![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-134_GGACTCCT-ACTGCATA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1908600 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
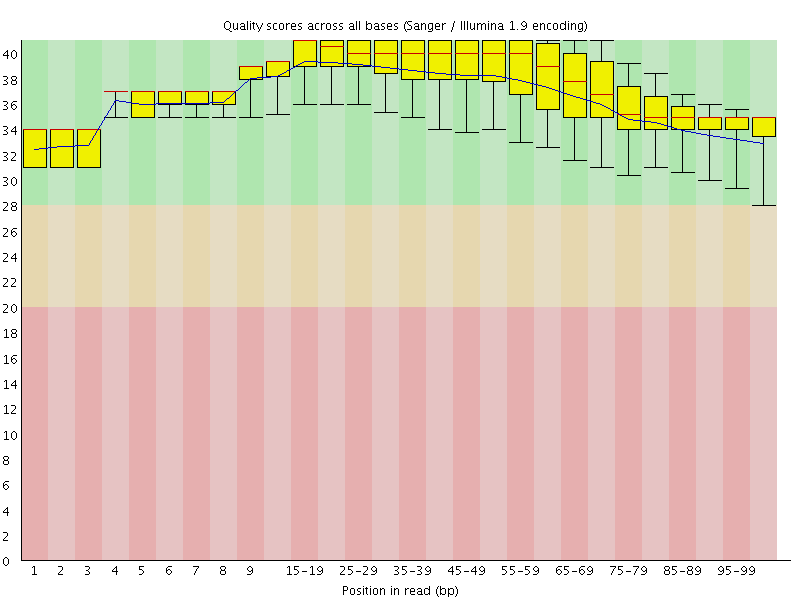
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
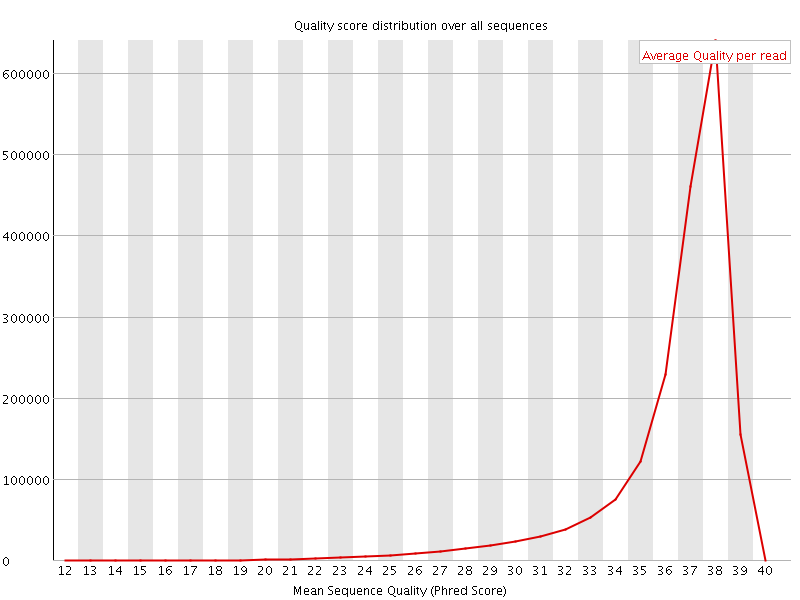
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
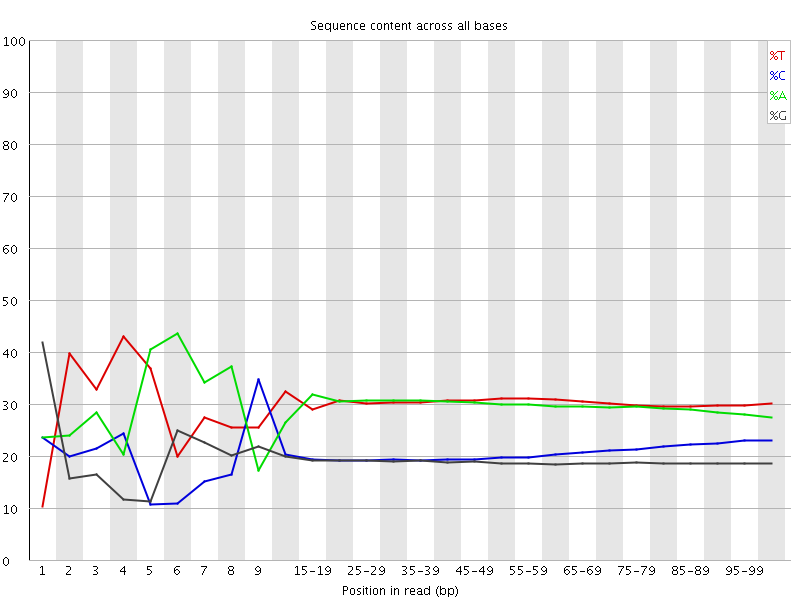
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
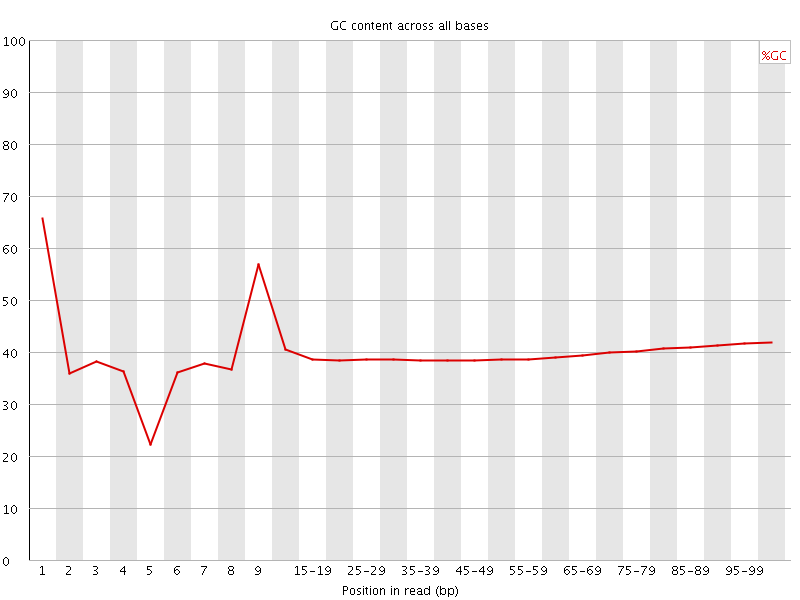
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
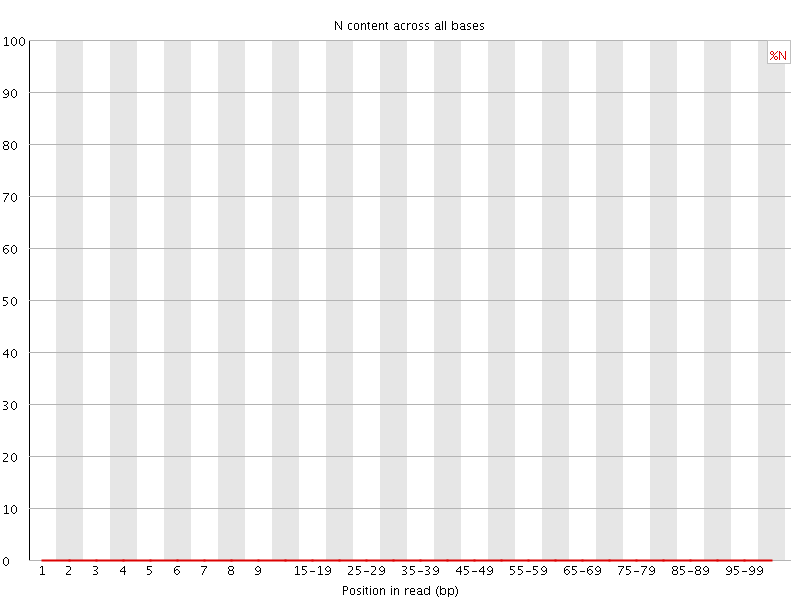
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
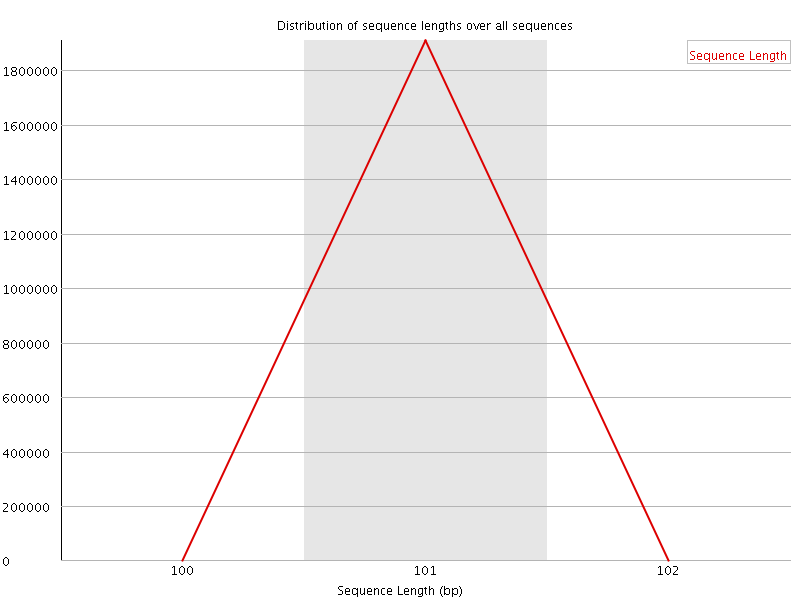
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
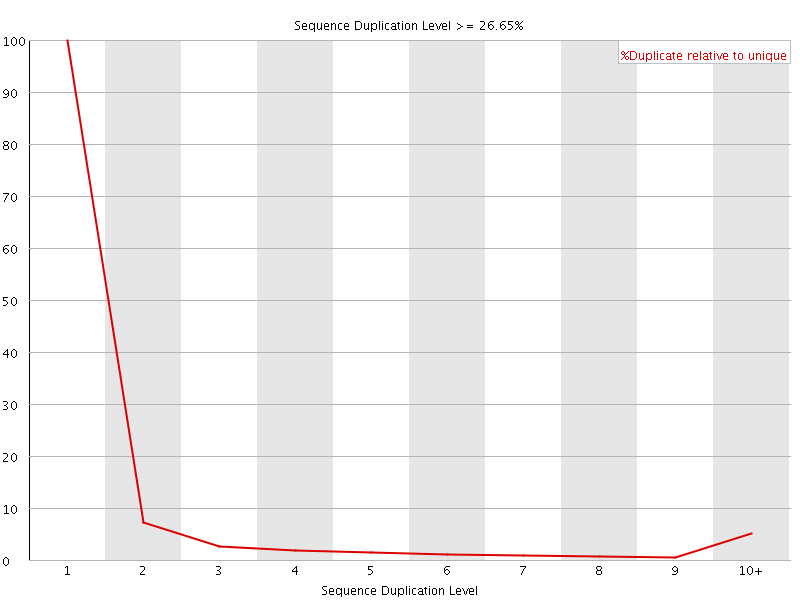
