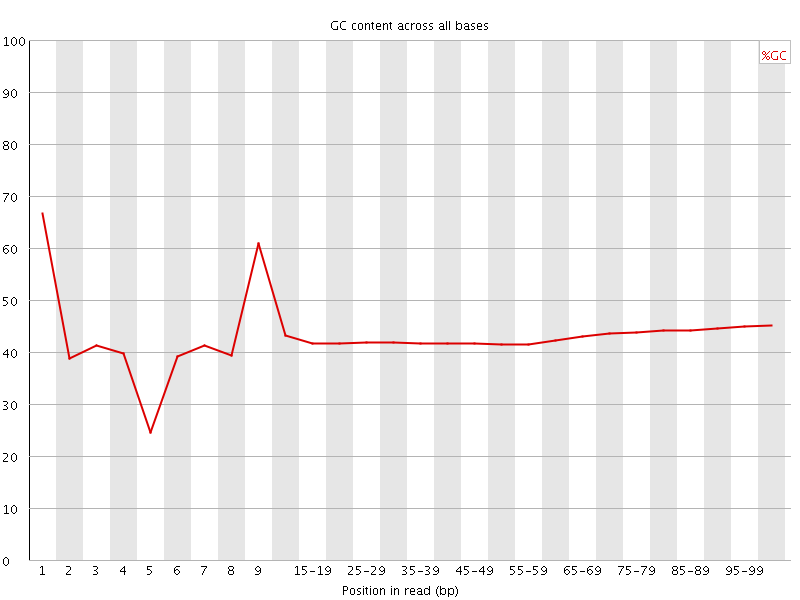
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
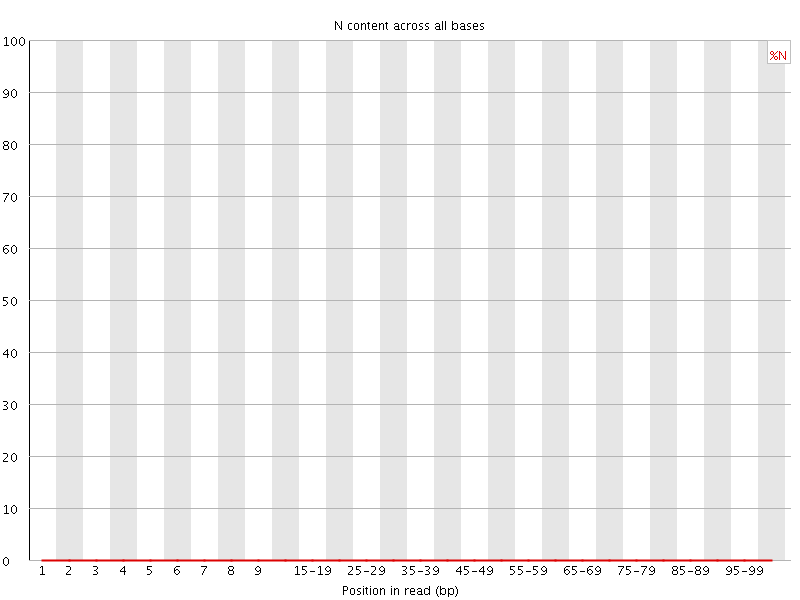
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
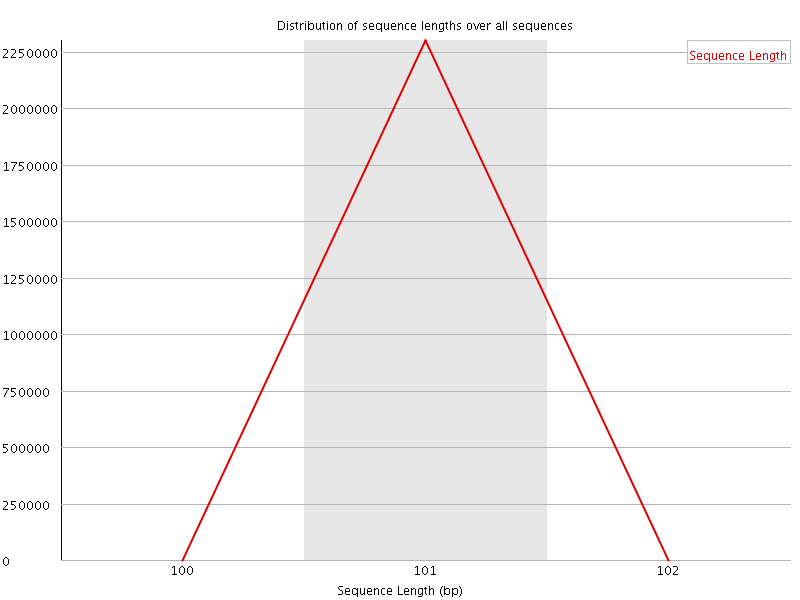
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
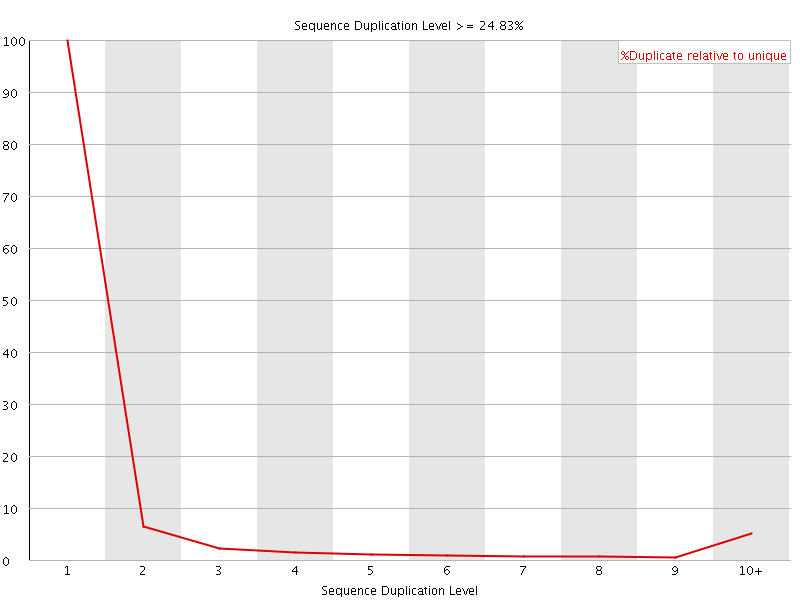
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
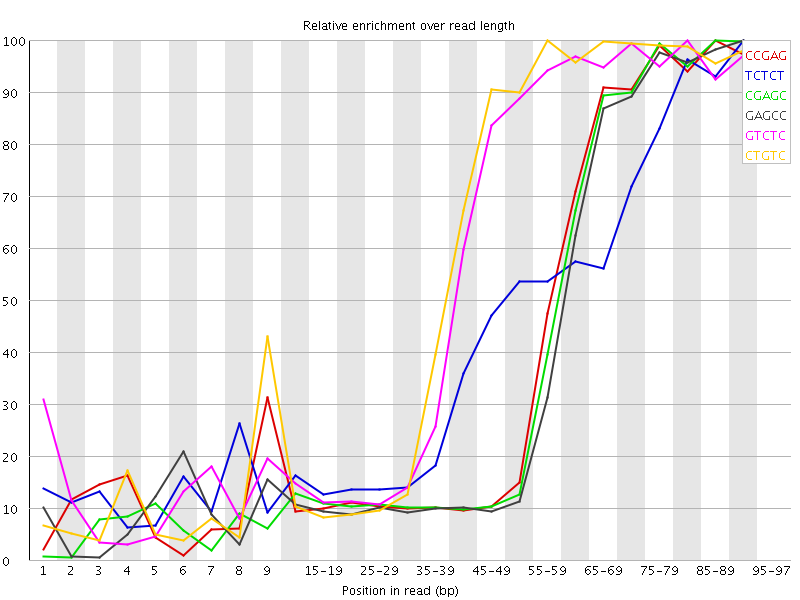
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 631380 | 4.7939653 | 10.736185 | 85-89 |
| TCTCT | 1375150 | 4.7575426 | 10.038269 | 90-94 |
| CGAGC | 616455 | 4.680642 | 10.697894 | 85-89 |
| GAGCC | 604845 | 4.5924892 | 10.792655 | 90-94 |
| GTCTC | 889605 | 4.464617 | 7.405394 | 80-84 |
| CTGTC | 874405 | 4.388333 | 7.098489 | 55-59 |
| CTCCG | 648805 | 4.1305094 | 9.000459 | 95-97 |
| TGTCT | 983095 | 3.889376 | 6.1696167 | 80-84 |
| ACATC | 1027535 | 3.8668265 | 10.183135 | 95-97 |
| AGCCC | 579385 | 3.8469756 | 9.432129 | 90-94 |
| GCCCA | 573525 | 3.8080666 | 9.49974 | 90-94 |
| CATCT | 1037065 | 3.7419784 | 9.8886 | 95-97 |
| ATCTC | 1019625 | 3.6790512 | 9.741636 | 95-97 |
| CTCTT | 1024335 | 3.5438442 | 5.5393634 | 60-64 |
| ACACA | 897820 | 3.5237908 | 6.1893063 | 70-74 |
