![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-147_CTCTCTAC-TATCCTCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3435235 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
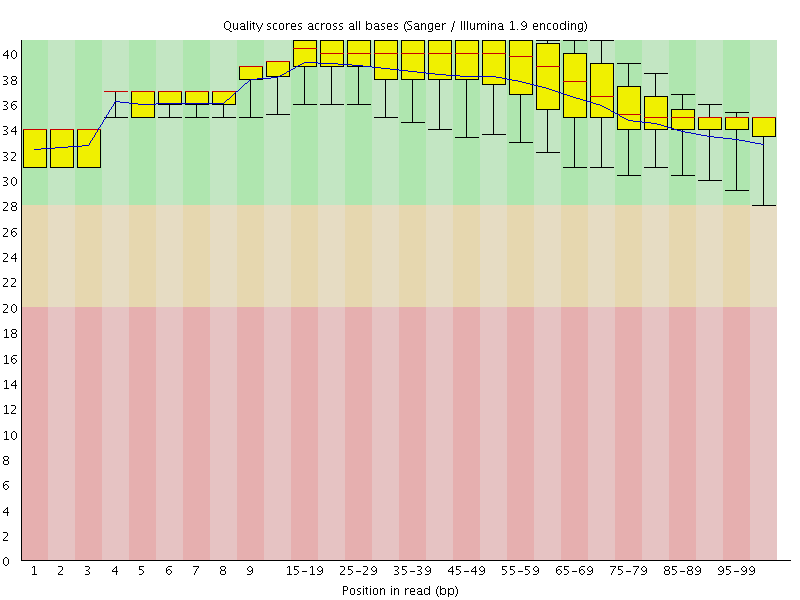
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
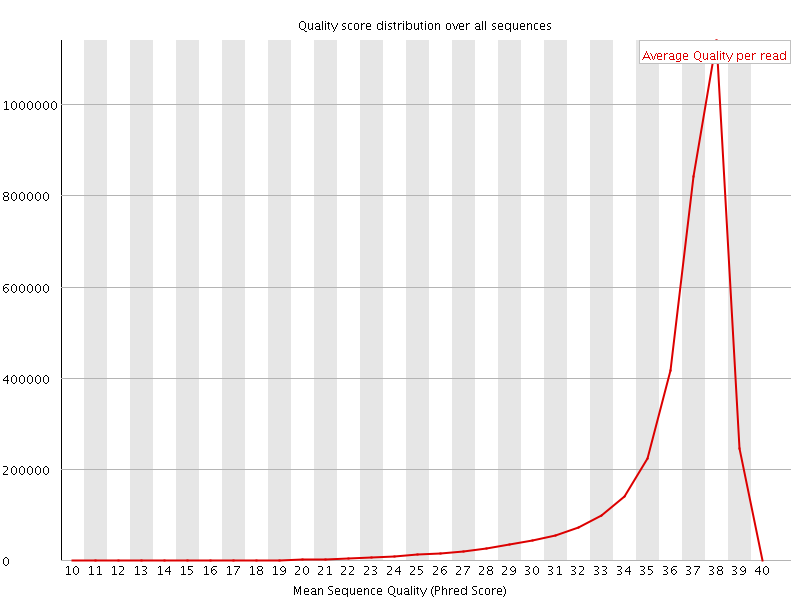
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
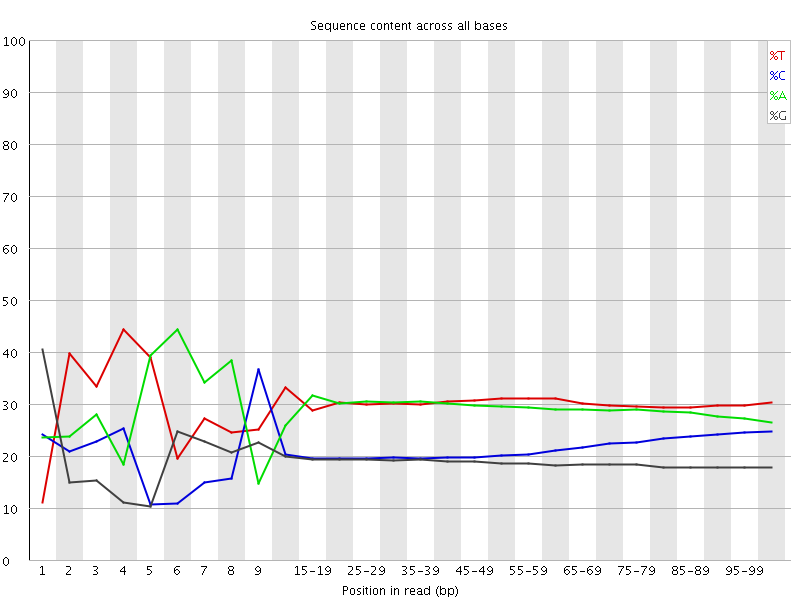
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
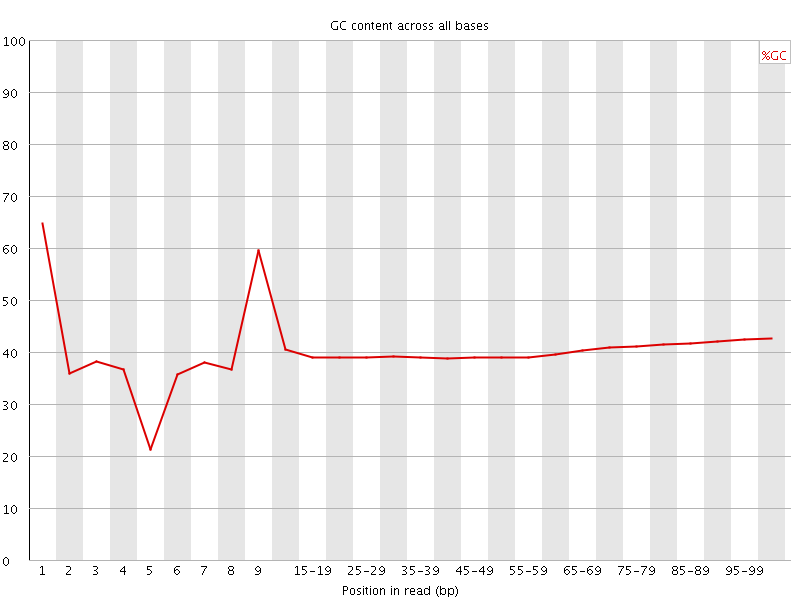
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
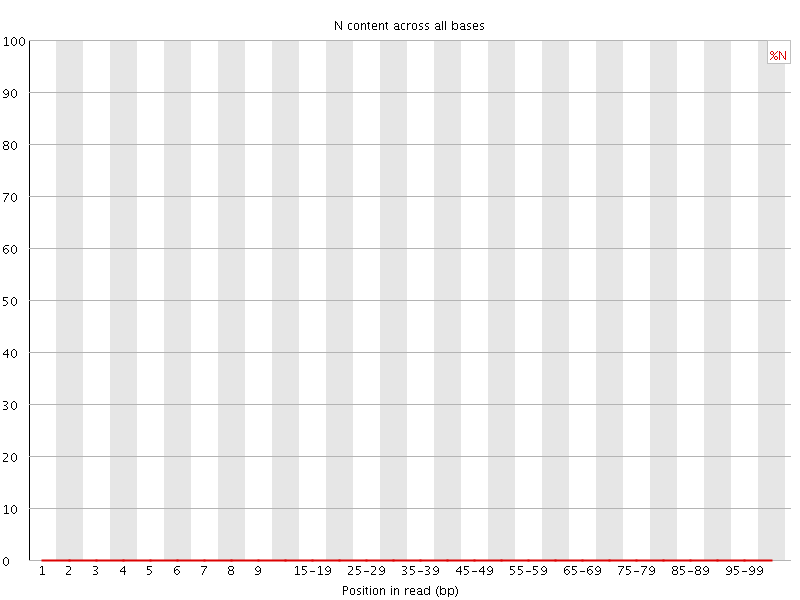
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
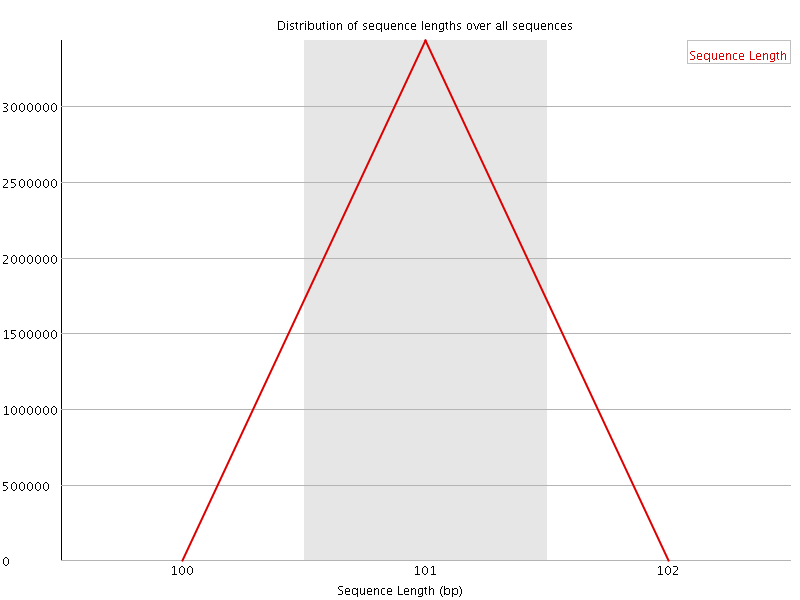
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
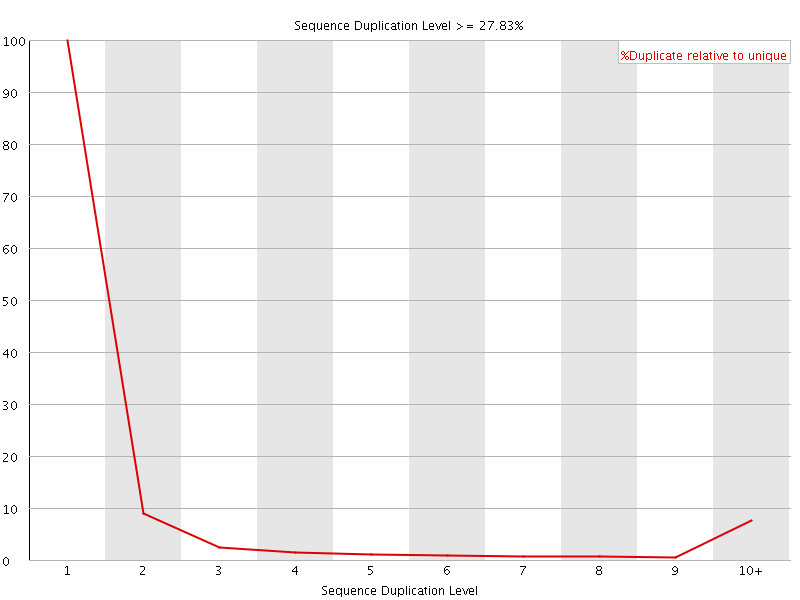
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
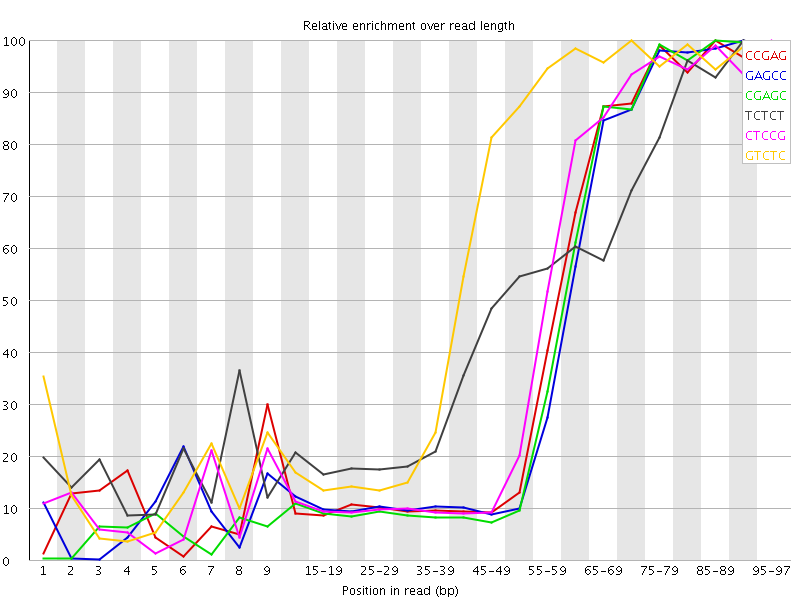
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 775305 | 4.8397007 | 11.179712 | 85-89 |
| GAGCC | 747110 | 4.663699 | 11.093031 | 90-94 |
