![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-146_CTCTCTAC-CTCTCTAT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4089092 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
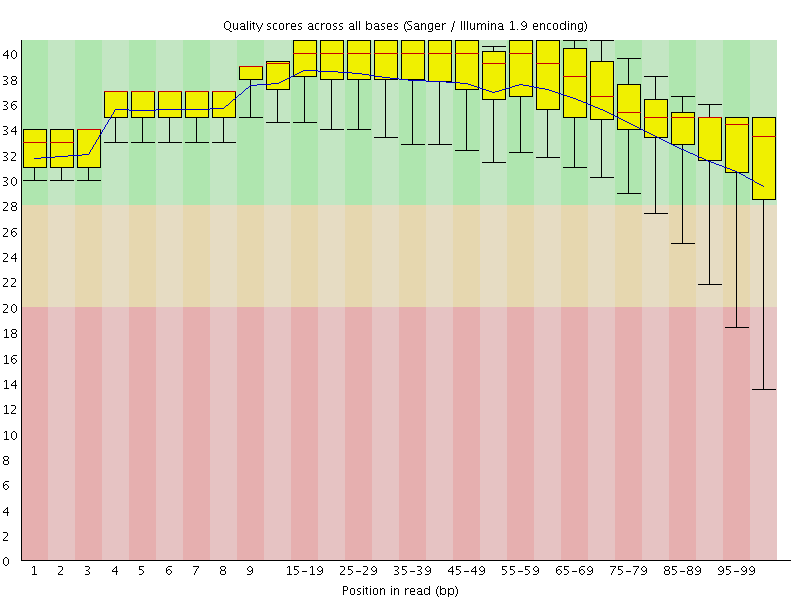
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
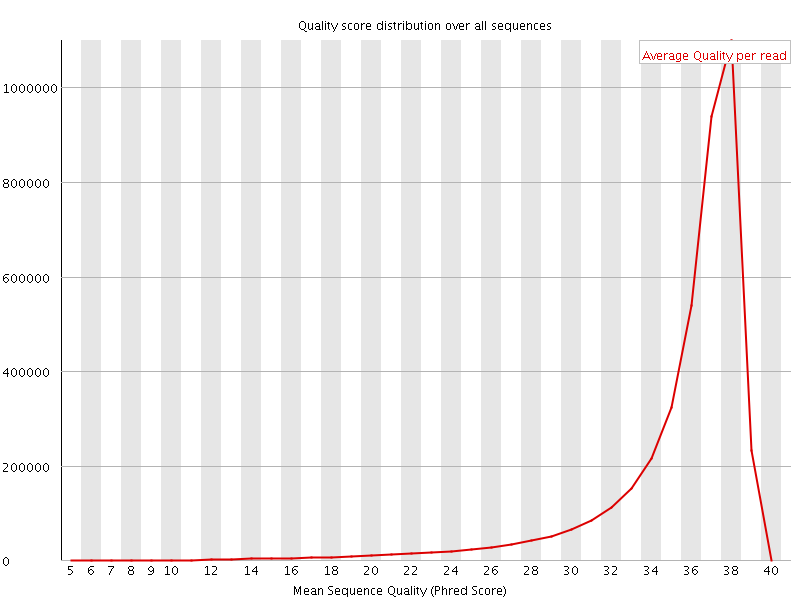
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
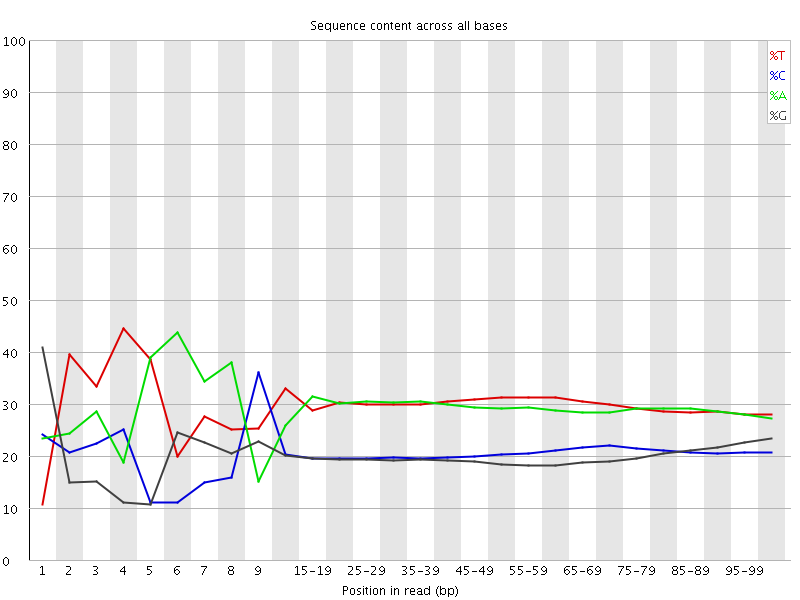
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
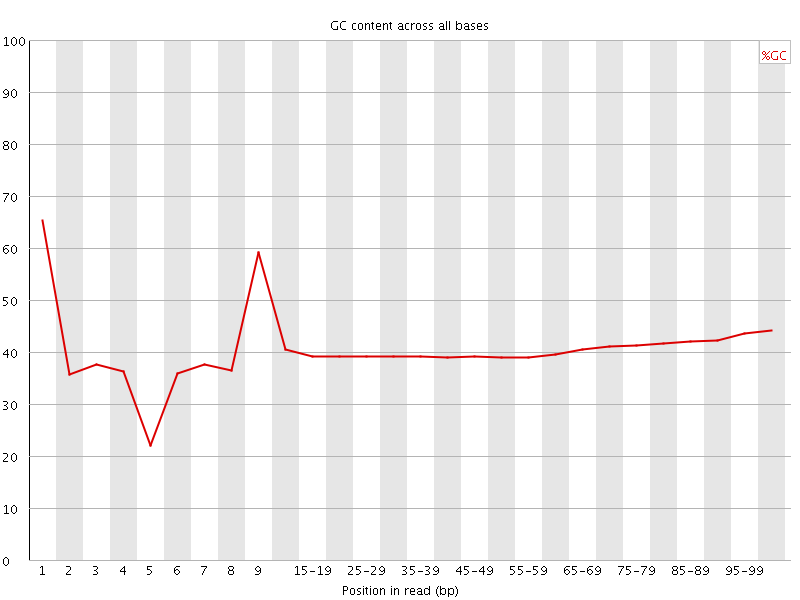
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
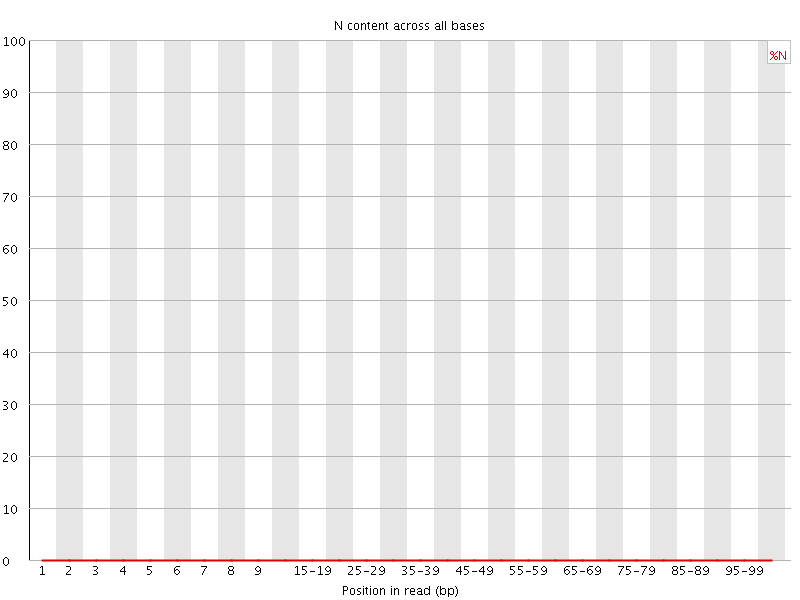
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
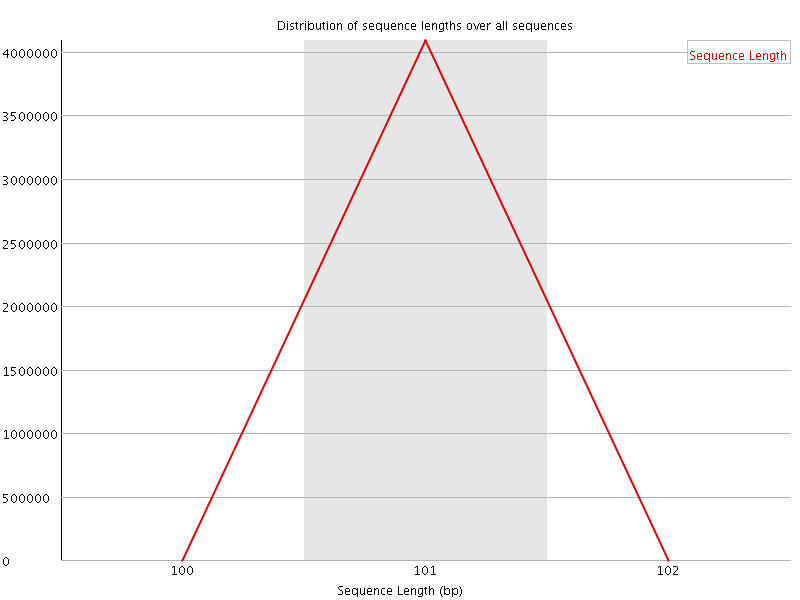
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
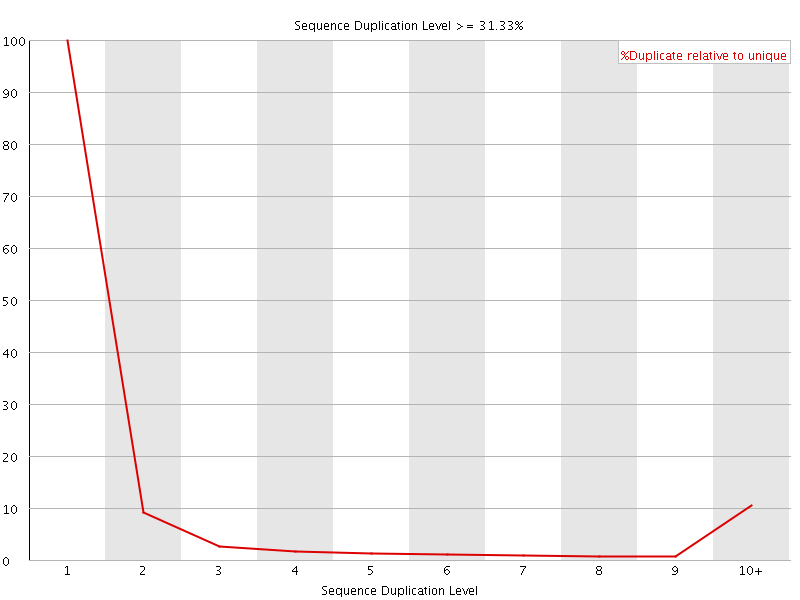
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
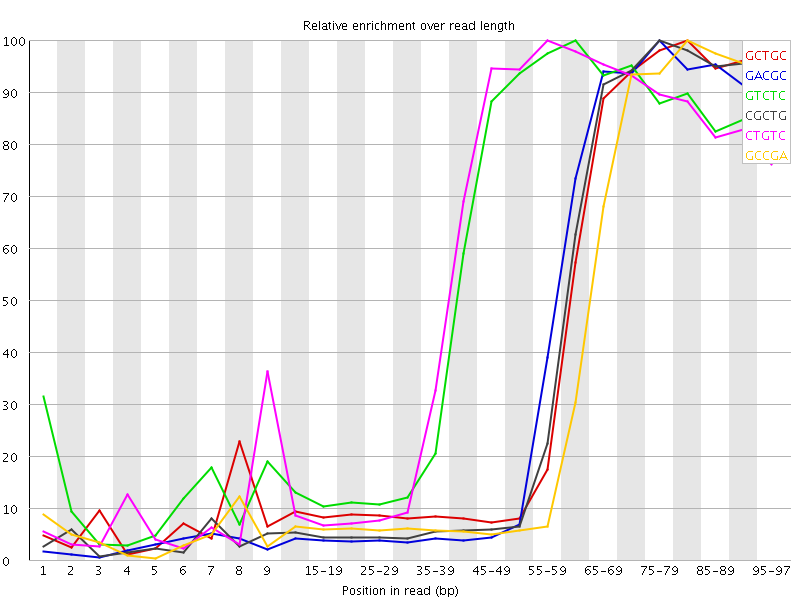
