![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-134_GGACTCCT-ACTGCATA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1908600 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
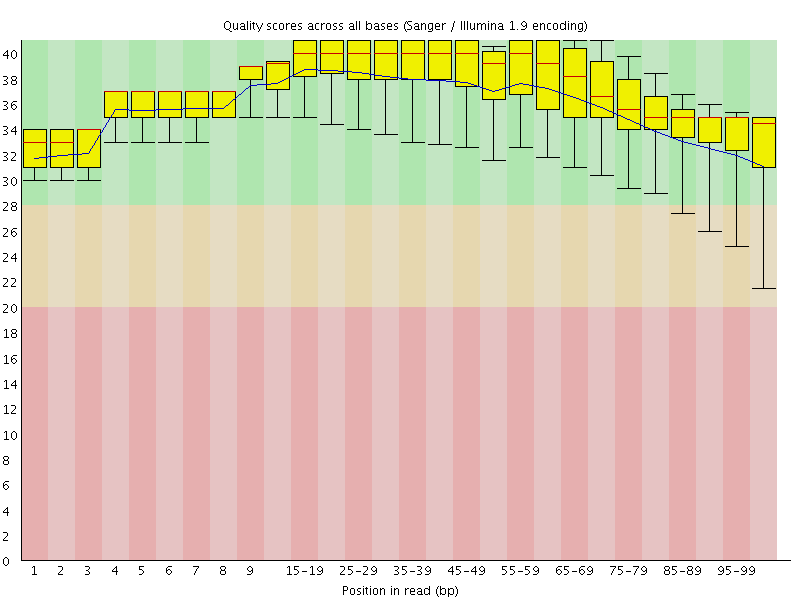
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
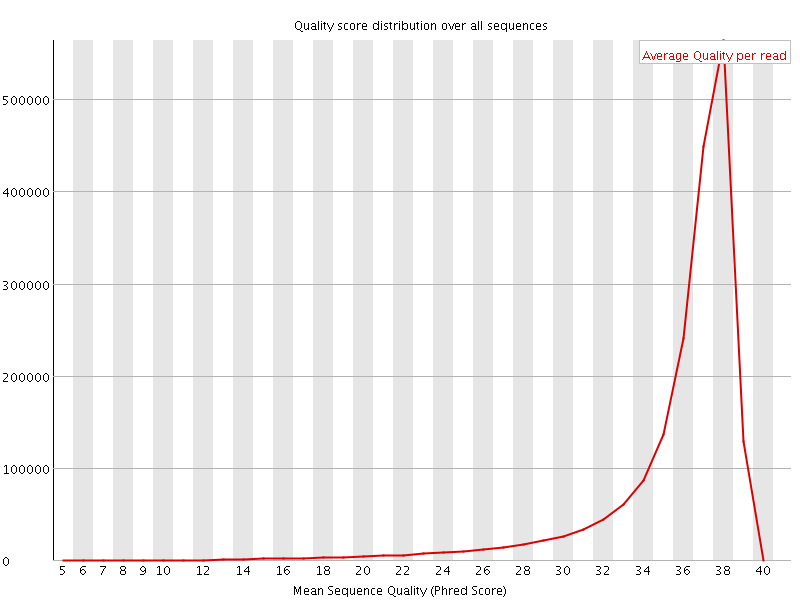
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
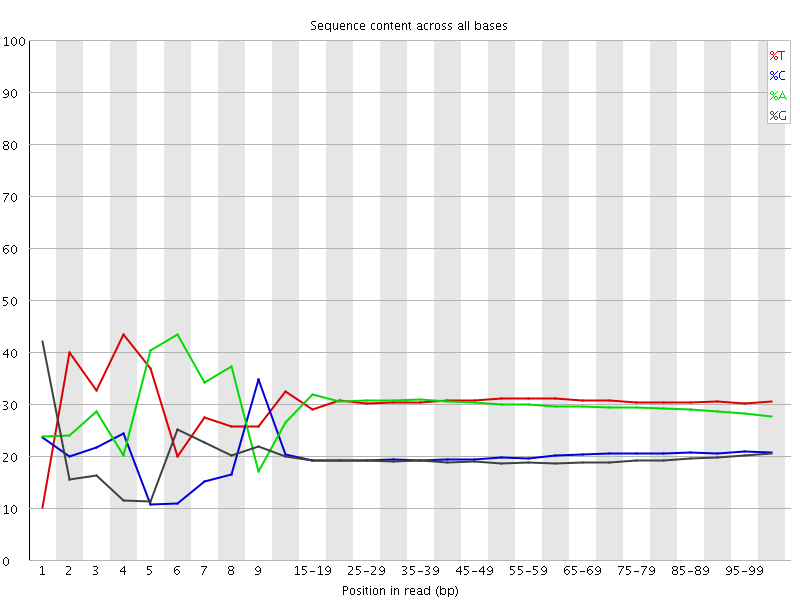
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
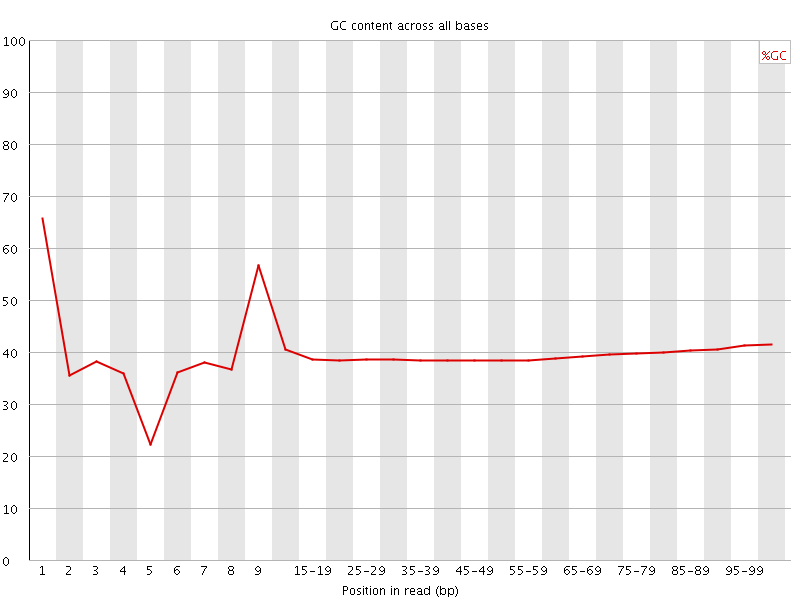
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
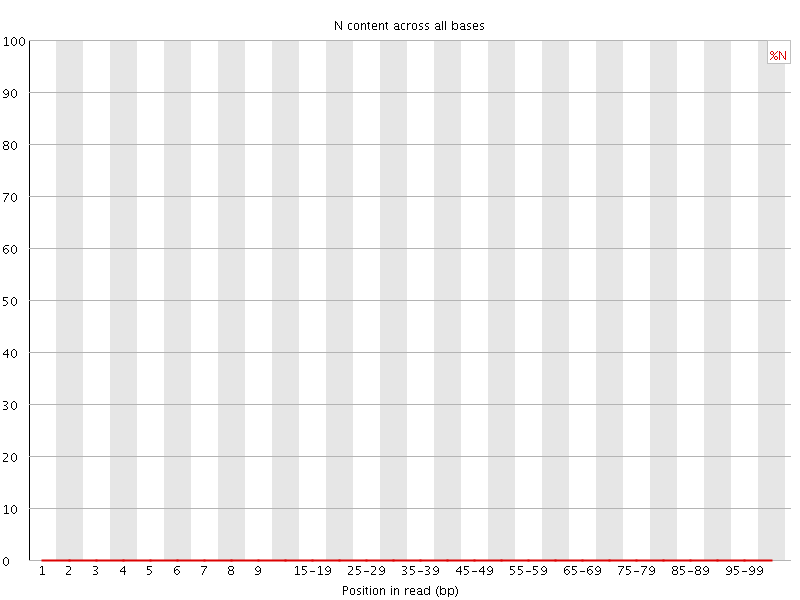
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
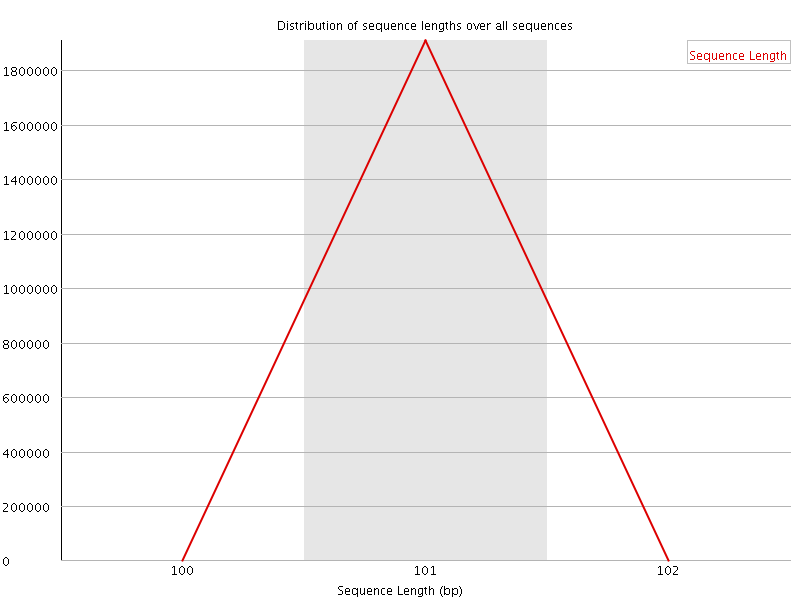