| CGAGC | 731410 | 4.5656943 | 10.920015 | 85-89 |
| TCTCT | 1871210 | 4.4121237 | 8.93845 | 90-94 |
| CTCCG | 809905 | 4.3416457 | 9.656052 | 95-97 |
| GTCTC | 1149295 | 4.3376884 | 7.1208997 | 70-74 |
| CTGTC | 1098325 | 4.145317 | 6.753465 | 65-69 |
| AGCCC | 711215 | 3.9394777 | 9.87717 | 90-94 |
| GCCCA | 704245 | 3.9008703 | 9.915307 | 90-94 |
| CCACG | 633710 | 3.5101712 | 9.687335 | 95-97 |
| TGTCT | 1300905 | 3.4568381 | 5.374593 | 90-94 |
| TCTCC | 1018515 | 3.4110336 | 6.613295 | 95-97 |
| CCCAC | 680495 | 3.3446786 | 8.704384 | 90-94 |
| ACATC | 1322080 | 3.3282635 | 8.639547 | 95-97 |
| CTCTT | 1409360 | 3.3231282 | 5.092705 | 60-64 |
| CATCT | 1352510 | 3.29521 | 8.520378 | 95-97 |
| TCCGA | 844265 | 3.2924793 | 7.2026134 | 85-89 |
| ATCTC | 1337775 | 3.25931 | 8.376259 | 95-97 |
| GAGAC | 695920 | 3.1603072 | 8.436073 | 95-97 |
| CGAGA | 694145 | 3.1522465 | 8.348422 | 85-89 |
| ACACA | 1197460 | 3.1148593 | 6.255428 | 6 |
| ACGAG | 666930 | 3.028658 | 8.274604 | 95-97 |
