![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-135_GGACTCCT-AAGGAGTA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1864430 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
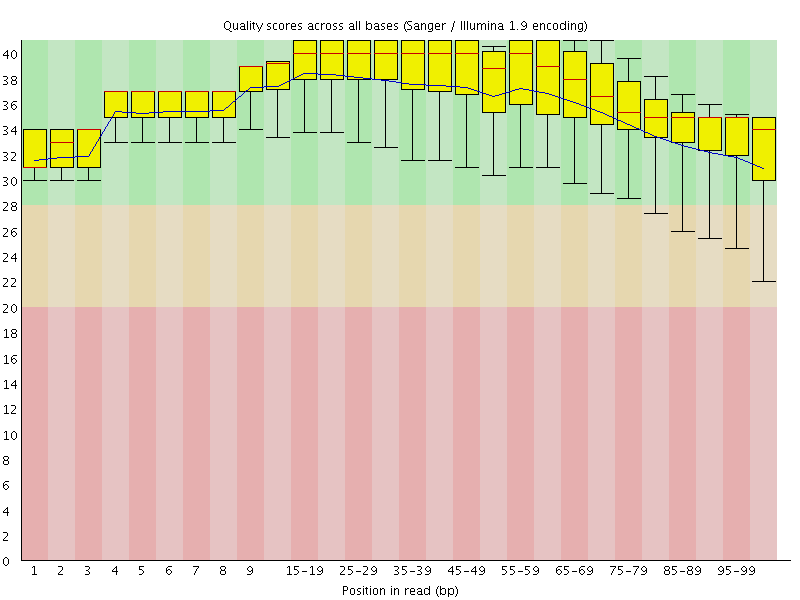
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
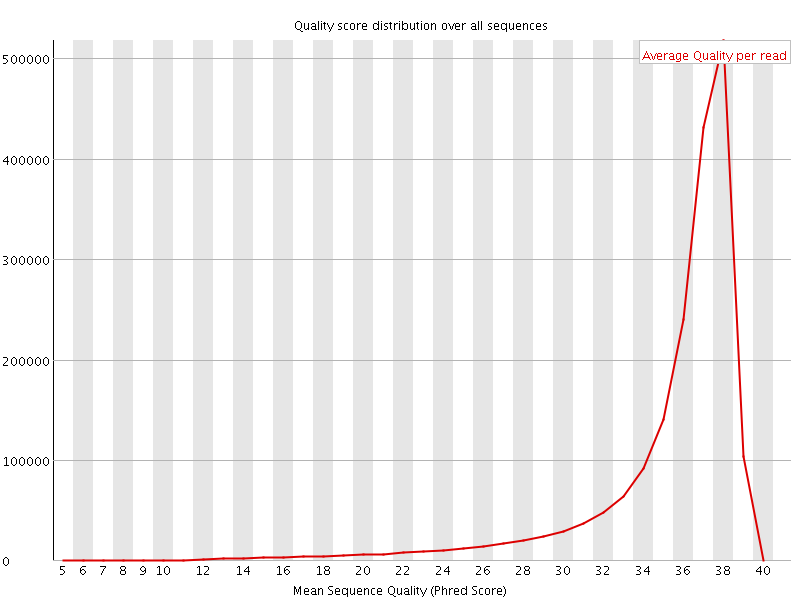
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
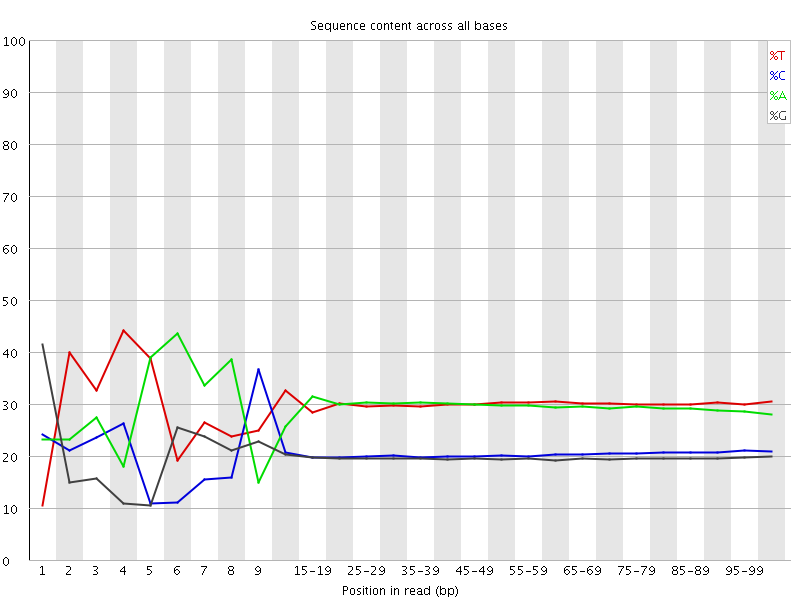
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
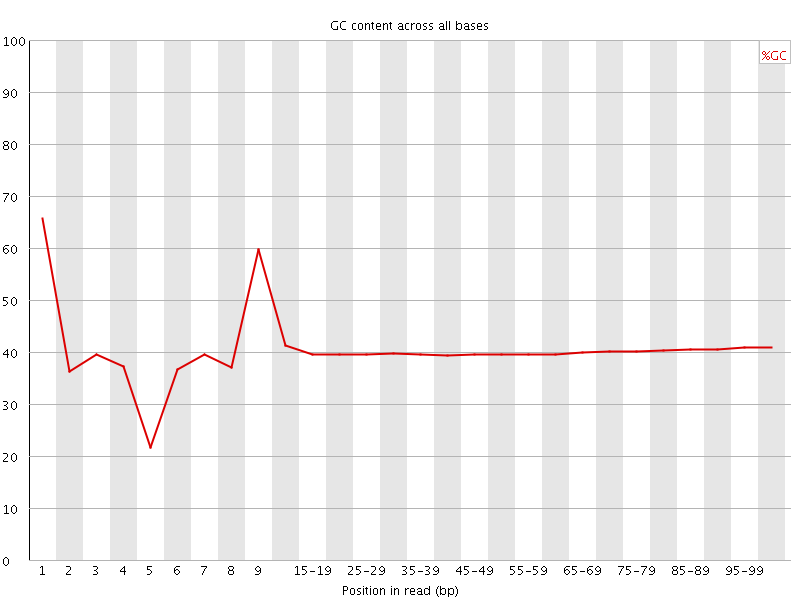
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
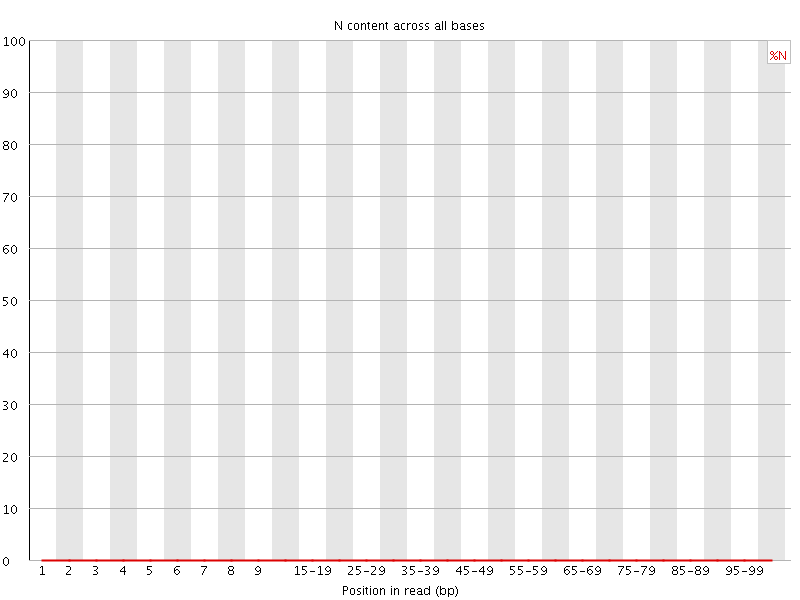
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
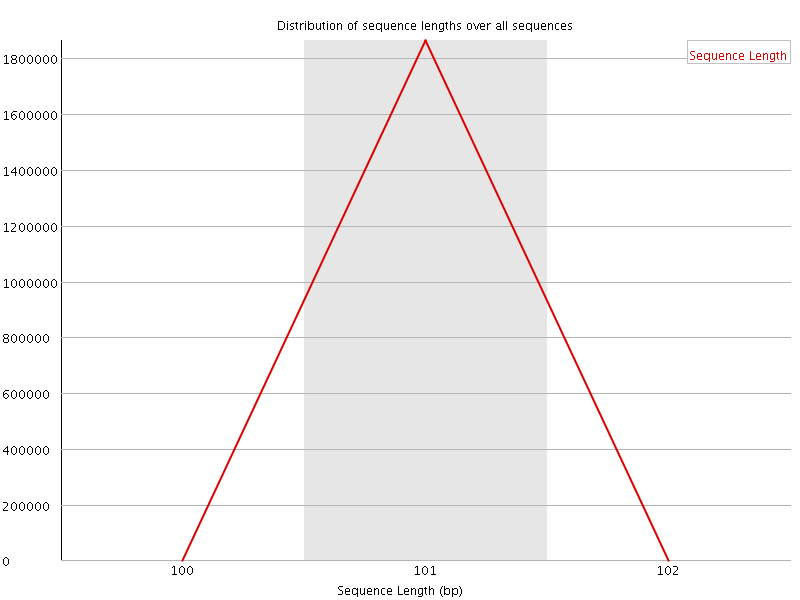
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
