![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-135_GGACTCCT-AAGGAGTA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1864430 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
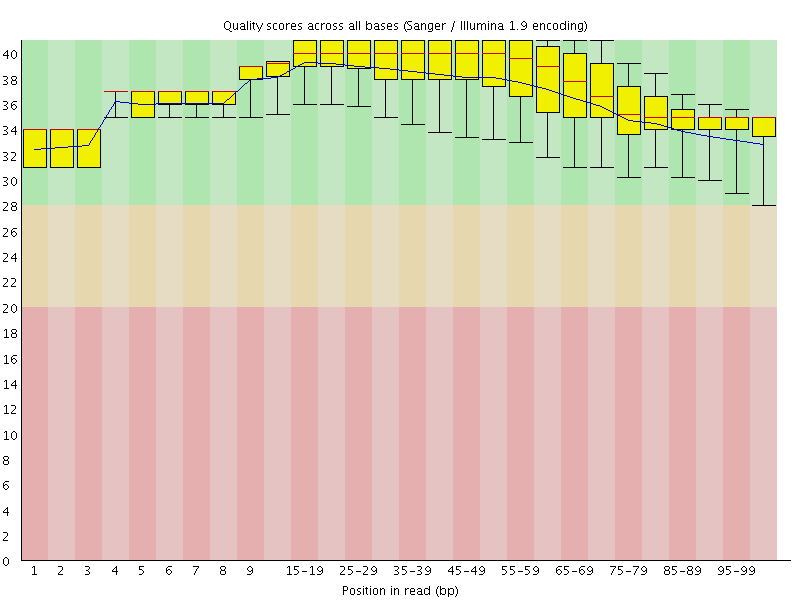
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
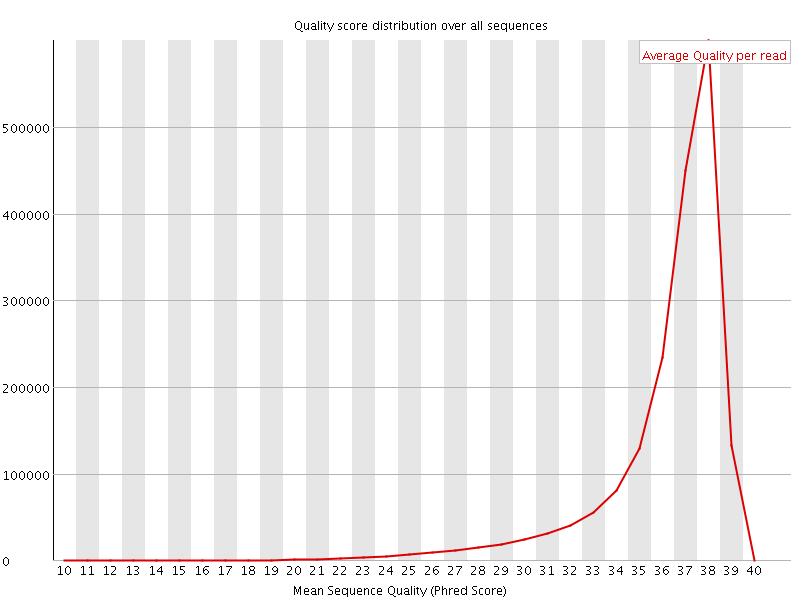
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
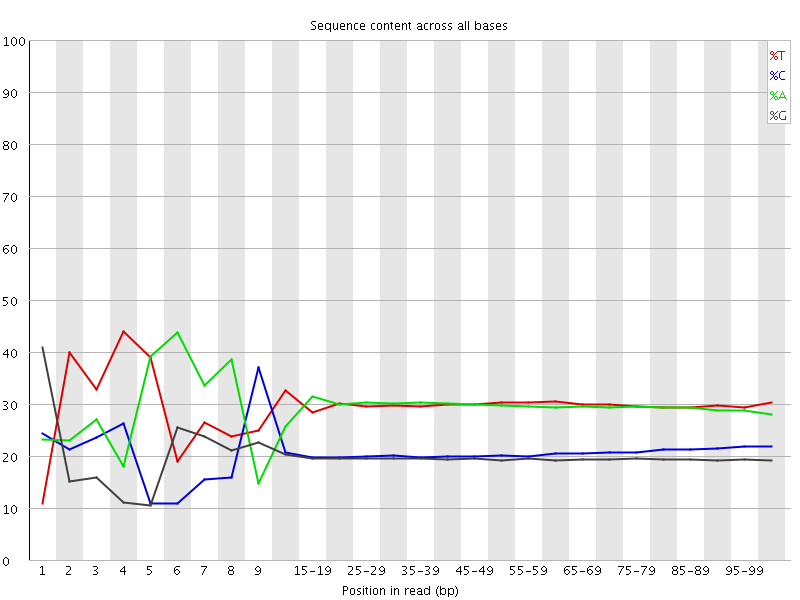
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
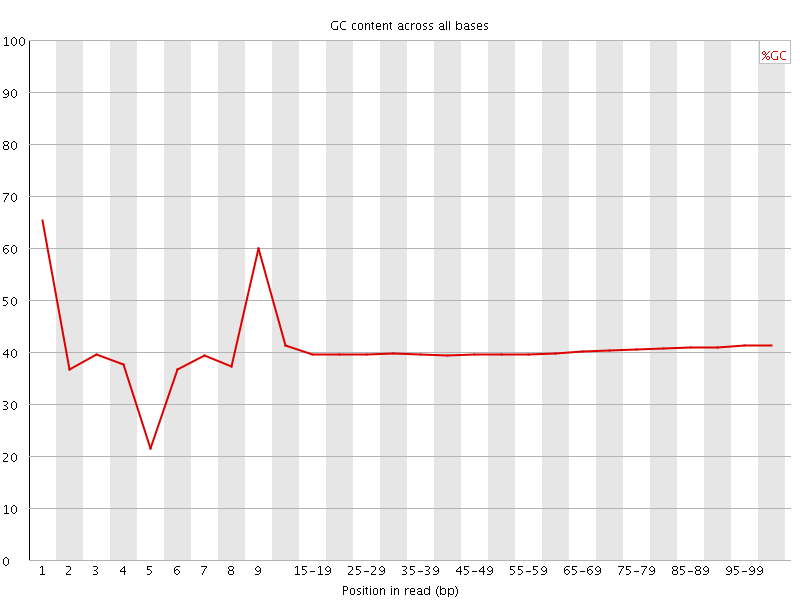
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
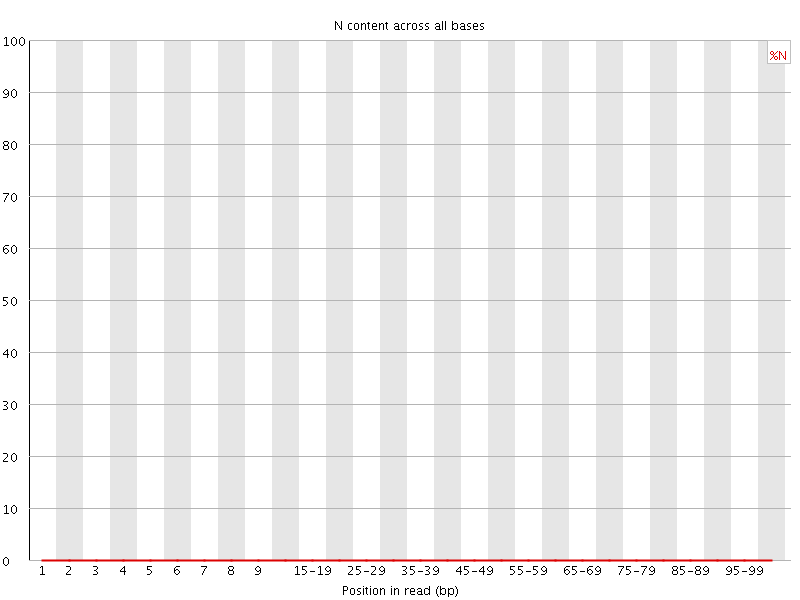
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
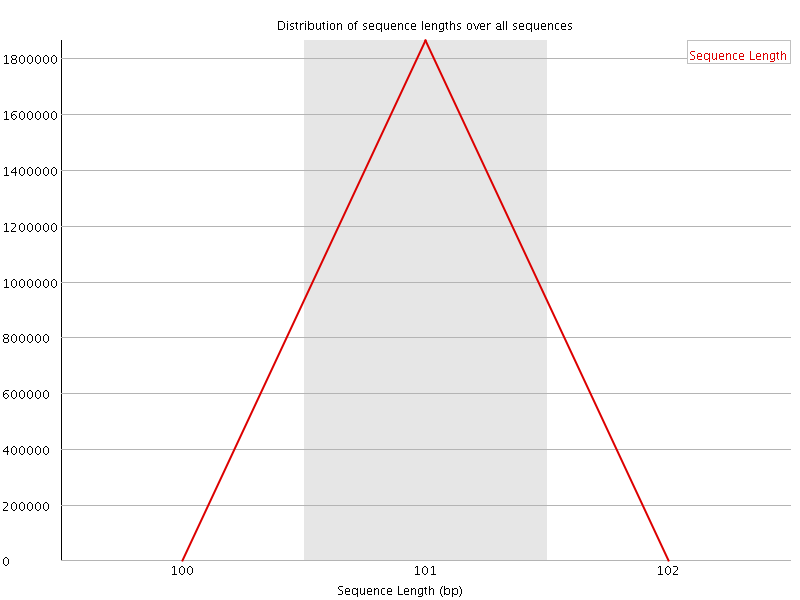
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
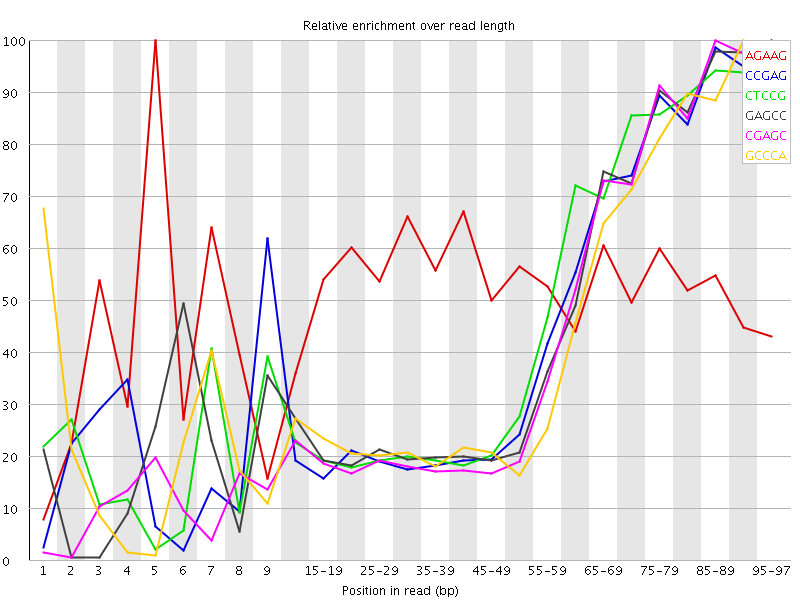
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 494465 | 2.7067144 | 5.162141 | 5 |
| CCGAG | 228710 | 2.569967 | 5.6463604 | 95-97 |
| CTCCG | 237025 | 2.5104141 | 5.310048 | 95-97 |
