![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-146_CTCTCTAC-CTCTCTAT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4089092 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
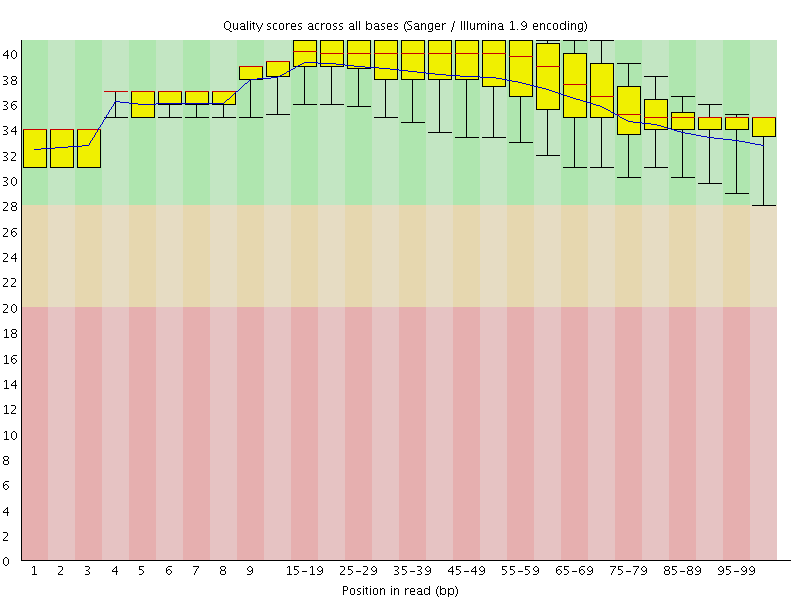
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
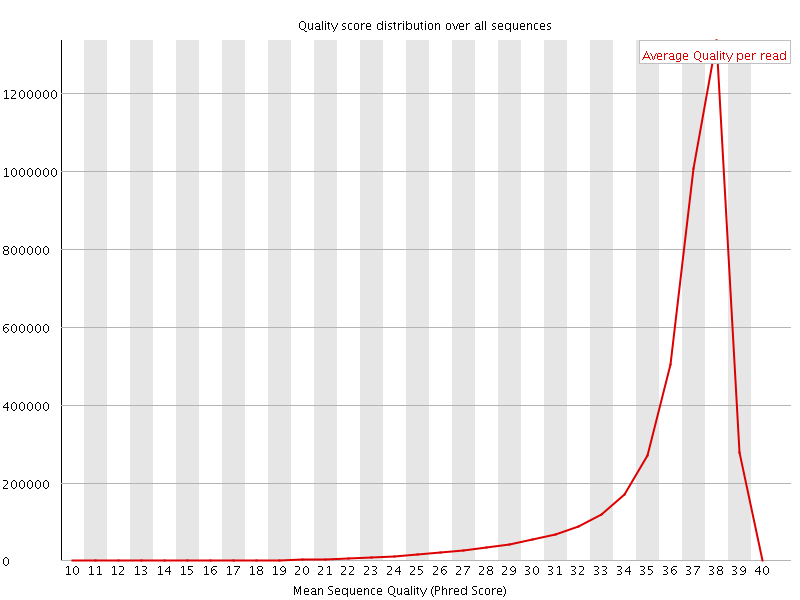
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
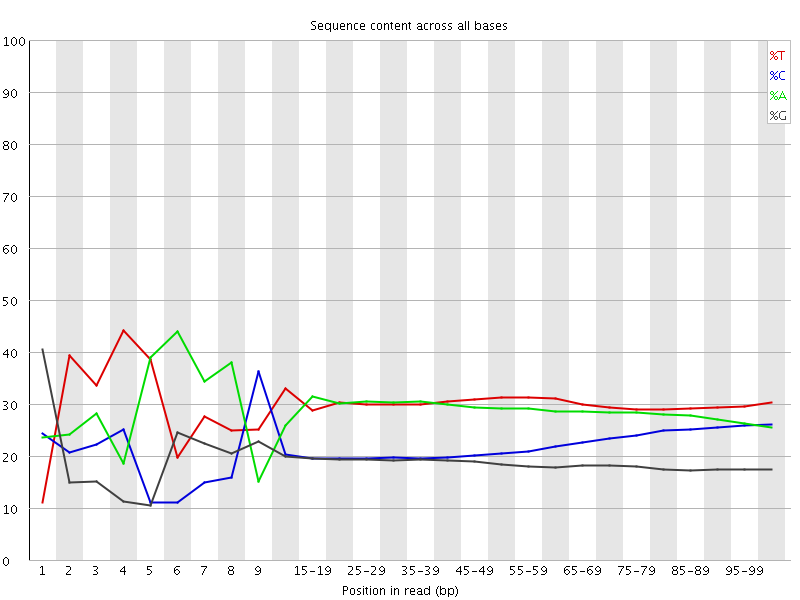
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
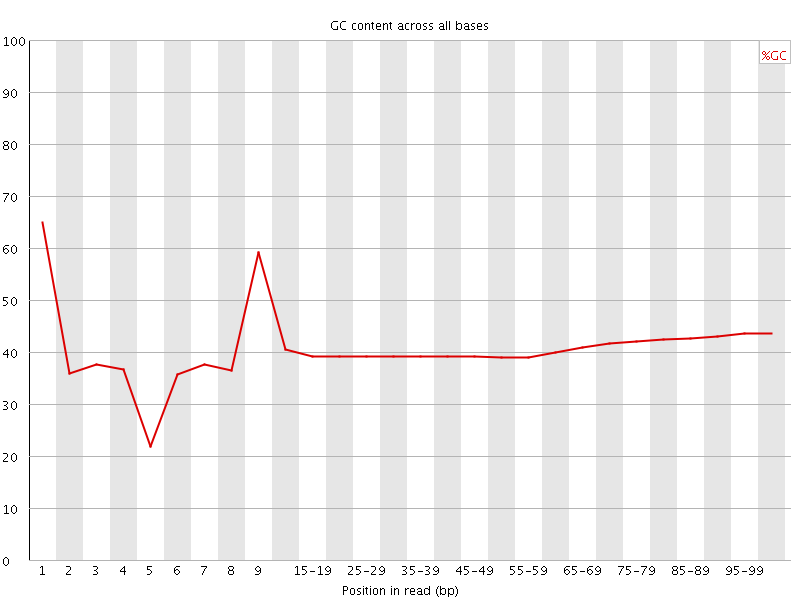
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
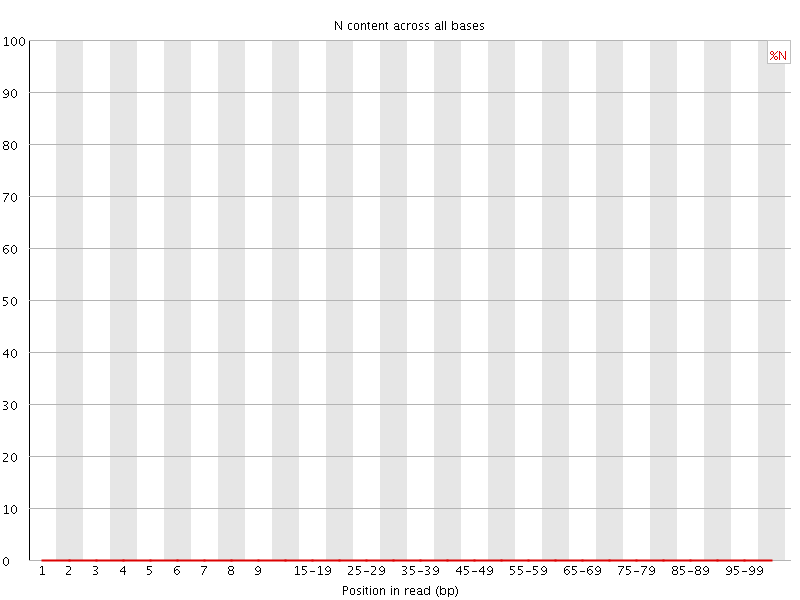
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
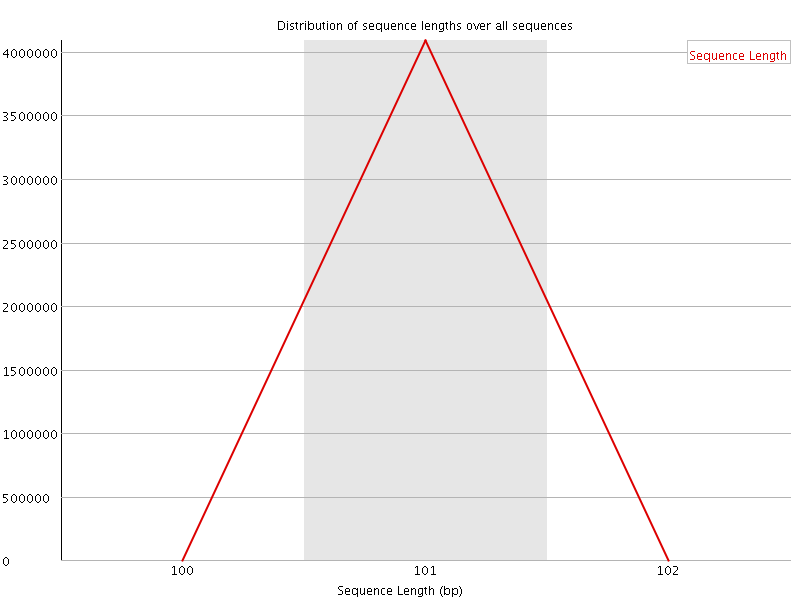
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
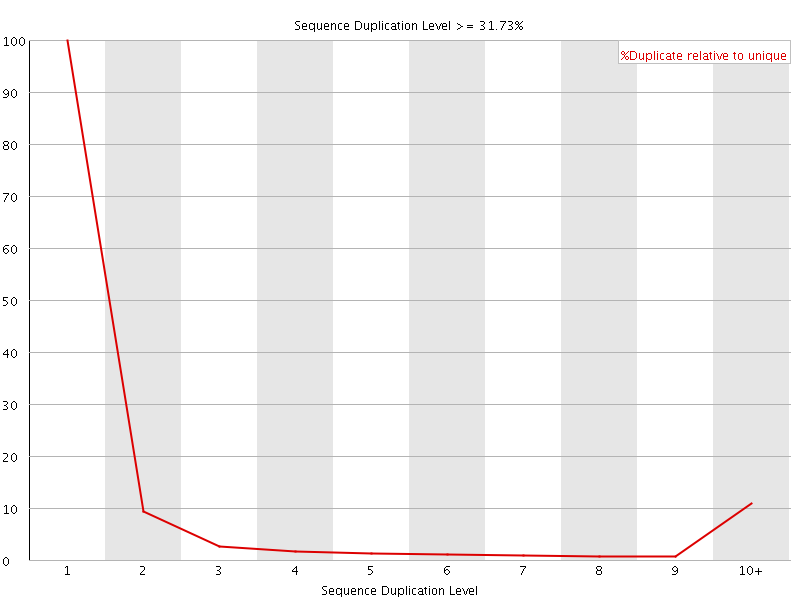
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
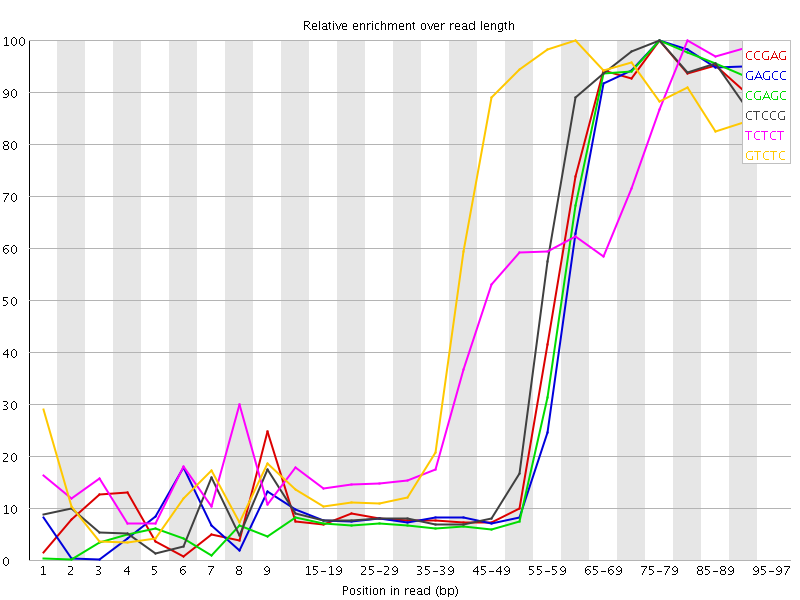
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 1173540 | 6.003707 | 14.160686 | 75-79 |
| GAGCC | 1133450 | 5.798611 | 14.039639 | 75-79 |
| CGAGC | 1114450 | 5.7014093 | 13.877911 | 75-79 |
