![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-136_GGACTCCT-CTAAGCCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1009776 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
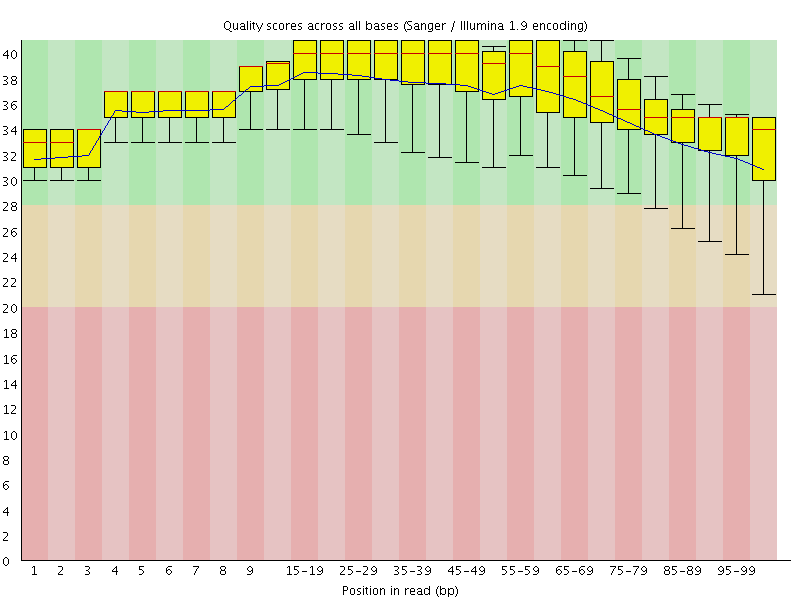
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
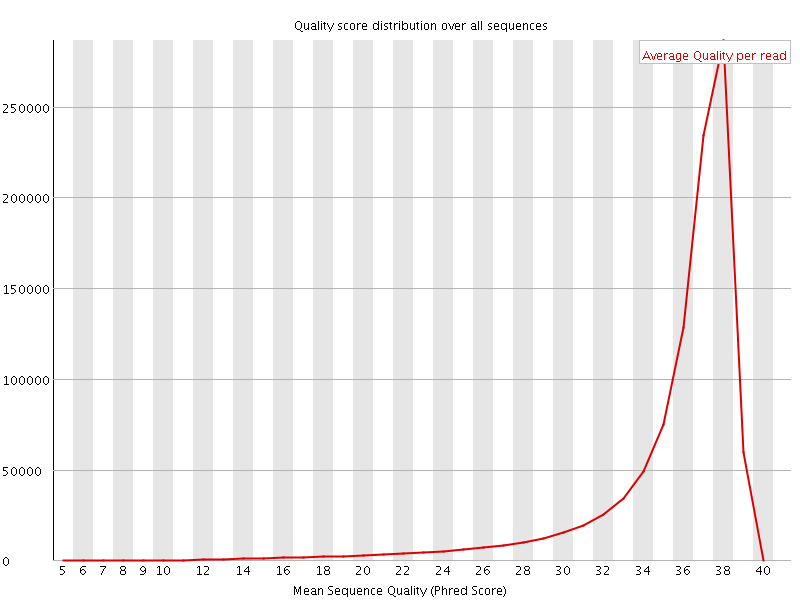
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
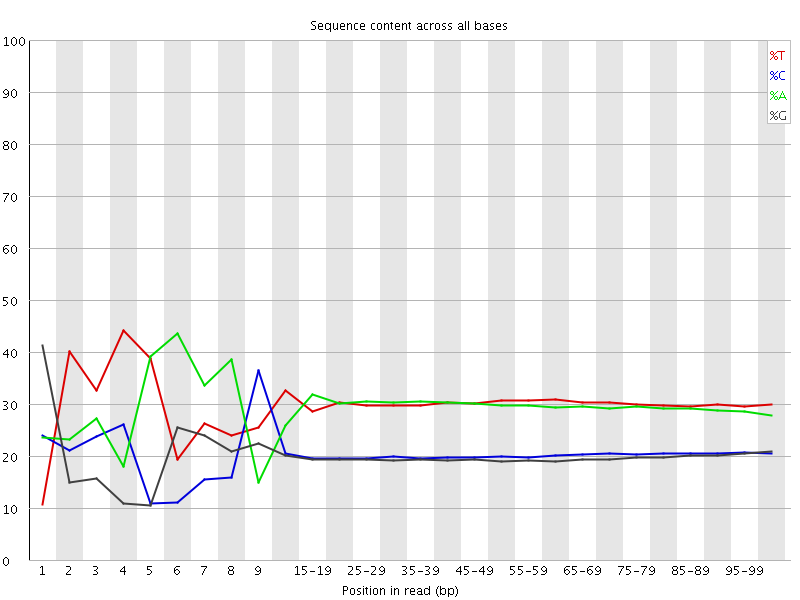
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
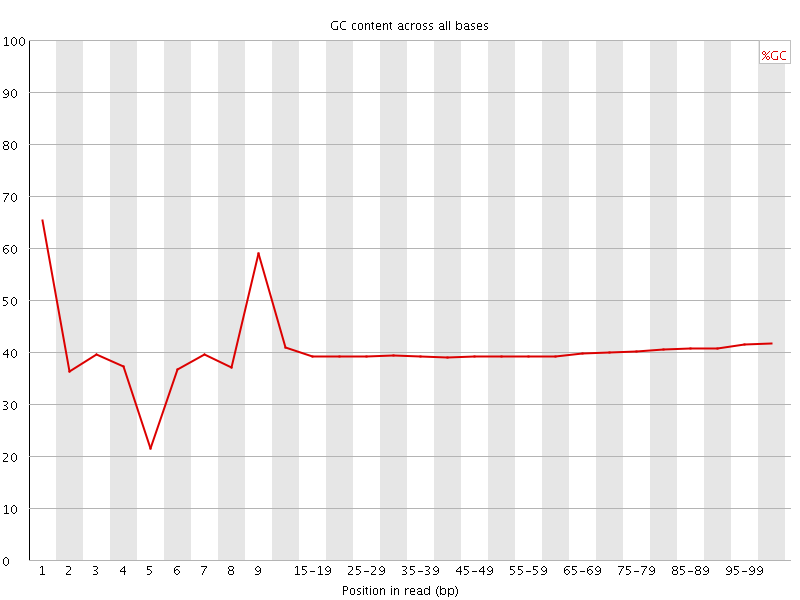
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
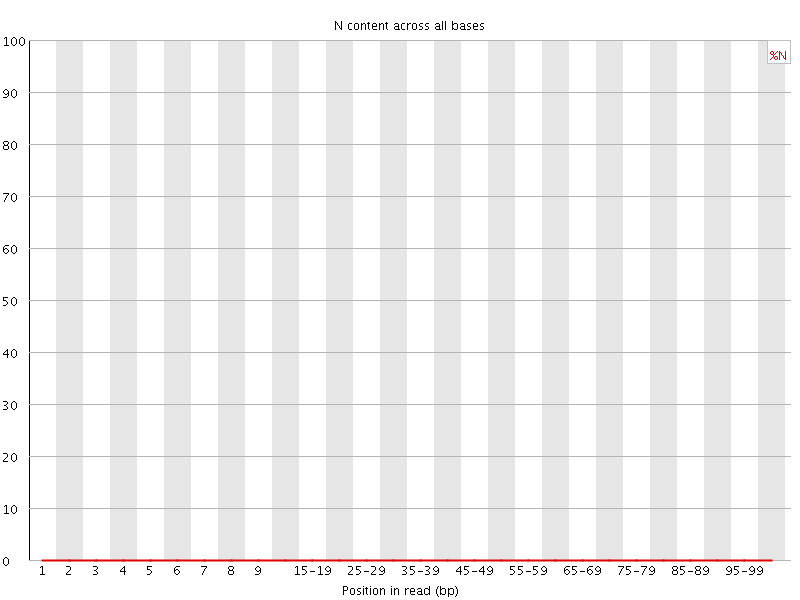
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
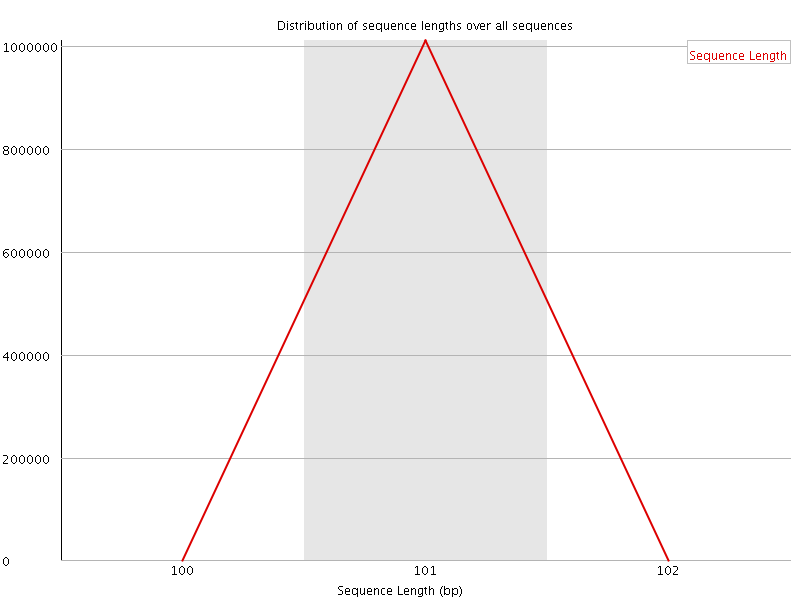
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
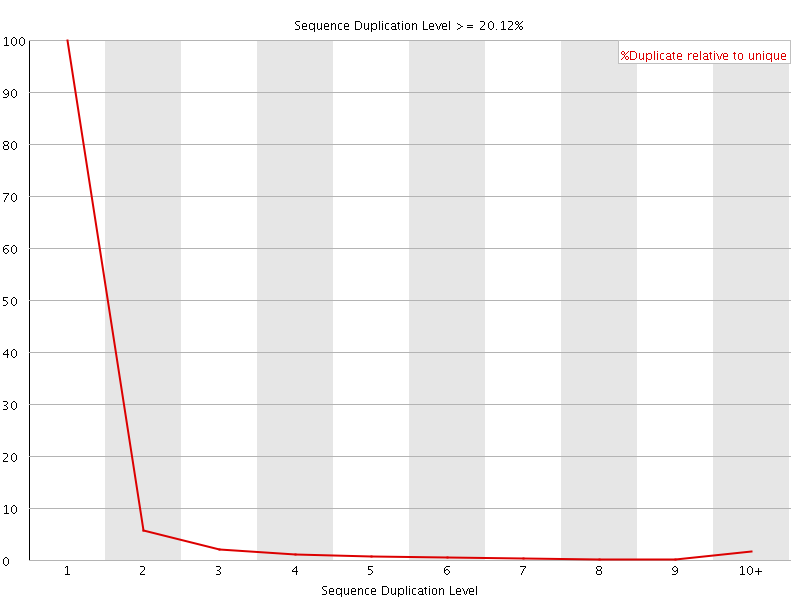
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
