![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-147_CTCTCTAC-TATCCTCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3435235 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
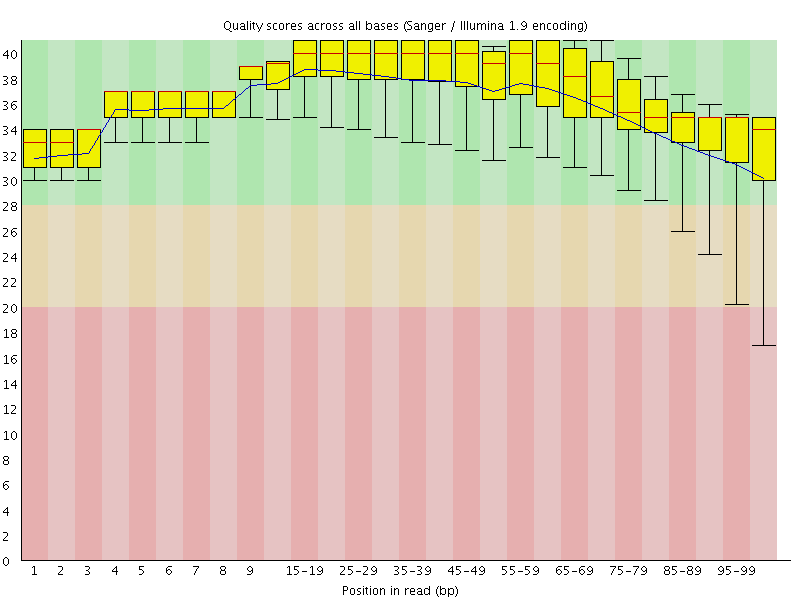
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
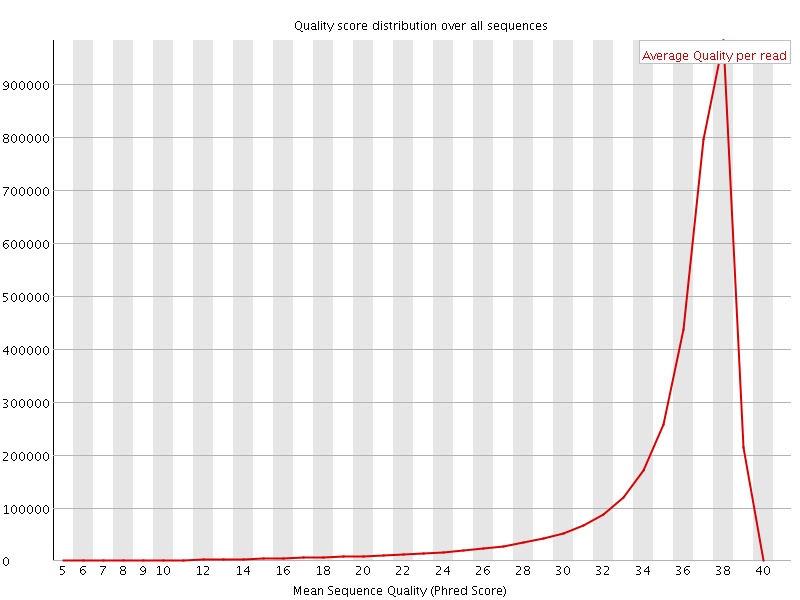
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
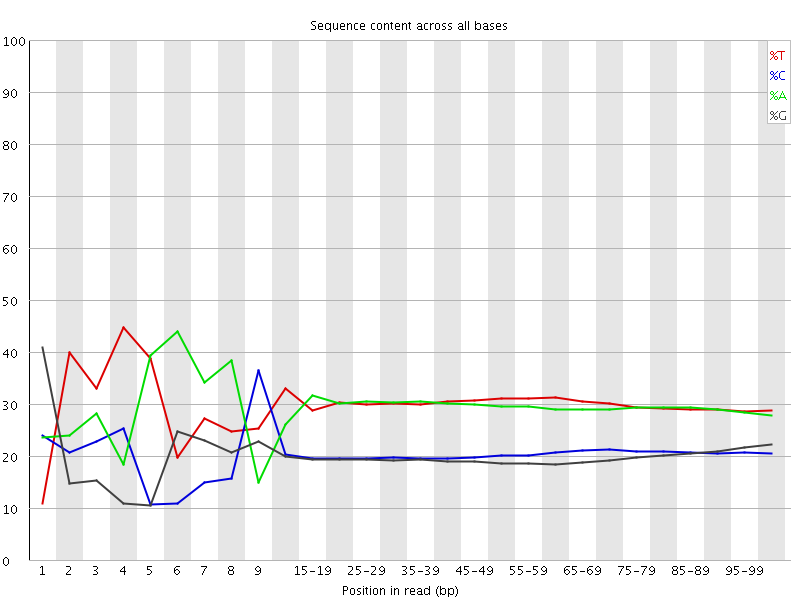
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
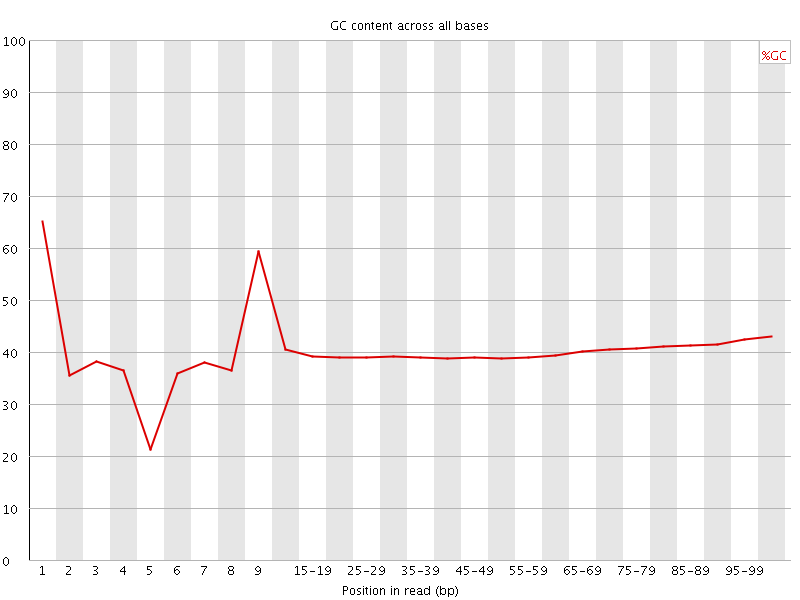
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
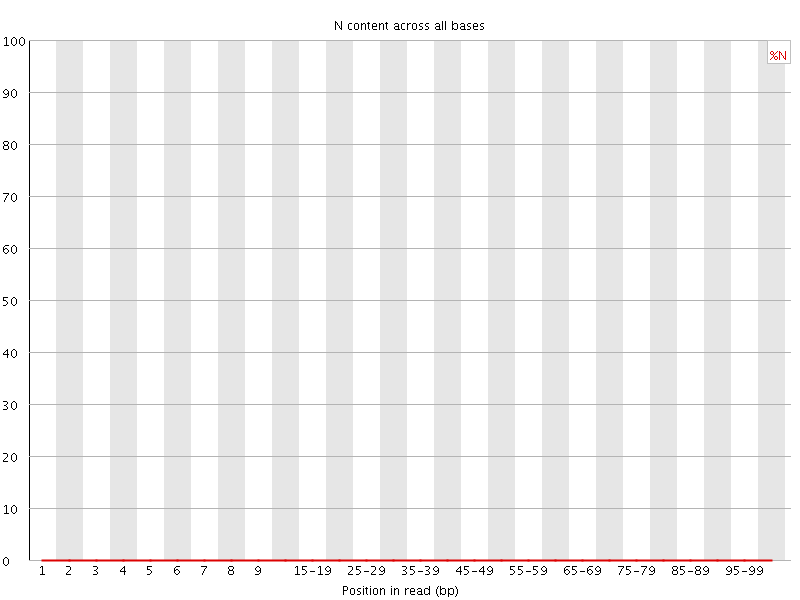
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
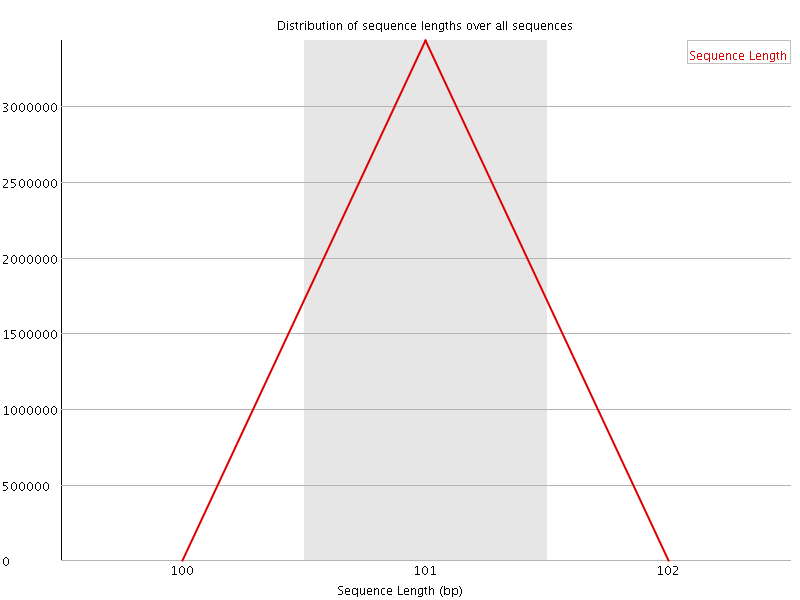
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
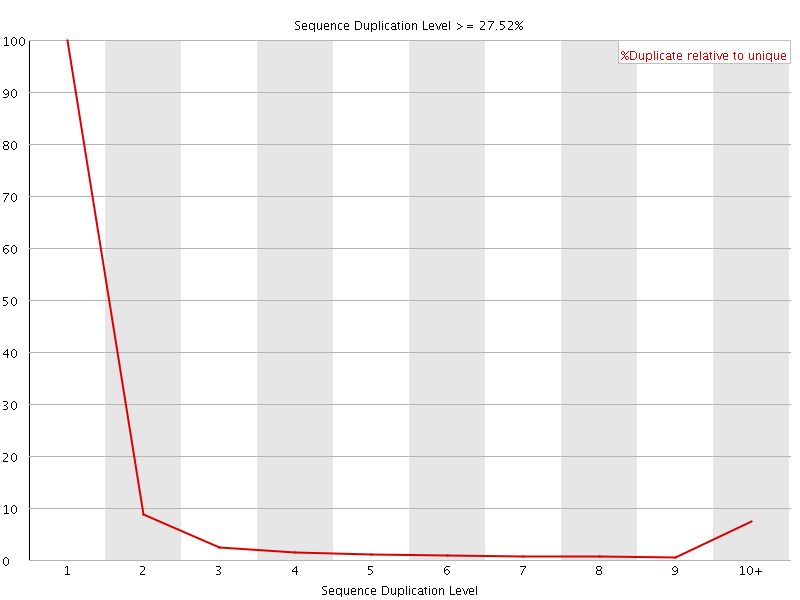
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
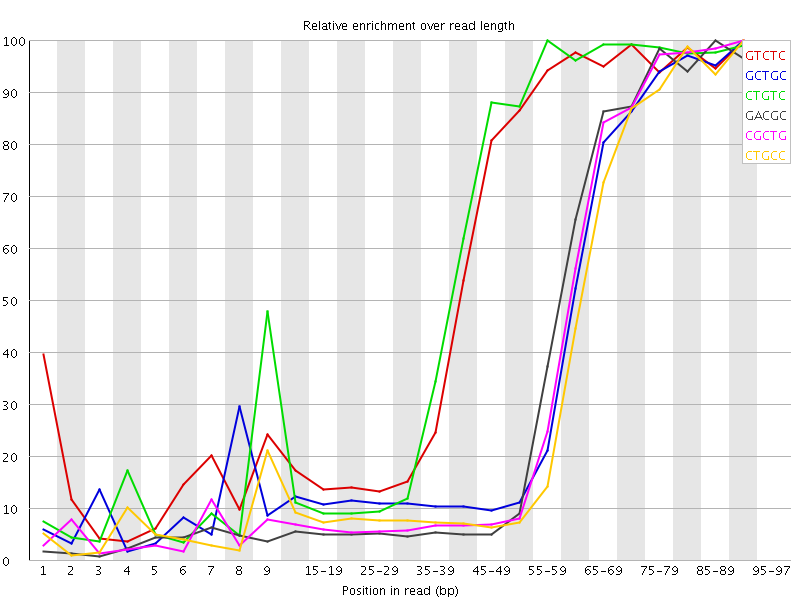