| CTCCG | 1218135 | 5.1265936 | 11.492078 | 75-79 |
| TCTCT | 2667110 | 5.0538306 | 10.182226 | 80-84 |
| GTCTC | 1655810 | 5.0493584 | 8.701464 | 60-64 |
| CTGTC | 1596435 | 4.868295 | 8.442168 | 55-59 |
| AGCCC | 1086505 | 4.7666717 | 11.915202 | 80-84 |
| GCCCA | 1066445 | 4.6786647 | 12.050489 | 80-84 |
| CCACG | 974745 | 4.2763624 | 11.790124 | 80-84 |
| TGTCT | 1830420 | 4.044535 | 6.612986 | 55-59 |
| ACATC | 1955330 | 4.0262423 | 10.431021 | 95-97 |
| TCCGA | 1258755 | 4.0014453 | 8.957243 | 75-79 |
| CGAGA | 1032110 | 3.9883182 | 10.922419 | 85-89 |
| GAGAC | 1027070 | 3.9688425 | 10.967763 | 85-89 |
| CATCT | 1989970 | 3.9307585 | 10.191862 | 95-97 |
| CCCAC | 1033500 | 3.8882654 | 10.298203 | 80-84 |
| ATCTC | 1968220 | 3.8877962 | 10.142058 | 95-97 |
| ACGAG | 1004860 | 3.8830175 | 10.612501 | 85-89 |
| TCTCC | 1458205 | 3.8133454 | 7.646809 | 70-74 |
| CTCTT | 1948880 | 3.692877 | 6.073147 | 60-64 |
| ACACA | 1676065 | 3.5976648 | 6.606293 | 70-74 |
| CACGA | 994220 | 3.294642 | 9.031515 | 80-84 |
| TACAC | 1580840 | 3.2551258 | 6.204765 | 5 |