| GAGCC | 222940 | 2.5051303 | 5.484723 | 95-97 |
| CGAGC | 210660 | 2.3671427 | 5.403843 | 85-89 |
| GCCCA | 213280 | 2.2891936 | 5.1960354 | 90-94 |
| ACACA | 451090 | 2.2529433 | 7.276708 | 6 |
| AGCCC | 207685 | 2.229141 | 5.088033 | 95-97 |
| CTTTG | 418015 | 2.1001256 | 8.264958 | 9 |
| GACTC | 263525 | 1.9480736 | 5.250789 | 7 |
| TACAC | 393405 | 1.9388568 | 9.548492 | 5 |
| AGAGA | 346885 | 1.8988575 | 5.02413 | 8 |
| GACGG | 157995 | 1.8586415 | 5.106975 | 95-97 |
| GAGAG | 223460 | 1.8347813 | 5.7763743 | 7 |
| GCTTT | 351195 | 1.7644188 | 5.2086487 | 1 |
| ATACA | 472050 | 1.6237762 | 5.071752 | 6 |
| GGACT | 204720 | 1.5843593 | 6.566514 | 6 |
| TTGAG | 289095 | 1.5409378 | 5.5382185 | 9 |
| GAGTC | 182350 | 1.4112345 | 7.0468073 | 9 |
| AAGAC | 269695 | 1.410165 | 5.759818 | 5 |
| GTCTT | 279705 | 1.4052501 | 6.531519 | 1 |
| CCATG | 185195 | 1.3690295 | 5.9174423 | 9 |
| TATAC | 400430 | 1.3592007 | 6.2439256 | 5 |