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 1150950 | 4.6003194 | 7.5795255 | 90-94 |
| GCTGC | 743405 | 4.5339994 | 11.015376 | 90-94 |
| CTGTC | 1096125 | 4.3811855 | 7.1539855 | 55-59 |
| GACGC | 698150 | 4.3470716 | 10.78313 | 85-89 |
| CGCTG | 691995 | 4.2204514 | 10.610729 | 90-94 |
| CTGCC | 674820 | 3.9997957 | 10.497612 | 90-94 |
| GCCGA | 637835 | 3.9715166 | 11.105869 | 95-97 |
| TCTCT | 1513005 | 3.9632142 | 5.9464397 | 60-64 |
| CGACG | 621725 | 3.8712068 | 11.196472 | 95-97 |
| TGCCG | 623560 | 3.8030694 | 10.517259 | 90-94 |
| CCGAC | 612950 | 3.7090864 | 10.59548 | 95-97 |
| CTCTT | 1409050 | 3.690911 | 5.6372266 | 60-64 |
| GAAGA | 1202775 | 3.549605 | 6.911401 | 85-89 |
| TGTCT | 1298360 | 3.4995196 | 5.4764457 | 90-94 |
| ACACA | 1192700 | 3.3244097 | 6.5502453 | 6 |
| CTGAC | 793660 | 3.238606 | 7.376571 | 95-97 |
| TGACG | 761570 | 3.197714 | 7.542811 | 95-97 |
| GAGGA | 718570 | 3.1695454 | 8.386717 | 90-94 |