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
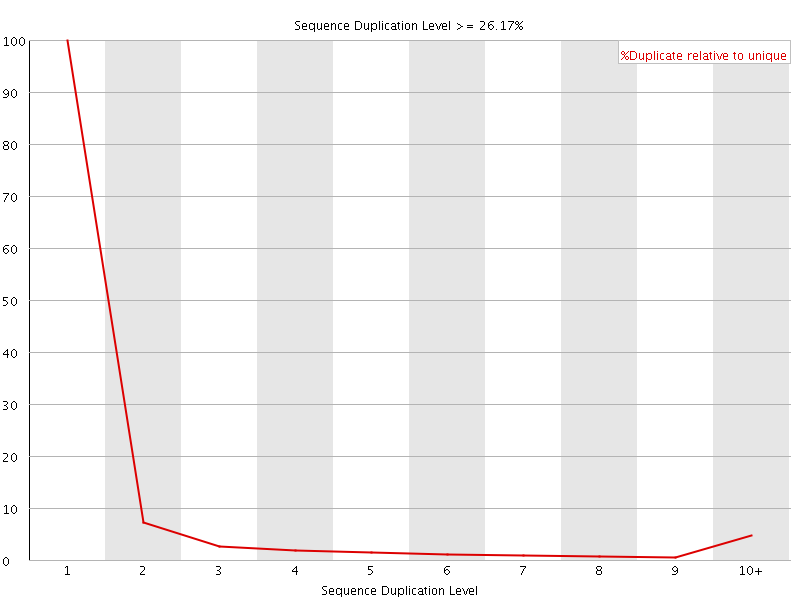
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
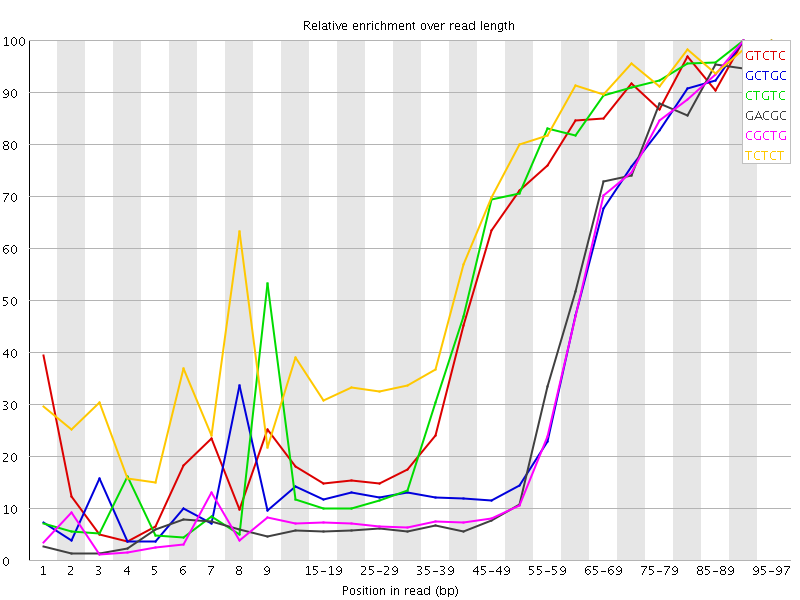
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 530175 | 3.89188 | 6.963754 | 90-94 |
| GCTGC | 327770 | 3.785322 | 9.548638 | 90-94 |
| CTGTC | 500005 | 3.6704097 | 6.5692835 | 90-94 |
| GACGC | 300270 | 3.558641 | 9.525047 | 95-97 |
| CGCTG | 297745 | 3.4385715 | 9.282028 | 90-94 |
| TCTCT | 731375 | 3.4126122 | 5.2294564 | 95-97 |
| GCCGA | 279305 | 3.310175 | 9.674505 | 95-97 |
| CTGCC | 294540 | 3.2960956 | 9.00076 | 90-94 |
| CGACG | 273925 | 3.246414 | 9.670673 | 95-97 |
| CTCTT | 676430 | 3.1562376 | 4.985033 | 95-97 |