| TACAC | 1111675 | 2.798581 | 7.577215 | 5 |
| CACAT | 1097245 | 2.762254 | 5.0090175 | 95-97 |
| AGAAG | 821535 | 2.7140636 | 5.8095093 | 5 |
| CACGA | 657015 | 2.6475062 | 7.202392 | 95-97 |
| CTCTC | 767880 | 2.5716505 | 6.5629964 | 90-94 |
| AGACC | 624410 | 2.5161214 | 7.25239 | 85-89 |
| GACCT | 620960 | 2.4216309 | 6.939699 | 90-94 |
| CCTCT | 683940 | 2.290533 | 6.3453994 | 90-94 |
| ACCTC | 612895 | 2.1209092 | 6.18642 | 90-94 |
| TATAC | 1164900 | 2.0646918 | 5.622655 | 5 |
| CAAAG | 699165 | 2.0495822 | 5.125182 | 4 |
| AGAGA | 610055 | 2.0154078 | 5.7758727 | 8 |
| GAGAG | 388025 | 1.9858111 | 6.714349 | 7 |
| CTTTG | 686755 | 1.824884 | 7.735209 | 9 |
| TGGAG | 357915 | 1.7727222 | 5.0933013 | 5 |
| AAGAG | 519110 | 1.7149575 | 5.2713246 | 5 |
| TCTCG | 442490 | 1.6700532 | 5.9962616 | 90-94 |
| GCTTT | 575665 | 1.5296893 | 5.0651746 | 1 |
| AGGAG | 294205 | 1.5056647 | 5.1337724 | 7 |
| TTGAG | 481345 | 1.4894146 | 5.0437636 | 9 |
| CTCGT | 385455 | 1.4547908 | 5.8524213 | 90-94 |