| TACAC | 1105310 | 3.0176952 | 7.790146 | 5 |
| CACAT | 1090720 | 2.9778616 | 5.456261 | 95-97 |
| ATCTG | 1080280 | 2.9726357 | 5.67906 | 95-97 |
| CATCT | 1101360 | 2.945293 | 5.372539 | 80-84 |
| AGAGG | 664535 | 2.9312024 | 8.237825 | 90-94 |
| GACGA | 679625 | 2.9133399 | 7.897442 | 95-97 |
| ACGCT | 707690 | 2.8877974 | 7.065768 | 75-79 |
| ACATC | 1051150 | 2.8698285 | 5.3160477 | 90-94 |
| TCTGA | 1017255 | 2.7992082 | 5.3186965 | 95-97 |
| CGAAG | 632470 | 2.7112012 | 7.6972647 | 85-89 |
| AAGAG | 910555 | 2.6872113 | 5.949895 | 90-94 |
| TATAC | 1159305 | 2.1343749 | 5.630131 | 5 |
| ACGAA | 712625 | 2.043857 | 5.302136 | 85-89 |
| AGGAT | 679220 | 1.9634234 | 5.3622355 | 90-94 |
| CTTTG | 690280 | 1.8605381 | 7.8008814 | 9 |
| AGAGA | 616060 | 1.8181036 | 5.143748 | 8 |
| GGTGG | 275215 | 1.7772166 | 6.5618124 | 95-97 |
| GAGAG | 392645 | 1.7319206 | 5.9509196 | 7 |