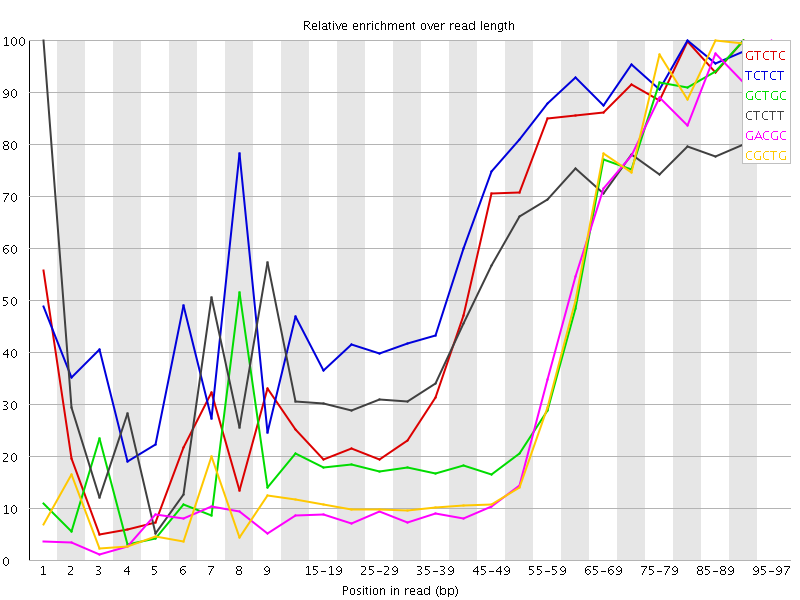
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 227160 | 3.1149921 | 5.219348 | 90-94 |
| TCTCT | 337650 | 3.0400398 | 4.3963337 | 80-84 |
| GCTGC | 144045 | 3.0084019 | 6.8168683 | 90-94 |
| CTCTT | 309535 | 2.7869055 | 5.035512 | 1 |
| GACGC | 123720 | 2.630612 | 6.7481394 | 95-97 |
| CGCTG | 123635 | 2.582136 | 6.3306665 | 85-89 |
| ACACA | 271545 | 2.579835 | 8.067344 | 6 |
| CTGCC | 124505 | 2.5429127 | 6.519806 | 90-94 |
| GCCGA | 117370 | 2.4955945 | 6.680208 | 90-94 |
| GAAGG | 167165 | 2.4843624 | 5.4505205 | 85-89 |
| CGACG | 116460 | 2.4762452 | 6.9543996 | 95-97 |
| CCGAC | 112890 | 2.347358 | 6.4647245 | 95-97 |
| TGCCG | 110180 | 2.3011262 | 6.458299 | 90-94 |
| CAAAG | 236135 | 2.294053 | 5.33921 | 4 |
| TACAC | 243440 | 2.271762 | 10.2211275 | 5 |
| CACAT | 237845 | 2.2195497 | 5.5529084 | 7 |