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 1112105 | 5.5375786 | 13.618951 | 80-84 |
| GACGC | 1077430 | 5.494847 | 13.6620865 | 75-79 |
| GTCTC | 1653385 | 5.4443197 | 9.426706 | 60-64 |
| CGCTG | 1069460 | 5.325234 | 13.460275 | 75-79 |
| CTGTC | 1590475 | 5.2371674 | 9.056999 | 55-59 |
| GCCGA | 981570 | 5.0059648 | 13.756308 | 80-84 |
| CTGCC | 1033180 | 4.9694304 | 13.175347 | 80-84 |
| CGACG | 953740 | 4.864033 | 13.775907 | 85-89 |
| TGCCG | 969160 | 4.8258038 | 13.383694 | 80-84 |
| CCGAC | 945365 | 4.657174 | 13.179946 | 80-84 |
| TCTCT | 2063790 | 4.4939747 | 7.29879 | 60-64 |
| CTCTT | 1942125 | 4.229045 | 6.936765 | 60-64 |
| TGTCT | 1824350 | 4.1126037 | 6.712984 | 55-59 |
| TGACG | 1156340 | 4.037301 | 9.573247 | 75-79 |
| CTGAC | 1196730 | 4.036065 | 9.175344 | 75-79 |
| ACACA | 1668730 | 3.9041662 | 7.1770477 | 70-74 |
| GAGAG | 997670 | 3.6934175 | 10.701762 | 95-97 |
| ACGCT | 1085385 | 3.6605458 | 9.083091 | 75-79 |