| GTATG | 462115 | 1.4299116 | 5.135044 | 95-97 |
| GTCTT | 500505 | 1.32997 | 5.8666344 | 1 |
| GAGTC | 298545 | 1.3120861 | 6.35629 | 9 |
| AAGAC | 441980 | 1.2956519 | 6.019176 | 5 |
| GCCGT | 206475 | 1.247372 | 8.423402 | 95-97 |
| TGCCG | 204205 | 1.2336583 | 8.907696 | 95-97 |
| GTGTT | 408140 | 1.222224 | 6.197607 | 1 |
| GATTG | 393190 | 1.2166386 | 5.7233667 | 7 |
| ATGCC | 305135 | 1.1899709 | 6.0357003 | 95-97 |
| CCATG | 302670 | 1.1803577 | 5.0143623 | 9 |
| TATGA | 590500 | 1.1794913 | 5.5762773 | 4 |
| CCGTC | 209875 | 1.1250737 | 6.8723006 | 95-97 |
| GACTT | 407955 | 1.1201162 | 5.674977 | 7 |
| GTTCT | 413540 | 1.0988818 | 5.124447 | 1 |
| TCATA | 599190 | 1.0620162 | 5.290822 | 2 |
| GTGTA | 342005 | 1.058258 | 9.532261 | 1 |
| TGGAC | 234995 | 1.0327879 | 6.562981 | 5 |
| GAGTA | 297815 | 0.9521889 | 5.423184 | 1 |
| GGACT | 215700 | 0.9479877 | 6.241225 | 6 |
| AGACT | 332650 | 0.9437476 | 5.3796506 | 6 |
| GTCCA | 239085 | 0.9323878 | 6.3066406 | 1 |
| GTATA | 361850 | 0.7227755 | 7.079308 | 1 |