| CTTTG | 226915 | 2.089146 | 7.8848367 | 9 |
| GACGA | 135815 | 1.9738959 | 5.155372 | 95-97 |
| ATACA | 290570 | 1.853452 | 5.0619497 | 4 |
| GAGAG | 120895 | 1.7967098 | 6.1622705 | 7 |
| GCTTT | 190055 | 1.7497854 | 5.1402316 | 1 |
| TGGAG | 115040 | 1.6793425 | 5.2328568 | 5 |
| CTCCA | 121335 | 1.6565223 | 5.172066 | 1 |
| CACCA | 113280 | 1.5745037 | 5.068895 | 7 |
| TATAC | 247765 | 1.5523559 | 5.9263864 | 5 |
| TTGAG | 156165 | 1.4967921 | 5.8101087 | 9 |
| GACTC | 103500 | 1.4449228 | 5.2469997 | 7 |
| GTGTA | 149770 | 1.4354982 | 11.297594 | 1 |
| GAGTC | 99370 | 1.418576 | 7.282987 | 9 |
| AAGAC | 145760 | 1.4160595 | 5.9282675 | 5 |
| CATGA | 143185 | 1.3663485 | 5.145946 | 9 |
| CCATG | 97245 | 1.3575991 | 5.8765235 | 9 |
| GTCTT | 143540 | 1.3215344 | 6.872995 | 1 |
| GACTT | 136265 | 1.2772299 | 6.409275 | 7 |
| TGAGT | 131925 | 1.2644594 | 5.02933 | 8 |