| GACGA | 1015620 | 3.6318607 | 10.1107 | 85-89 |
| ATCTG | 1559950 | 3.6017346 | 7.145058 | 70-74 |
| TACAC | 1573950 | 3.5953472 | 6.6772323 | 5 |
| CACAT | 1547285 | 3.5344367 | 6.6104894 | 70-74 |
| CATCT | 1560230 | 3.4797351 | 6.7280746 | 70-74 |
| ACATC | 1498685 | 3.4234202 | 6.701116 | 70-74 |
| TCTGA | 1450040 | 3.347966 | 6.6874895 | 70-74 |
| AGAGA | 1280905 | 3.2117836 | 7.657441 | 90-94 |
| AGAGG | 854900 | 3.164877 | 10.524596 | 95-97 |
| TATAC | 1643985 | 2.5709095 | 5.346534 | 5 |
| ACGAA | 1044085 | 2.528842 | 6.869229 | 85-89 |
| GAGGT | 673800 | 2.4354548 | 8.520845 | 95-97 |
| CGAAT | 1019510 | 2.4109318 | 6.600143 | 85-89 |
| TAGAG | 943885 | 2.3107667 | 6.970577 | 90-94 |
| AGGTG | 632635 | 2.2866638 | 8.867383 | 95-97 |
| GGTGT | 600485 | 2.1191363 | 7.1822376 | 95-97 |
| GGTGG | 389855 | 2.0804842 | 10.43378 | 95-97 |
| CTTTG | 788705 | 1.7779653 | 7.983779 | 9 |
