![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-136_GGACTCCT-CTAAGCCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1009776 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
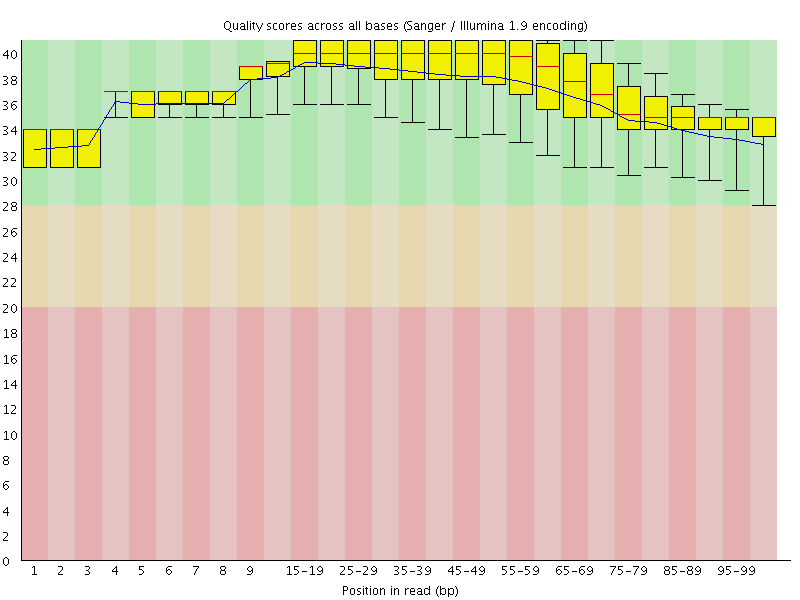
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
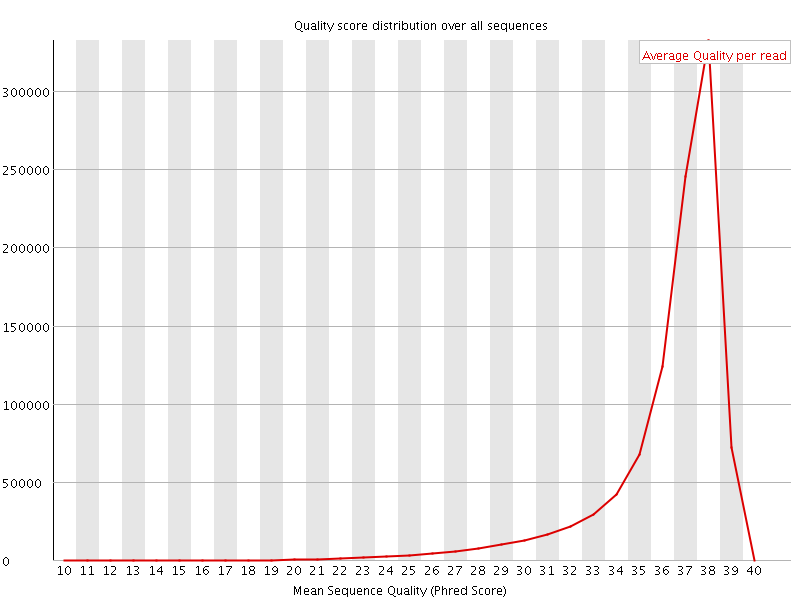
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
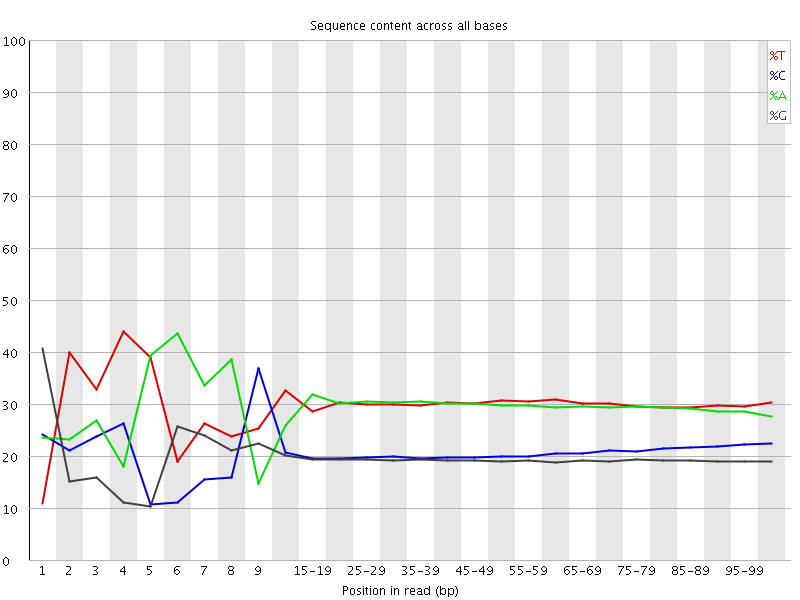
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
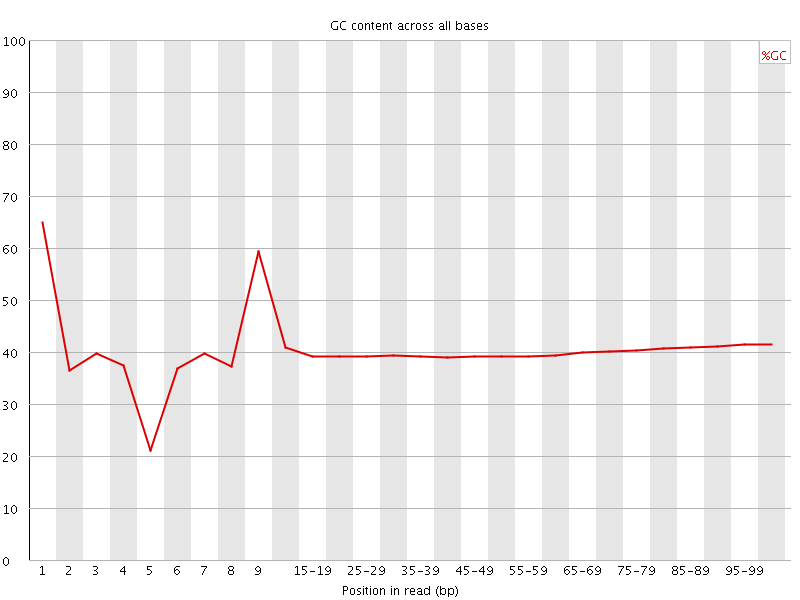
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
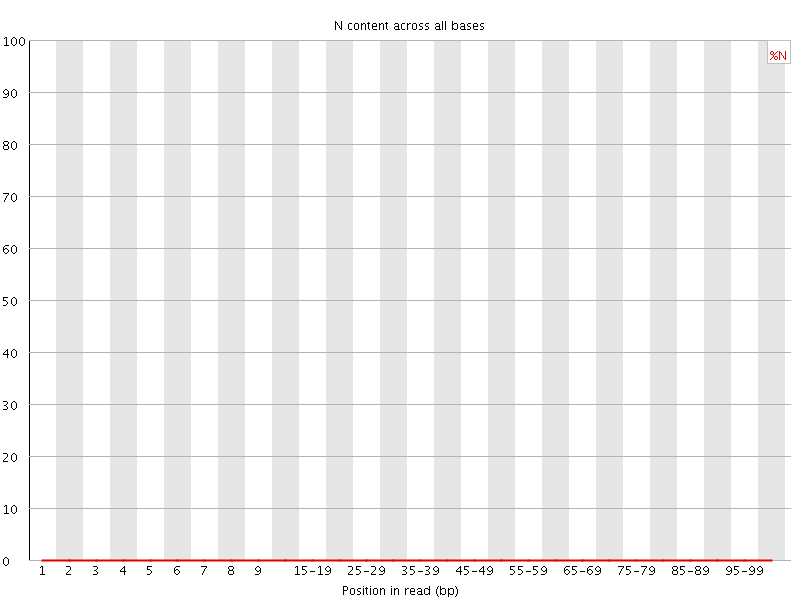
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
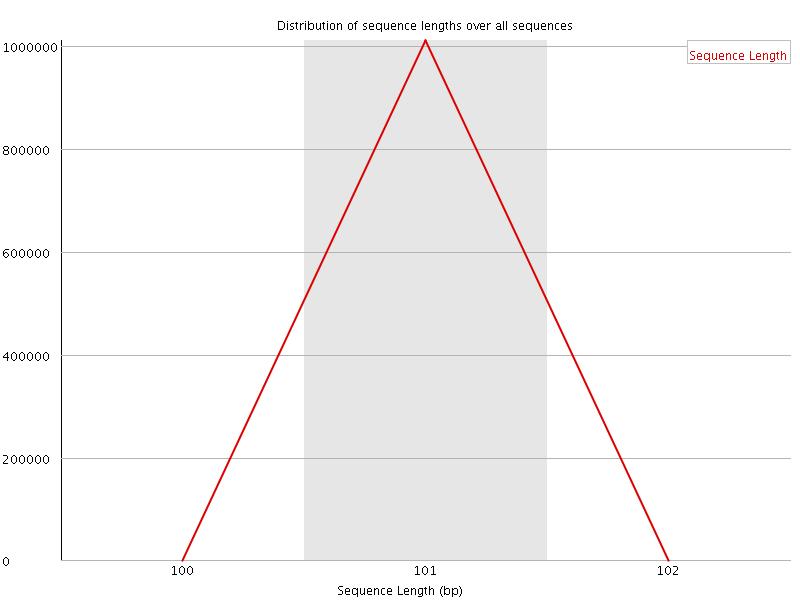
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
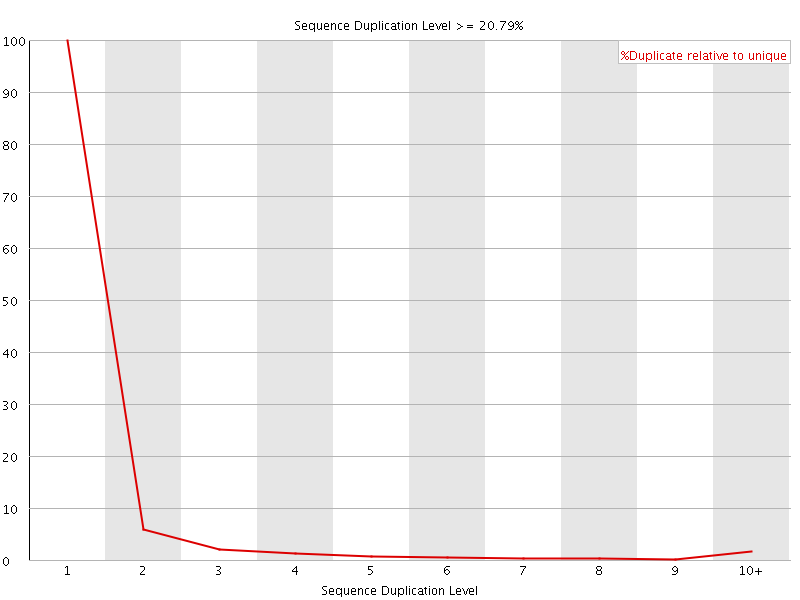
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
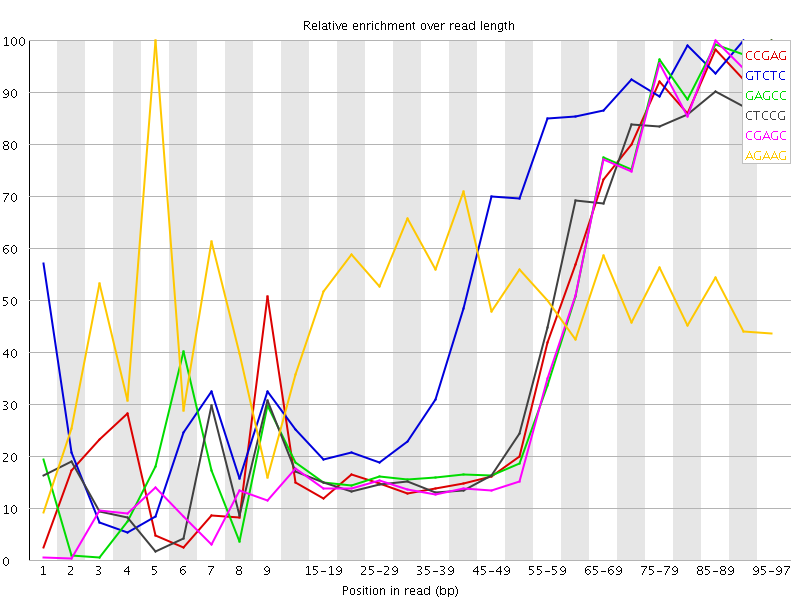