| GTGTT | 248645 | 1.3078052 | 6.3252687 | 1 |
| TGAGT | 243370 | 1.2972138 | 5.277201 | 8 |
| TATGA | 359995 | 1.2792733 | 6.2732267 | 4 |
| GACTT | 249685 | 1.2712386 | 6.5514727 | 7 |
| TCATA | 364825 | 1.2383447 | 6.0604477 | 2 |
| AACAC | 246710 | 1.232179 | 5.0682697 | 5 |
| GATTG | 229740 | 1.224563 | 5.8457537 | 7 |
| GTGTA | 224425 | 1.1962329 | 11.22014 | 1 |
| GTTCT | 236860 | 1.1899949 | 5.169669 | 1 |
| GTATG | 216545 | 1.1542308 | 5.548556 | 3 |
| ACATG | 221700 | 1.1438826 | 5.163318 | 8 |
| ACCAT | 228885 | 1.1280366 | 5.1159463 | 8 |
| ACACC | 155635 | 1.1136842 | 5.3116984 | 6 |
| TGGAC | 142780 | 1.1049962 | 7.028046 | 5 |
| GTCCA | 145170 | 1.0731499 | 7.531362 | 1 |
| ACACT | 213145 | 1.0504636 | 5.4146347 | 6 |
| AGACT | 199370 | 1.0286688 | 5.7837167 | 6 |
| GAGTA | 185330 | 1.001086 | 5.3041368 | 1 |
| TGTAT | 284845 | 0.9988362 | 5.114786 | 2 |
| GTATA | 221410 | 0.78679955 | 7.499283 | 1 |
| ATACT | 222625 | 0.7556678 | 5.0063314 | 6 |