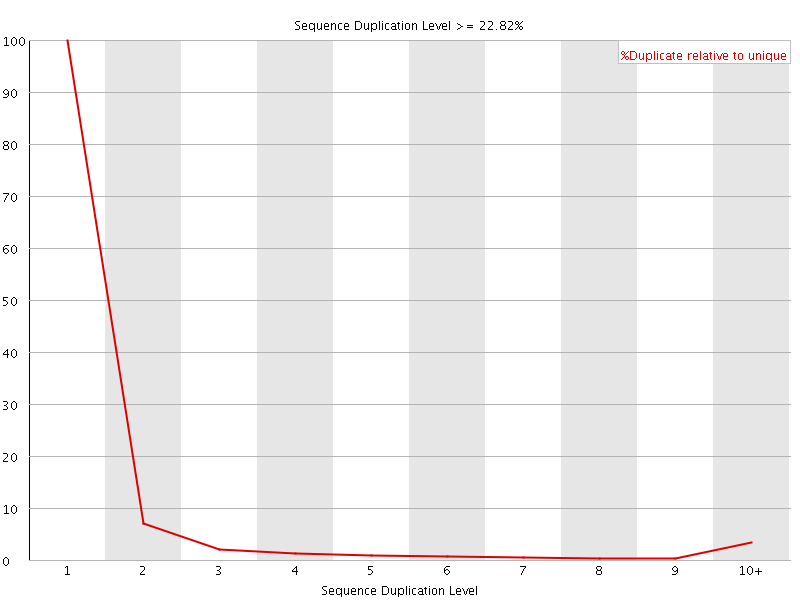
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
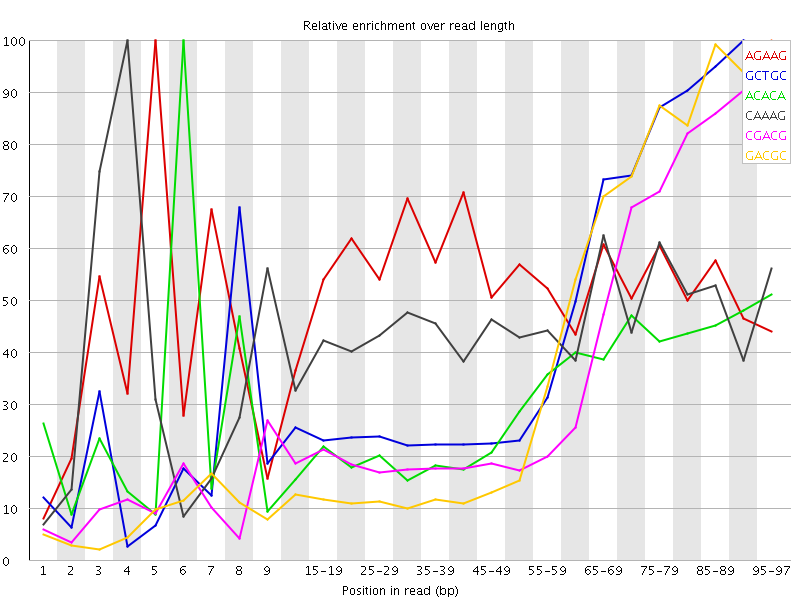
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 492310 | 2.6784558 | 5.0197515 | 5 |
| GCTGC | 229320 | 2.5566175 | 5.47726 | 90-94 |
| ACACA | 450405 | 2.3052876 | 7.458417 | 6 |
| CAAAG | 428620 | 2.2618108 | 5.0198107 | 4 |
| CGACG | 185735 | 2.1119382 | 5.566606 | 95-97 |
| GACGC | 185095 | 2.1046612 | 5.2375364 | 95-97 |
| GCCGA | 184385 | 2.0965877 | 5.364647 | 90-94 |
| CGCTG | 187925 | 2.0951178 | 5.0703826 | 95-97 |
| CTTTG | 418515 | 2.0816324 | 8.403713 | 9 |
| CTGCC | 192360 | 2.0800638 | 5.124783 | 90-94 |
| TACAC | 394645 | 1.9804547 | 9.584062 | 5 |
| CACAT | 380105 | 1.9074882 | 5.0643196 | 7 |
| AGAGA | 348635 | 1.8967793 | 5.013047 | 8 |