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 146100 | 3.09415 | 7.078827 | 95-97 |
| GTCTC | 227050 | 3.0495944 | 5.1060514 | 90-94 |
| GAGCC | 141905 | 3.0053074 | 6.815254 | 95-97 |
| CTCCG | 153030 | 2.9975297 | 6.895478 | 95-97 |
| CGAGC | 134955 | 2.8581183 | 6.7858696 | 85-89 |
| AGAAG | 271745 | 2.8067327 | 5.483636 | 5 |
| TCTCC | 213760 | 2.7044764 | 5.114345 | 95-97 |
| GCCCA | 133205 | 2.6573448 | 6.62236 | 90-94 |
| AGCCC | 132010 | 2.6335056 | 6.287664 | 90-94 |
| ACACA | 272745 | 2.499589 | 8.269385 | 6 |
| CCCAC | 129235 | 2.4285312 | 6.1669793 | 90-94 |
| ATCTC | 274380 | 2.424283 | 5.665166 | 95-97 |
| CCACG | 116230 | 2.3187056 | 6.231558 | 90-94 |
| CAAAG | 233260 | 2.2694254 | 5.0380435 | 4 |
| GACGG | 99110 | 2.2282941 | 6.6217704 | 85-89 |
| TACAC | 244665 | 2.2016242 | 10.144001 | 5 |
| GACTC | 157620 | 2.1561172 | 5.677405 | 7 |
| GAGAC | 145690 | 2.154741 | 5.321206 | 95-97 |
| CACAT | 237650 | 2.1384995 | 5.767901 | 7 |