| GGATA | 591080 | 1.7086368 | 5.0905113 | 95-97 |
| GCTTT | 578765 | 1.5599675 | 5.044419 | 1 |
| CTCGG | 243330 | 1.4840606 | 6.4902906 | 95-97 |
| GTGTA | 522890 | 1.4805458 | 9.046255 | 1 |
| AAGAC | 441160 | 1.2652768 | 5.609695 | 5 |
| GAGTC | 295985 | 1.2427949 | 6.2411094 | 9 |
| CCATG | 299680 | 1.2228732 | 5.0600634 | 9 |
| CGGTG | 192615 | 1.2087938 | 5.656624 | 95-97 |
| GTCTT | 439060 | 1.1834153 | 6.2996397 | 1 |
| GTGTT | 409535 | 1.1358225 | 5.665527 | 1 |
| TATGA | 593645 | 1.1246204 | 5.3990374 | 4 |
| GACTT | 405610 | 1.116128 | 5.602048 | 7 |
| GATTG | 393490 | 1.1141537 | 5.189044 | 7 |
| GTTCT | 409885 | 1.1047788 | 5.100642 | 1 |
| TCATA | 595245 | 1.0958945 | 5.39609 | 2 |
| TGGAC | 238655 | 1.0020752 | 6.4527407 | 5 |
| GTCCA | 235710 | 0.96183735 | 6.88476 | 1 |
| AGACT | 332005 | 0.93270016 | 5.066574 | 6 |
| GGACT | 217660 | 0.9139204 | 5.835751 | 6 |
| GTATA | 364440 | 0.69040704 | 6.8897305 | 1 |