| CCACG | 524065 | 3.4796643 | 9.344086 | 95-97 |
| TCCGA | 652595 | 3.415808 | 7.486877 | 85-89 |
| TCTCC | 766575 | 3.3642616 | 6.7067595 | 95-97 |
| CGAGA | 538520 | 3.3617556 | 8.846315 | 85-89 |
| GAGAC | 526095 | 3.2841916 | 8.992994 | 95-97 |
| ACGAG | 525895 | 3.2829428 | 8.867881 | 95-97 |
| TACAC | 850005 | 3.1987445 | 7.0411735 | 5 |
| CCCAC | 550560 | 3.196723 | 8.195584 | 95-97 |
| CACAT | 829320 | 3.1209023 | 5.7259603 | 95-97 |
| CACGA | 524365 | 2.862504 | 7.748577 | 95-97 |
| AGAAG | 527270 | 2.706186 | 5.181046 | 5 |
| AGACC | 485630 | 2.6510499 | 7.7099524 | 85-89 |
| GACCT | 495425 | 2.59315 | 7.418349 | 85-89 |
| TATAC | 870725 | 2.5830727 | 5.264399 | 5 |
| CTCTC | 543210 | 2.3839815 | 6.559113 | 90-94 |
| CCTCT | 511870 | 2.2464397 | 6.379504 | 90-94 |
| ACCTC | 476040 | 2.1789198 | 6.4004736 | 90-94 |
| CAAAG | 460465 | 2.0666611 | 5.048621 | 4 |
| CTTTG | 452780 | 1.7913141 | 7.8934426 | 9 |
| AGAGA | 348190 | 1.7870669 | 5.098925 | 8 |
| TCTCG | 341010 | 1.7114099 | 6.4522533 | 90-94 |
| CTACA | 442095 | 1.6636947 | 5.4054184 | 95-97 |
| CTCTA | 453365 | 1.6358495 | 5.06341 | 90-94 |
| GAGAG | 228750 | 1.6329666 | 5.347574 | 7 |
| GTATG | 335170 | 1.581482 | 6.223842 | 95-97 |