| CTCGG | 339090 | 1.6884537 | 8.934395 | 95-97 |
| GTGTA | 692405 | 1.6550258 | 7.9263186 | 1 |
| GCTTT | 658305 | 1.4840066 | 5.0280843 | 1 |
| TCTCG | 428075 | 1.4095792 | 5.76826 | 95-97 |
| CGGTG | 269805 | 1.3908094 | 8.439212 | 95-97 |
| GGTCG | 240605 | 1.2402875 | 5.0005207 | 95-97 |
| AAGAC | 509040 | 1.2329279 | 5.4809384 | 5 |
| GAGTC | 345720 | 1.2070633 | 6.5339956 | 9 |
| CGCCG | 161125 | 1.1719202 | 5.19173 | 95-97 |
| GTCTT | 502920 | 1.1337247 | 6.2189465 | 1 |
| GACTT | 469925 | 1.0849997 | 5.6365566 | 7 |
| GTTCT | 477285 | 1.0759361 | 5.002933 | 1 |
| GTGTT | 454270 | 1.0601475 | 5.6117206 | 1 |
| TCGGT | 301625 | 1.028207 | 5.3818517 | 95-97 |
| TGGAC | 277775 | 0.9698369 | 6.3139176 | 5 |
| GTCCA | 276435 | 0.93229866 | 6.8401732 | 1 |
| AGACT | 381565 | 0.9023228 | 5.0459075 | 6 |
| GGACT | 255765 | 0.8929902 | 6.1157045 | 6 |
| GTATA | 414630 | 0.6712638 | 6.7821918 | 1 |