| CTTTG | 229955 | 2.117859 | 8.395123 | 9 |
| CGAGA | 139335 | 2.0607512 | 5.1395297 | 95-97 |
| CGGAC | 92760 | 1.9644995 | 6.102003 | 90-94 |
| ACGAG | 130620 | 1.9318572 | 5.010444 | 95-97 |
| AGAGA | 187025 | 1.9316978 | 5.233242 | 8 |
| GAGAG | 118865 | 1.8663074 | 6.2729445 | 7 |
| GGACT | 126885 | 1.8426174 | 7.343857 | 6 |
| ATACA | 290185 | 1.8235615 | 5.0578804 | 4 |
| TGGAG | 113760 | 1.7537928 | 5.4267564 | 5 |
| GCCAA | 113995 | 1.5881346 | 5.006072 | 1 |
| TTGAG | 159010 | 1.5833719 | 5.885417 | 9 |
| CACCA | 117285 | 1.5391471 | 5.274815 | 7 |
| TATAC | 249150 | 1.5373256 | 5.6573305 | 5 |
| ATGGA | 143920 | 1.4595538 | 5.0597534 | 4 |
| GAGTC | 100315 | 1.4567692 | 7.146706 | 9 |
| AGGAG | 92030 | 1.4449693 | 5.0396714 | 5 |
| GTCTT | 153310 | 1.4119675 | 6.132 | 1 |
| AAGAC | 143400 | 1.3951625 | 6.1984916 | 5 |
| CATGA | 142045 | 1.3569415 | 5.4053154 | 9 |
| CCATG | 97365 | 1.3318763 | 5.7503614 | 9 |