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
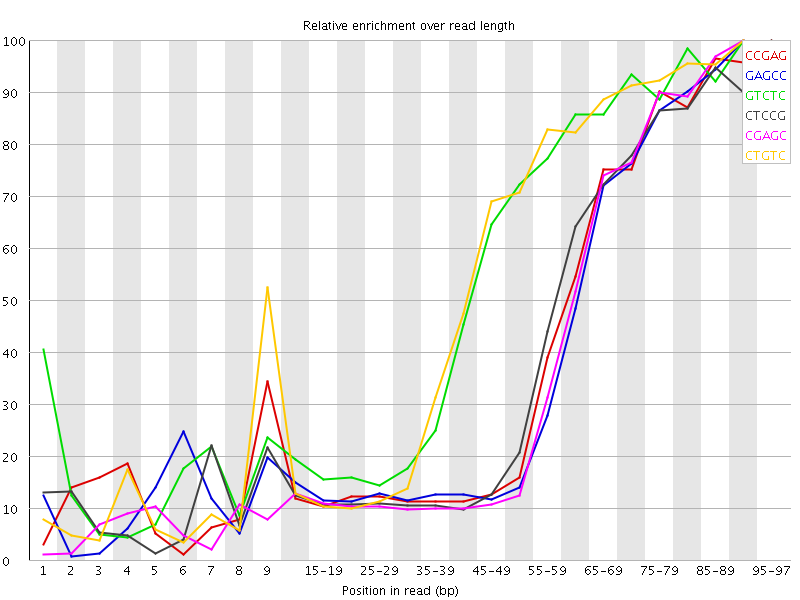
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 347355 | 4.0342007 | 9.699199 | 95-97 |
| GAGCC | 334040 | 3.8795595 | 9.5398855 | 90-94 |
| GTCTC | 530900 | 3.8169951 | 6.7318892 | 90-94 |
| CTCCG | 358030 | 3.7983513 | 9.137658 | 95-97 |
| CGAGC | 321755 | 3.7368803 | 9.306737 | 90-94 |
| CTGTC | 501655 | 3.6067333 | 6.432074 | 90-94 |
| AGCCC | 313640 | 3.3899255 | 8.7093 | 90-94 |
| GCCCA | 311305 | 3.364688 | 8.792106 | 90-94 |
| TCTCT | 735560 | 3.3352993 | 5.08469 | 95-97 |
| TCTCC | 470265 | 3.1464891 | 6.4982924 | 95-97 |
| CTCTT | 675410 | 3.062557 | 4.7886086 | 95-97 |
| CCCAC | 298170 | 2.9991446 | 8.205031 | 95-97 |