| TGCCG | 211120 | 1.5369887 | 9.043942 | 95-97 |
| CTCGT | 300305 | 1.5071259 | 6.340311 | 90-94 |
| TCTAC | 416050 | 1.501208 | 5.088349 | 95-97 |
| TTGAG | 310400 | 1.4646058 | 5.261857 | 9 |
| GCCGT | 196570 | 1.4310623 | 8.432071 | 95-97 |
| GTCTT | 341165 | 1.3497365 | 5.726399 | 1 |
| ATGCC | 252205 | 1.3200896 | 6.6892514 | 95-97 |
| TATGA | 381470 | 1.2941 | 6.1582236 | 4 |
| AAGAC | 286850 | 1.2874416 | 5.821148 | 5 |
| ATGAG | 252450 | 1.2423307 | 5.1275554 | 7 |
| GAGTC | 201570 | 1.2065004 | 6.094429 | 9 |
| CCGTC | 189320 | 1.2052745 | 6.7778697 | 95-97 |
| GTGTT | 262440 | 1.1873162 | 5.903408 | 1 |
| GATTG | 251135 | 1.1849673 | 5.95505 | 7 |
| TGAGT | 250905 | 1.1838819 | 5.152045 | 8 |
| TCATA | 381580 | 1.1319864 | 5.5842376 | 2 |
| TCGTA | 269580 | 1.112333 | 5.0931454 | 95-97 |
| GACTT | 267660 | 1.104411 | 5.755715 | 7 |
| TATGC | 262870 | 1.0846466 | 5.2418656 | 95-97 |
| GTGTA | 224990 | 1.0616033 | 9.929824 | 1 |
| CGTAT | 247255 | 1.0202165 | 5.1371617 | 95-97 |
| TGGAC | 169900 | 1.016939 | 6.9244323 | 5 |
| GGACT | 151850 | 0.90890056 | 6.3265977 | 6 |
| GTCCA | 170155 | 0.8906241 | 6.40861 | 1 |
| GTATA | 217545 | 0.7380004 | 6.8908153 | 1 |