| TGCCG | 169935 | 1.8945527 | 5.1387844 | 90-94 |
| GAGAG | 226270 | 1.8348713 | 6.016734 | 7 |
| GCTTT | 353280 | 1.757163 | 5.0115457 | 1 |
| CACCA | 220555 | 1.6319604 | 5.091736 | 7 |
| ATACA | 467915 | 1.6242557 | 5.068488 | 6 |
| TGGCT | 209420 | 1.5834615 | 5.0929785 | 6 |
| TTGAG | 289430 | 1.5137776 | 5.222458 | 9 |
| GACTC | 191455 | 1.4320487 | 5.0237913 | 7 |
| AAGAC | 269630 | 1.4228269 | 5.859432 | 5 |
| GAGTC | 181810 | 1.4020736 | 7.2777305 | 9 |
| CCATG | 186500 | 1.3949862 | 6.064953 | 9 |
| GTGTA | 264810 | 1.38501 | 10.443261 | 1 |
| TATAC | 396835 | 1.3506218 | 6.1978908 | 5 |
| GTCTT | 267615 | 1.3310778 | 6.57509 | 1 |
| GACTT | 250715 | 1.2718531 | 6.438001 | 7 |
| TGAGT | 242725 | 1.269501 | 5.3137975 | 8 |