| TGTCT | 610865 | 2.9763713 | 5.029474 | 90-94 |
| CCACG | 274820 | 2.9703455 | 8.664705 | 95-97 |
| GACGG | 233605 | 2.915354 | 9.222079 | 95-97 |
| ATCTC | 609735 | 2.816701 | 7.1513476 | 95-97 |
| TCCGA | 383475 | 2.8088517 | 6.3053093 | 95-97 |
| ACACA | 565265 | 2.7103014 | 5.828093 | 6 |
| GAGAC | 319135 | 2.5590243 | 6.6654572 | 95-97 |
| CGAGA | 315100 | 2.526669 | 6.6382403 | 95-97 |
| AGAAG | 456115 | 2.5251684 | 5.42755 | 5 |
| CGGAC | 217075 | 2.521121 | 8.482275 | 90-94 |
| TACAC | 512750 | 2.4131718 | 6.790819 | 5 |
| ACGAG | 298595 | 2.3943217 | 6.646017 | 95-97 |
| GACTC | 302040 | 2.212362 | 5.7690144 | 95-97 |
| CACGA | 288090 | 2.149824 | 6.183744 | 95-97 |
| GGACT | 262695 | 2.0676107 | 6.424241 | 95-97 |
| AGACG | 256000 | 2.0527685 | 6.2248087 | 95-97 |