| CACAT | 1553405 | 3.1986341 | 6.000886 | 70-74 |
| AGACC | 938860 | 3.11119 | 9.17086 | 85-89 |
| GACCT | 935180 | 2.9728355 | 8.879519 | 85-89 |
| CTCTC | 1071395 | 2.8018005 | 7.7717133 | 90-94 |
| AGAAG | 949665 | 2.7718928 | 5.8220325 | 5 |
| CCTCT | 988185 | 2.5841985 | 7.595712 | 90-94 |
| ACCTC | 907640 | 2.474294 | 7.5405393 | 90-94 |
| TATAC | 1652605 | 2.4657052 | 5.320581 | 5 |
| AGAGA | 698970 | 2.0401614 | 6.12488 | 8 |
| GAGAG | 442675 | 1.9947426 | 6.997825 | 7 |
| CAAAG | 794105 | 1.9876772 | 5.0254793 | 4 |
| TCTCG | 642835 | 1.9603118 | 7.9784966 | 90-94 |
| TGGAG | 413360 | 1.786818 | 5.2647266 | 5 |
| CTCTA | 903040 | 1.7837615 | 5.617911 | 95-97 |
| CTACA | 854680 | 1.7598814 | 5.7741194 | 95-97 |
| CTTTG | 789340 | 1.7441425 | 7.899967 | 9 |
| AAGAG | 594965 | 1.7365905 | 5.4286137 | 5 |
| CTCGT | 562815 | 1.7162926 | 7.9048605 | 90-94 |
| GTATG | 616985 | 1.6572217 | 7.235956 | 95-97 |
| TCTAC | 824385 | 1.6283956 | 5.5317125 | 95-97 |
| AGGAG | 346235 | 1.5601733 | 5.197573 | 7 |
| TGCCG | 302600 | 1.485048 | 12.276223 | 95-97 |
| GCCGT | 298935 | 1.4670615 | 11.7654085 | 95-97 |