| GTGTT | 246690 | 1.2650464 | 6.057537 | 1 |
| TATGA | 359725 | 1.262282 | 6.1109123 | 4 |
| AACAC | 246440 | 1.2613426 | 5.2437468 | 5 |
| TCATA | 362110 | 1.2324361 | 6.008026 | 2 |
| GATTG | 230945 | 1.2078893 | 5.8817983 | 7 |
| GTTCT | 235335 | 1.1705219 | 5.1756215 | 1 |
| ACACC | 157140 | 1.1627316 | 5.576016 | 6 |
| ACCAT | 231455 | 1.1615151 | 5.198307 | 8 |
| ACATG | 221845 | 1.1478097 | 5.296767 | 8 |
| TGGAC | 142755 | 1.100891 | 7.2312064 | 5 |
| GTCCA | 144775 | 1.0828907 | 7.917498 | 1 |
| ACACT | 212620 | 1.066995 | 5.4559975 | 6 |
| AGACT | 198000 | 1.0244374 | 5.419438 | 6 |
| GGACT | 131940 | 1.0174885 | 6.918389 | 6 |
| GAGTA | 185355 | 0.9887501 | 5.330791 | 1 |
| GTATG | 186090 | 0.9732885 | 5.335577 | 3 |
| GTATA | 222245 | 0.779862 | 7.5938177 | 1 |