| TGCCG | 268520 | 3.1010604 | 9.214501 | 95-97 |
| CCGAC | 266480 | 3.0602632 | 9.30957 | 95-97 |
| TGTCT | 609145 | 2.9332263 | 5.02062 | 90-94 |
| ACACA | 564670 | 2.8474588 | 5.8812466 | 6 |
| CTGAC | 351440 | 2.6474638 | 6.592771 | 95-97 |
| TGACG | 335110 | 2.6052196 | 6.53222 | 95-97 |
| TACAC | 511710 | 2.5144794 | 6.6410336 | 5 |
| GACGA | 308195 | 2.4587884 | 6.610595 | 95-97 |
| AGAAG | 457035 | 2.45453 | 5.12192 | 5 |
| ATCTG | 494215 | 2.4421902 | 5.165153 | 95-97 |
| ACGCT | 306080 | 2.3057585 | 5.9136114 | 90-94 |
| TGCAG | 261100 | 2.0298493 | 5.7740417 | 90-94 |
| GGTGG | 147625 | 1.8157233 | 5.641573 | 95-97 |
| CTTTG | 373885 | 1.8003749 | 6.919131 | 9 |
| GAGAG | 216050 | 1.7788032 | 5.573937 | 7 |
| GTGTG | 217110 | 1.6973686 | 5.490821 | 95-97 |
| GCAGT | 211675 | 1.6456084 | 5.525426 | 95-97 |
| CAGTG | 208035 | 1.6173103 | 5.651118 | 95-97 |
| CTCGG | 129685 | 1.497695 | 5.9189415 | 95-97 |
| GTGTA | 271140 | 1.3827231 | 9.22508 | 1 |
| GAGTC | 161870 | 1.2584133 | 6.9709 | 9 |
| AAGAC | 239430 | 1.246004 | 5.2256656 | 5 |
| CGGTG | 104305 | 1.2431307 | 5.1487203 | 95-97 |
| GTCTT | 241255 | 1.1617193 | 5.5104737 | 1 |
| TGAGT | 216535 | 1.1042559 | 5.003167 | 8 |
| GTGTT | 221490 | 1.1006699 | 5.6940193 | 1 |
| TGGAC | 133145 | 1.0350988 | 5.690843 | 5 |
| GTCCA | 134400 | 1.0124606 | 7.177223 | 1 |
| GGACT | 119350 | 0.9278534 | 5.161215 | 6 |
| GAGTA | 169660 | 0.8878911 | 5.455729 | 1 |
| GTATA | 211745 | 0.70436865 | 6.9098663 | 1 |