| CTCCT | 303005 | 2.0273716 | 5.582324 | 95-97 |
| ACGGA | 246535 | 1.9768718 | 5.9839005 | 90-94 |
| ACTCC | 278840 | 1.9007351 | 5.337941 | 95-97 |
| GAGAG | 218410 | 1.8819052 | 5.8361745 | 7 |
| CTTTG | 371770 | 1.811408 | 6.848521 | 9 |
| GTCTT | 265830 | 1.2952272 | 5.315967 | 1 |
| GAGTC | 161975 | 1.2748674 | 6.8995714 | 9 |
| AAGAC | 243760 | 1.2558947 | 5.488169 | 5 |
| TGAGT | 217170 | 1.1583772 | 5.211129 | 8 |
| GTGTT | 219105 | 1.147148 | 5.884891 | 1 |
| GCCGT | 96070 | 1.0951883 | 5.997434 | 95-97 |
| TGCCG | 92670 | 1.0564287 | 6.423058 | 95-97 |
| TGGAC | 132300 | 1.0413023 | 5.8425016 | 5 |
| GTGTA | 188450 | 1.0051856 | 9.681169 | 1 |
| GTCCA | 135330 | 0.9912561 | 6.6631947 | 1 |
| GAGTA | 171075 | 0.92965037 | 5.538902 | 1 |
| GTATA | 211585 | 0.7251466 | 7.046357 | 1 |