| TGAGT | 132660 | 1.3209867 | 5.1901746 | 8 |
| GTGTT | 134570 | 1.3157285 | 6.206333 | 1 |
| TATGA | 198635 | 1.3011417 | 6.491794 | 4 |
| GACTT | 137020 | 1.2852235 | 6.80364 | 7 |
| GTATG | 125785 | 1.2525277 | 5.9143853 | 3 |
| GATTG | 125265 | 1.24735 | 6.426162 | 7 |
| TCATA | 198405 | 1.2242149 | 6.534841 | 2 |
| GTGTA | 120870 | 1.2035857 | 11.772524 | 1 |
| GTTCT | 128825 | 1.1864635 | 5.118188 | 1 |
| ACATG | 119680 | 1.1432909 | 5.9935546 | 8 |
| TGGAC | 77870 | 1.130824 | 8.047965 | 5 |
| ACACC | 82395 | 1.081281 | 5.134832 | 6 |
| GTCCA | 79025 | 1.0809996 | 7.455981 | 1 |
| AGACT | 107485 | 1.0267935 | 5.8870234 | 6 |
| ACACT | 113810 | 1.0241221 | 5.117812 | 6 |
| TGTAT | 157165 | 1.0108441 | 5.183663 | 2 |
| GAGTA | 98990 | 1.0038997 | 5.463745 | 1 |
| TGTAG | 98545 | 0.9812803 | 5.0170846 | 2 |
| TGCCG | 43360 | 0.9016542 | 5.07177 | 95-97 |
| CATAC | 100120 | 0.9009325 | 5.6413746 | 3 |
| GTATA | 118120 | 0.77373505 | 7.077199 | 1 |