| AACAC | 132970 | 1.263292 | 5.198322 | 5 |
| TATGA | 196755 | 1.2605793 | 6.4548316 | 4 |
| GTGTT | 133280 | 1.2547685 | 6.101083 | 1 |
| TCATA | 197080 | 1.2347921 | 6.506771 | 2 |
| GATTG | 125640 | 1.2042197 | 6.247153 | 7 |
| GTTCT | 127650 | 1.1752393 | 5.305469 | 1 |
| ACATG | 120475 | 1.1496373 | 5.77542 | 8 |
| ACACC | 81570 | 1.1337594 | 5.59452 | 6 |
| GTCCA | 79305 | 1.1071459 | 7.8350096 | 1 |
| TGGAC | 77195 | 1.1020125 | 7.395586 | 5 |
| ACACT | 114890 | 1.0721439 | 5.5844493 | 6 |
| AGACT | 108485 | 1.0352224 | 5.548524 | 6 |
| GGACT | 71585 | 1.0219257 | 6.895265 | 6 |
| TGTAT | 155080 | 0.9759352 | 5.1893344 | 2 |
| GAGTA | 99995 | 0.9757425 | 5.5804935 | 1 |
| GTATG | 100165 | 0.96004987 | 5.9276614 | 3 |
| CATAC | 99180 | 0.9255396 | 5.6264887 | 3 |
| GTATA | 118640 | 0.7601084 | 7.0918884 | 1 |
| ATACT | 120670 | 0.7560502 | 5.019445 | 6 |