| ATGCC | 429450 | 1.365175 | 8.137341 | 95-97 |
| GTCTT | 606895 | 1.3410083 | 5.77846 | 1 |
| GAGTC | 347860 | 1.2894909 | 6.655364 | 9 |
| AAGAC | 507510 | 1.2703184 | 5.680819 | 5 |
| ATGAG | 448885 | 1.256872 | 5.1641736 | 7 |
| CCGTC | 297750 | 1.2530986 | 9.213378 | 95-97 |
| GATTG | 454745 | 1.2214451 | 5.3224916 | 7 |
| TCGTA | 520255 | 1.1983513 | 5.874641 | 95-97 |
| TGAGT | 442405 | 1.1882998 | 5.355049 | 8 |
| GTGTT | 454805 | 1.1718748 | 5.9786305 | 1 |
| TATGC | 502260 | 1.1569017 | 5.952821 | 95-97 |
| CGTAT | 476730 | 1.0980961 | 5.933834 | 95-97 |
| GACTT | 473160 | 1.0898732 | 5.7068067 | 7 |
| ATGTG | 401045 | 1.0772069 | 5.1050076 | 7 |
| GTGTA | 389605 | 1.0464789 | 8.578108 | 1 |
| TGGAC | 276220 | 1.0239269 | 6.849471 | 5 |
| GAGTA | 340330 | 0.95291954 | 5.060334 | 1 |
| CGTCT | 309755 | 0.94459146 | 6.1219344 | 95-97 |
| GGACT | 253125 | 0.9383154 | 6.4307027 | 6 |
| AGACT | 383500 | 0.9208381 | 5.1538496 | 6 |
| GGATA | 315915 | 0.88455784 | 5.262639 | 1 |
| GTCCA | 277840 | 0.8832232 | 6.324716 | 1 |
| GTATA | 412805 | 0.71821606 | 7.0910687 | 1 |