![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-133_GGACTCCT-GTAAGGAG_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1307432 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
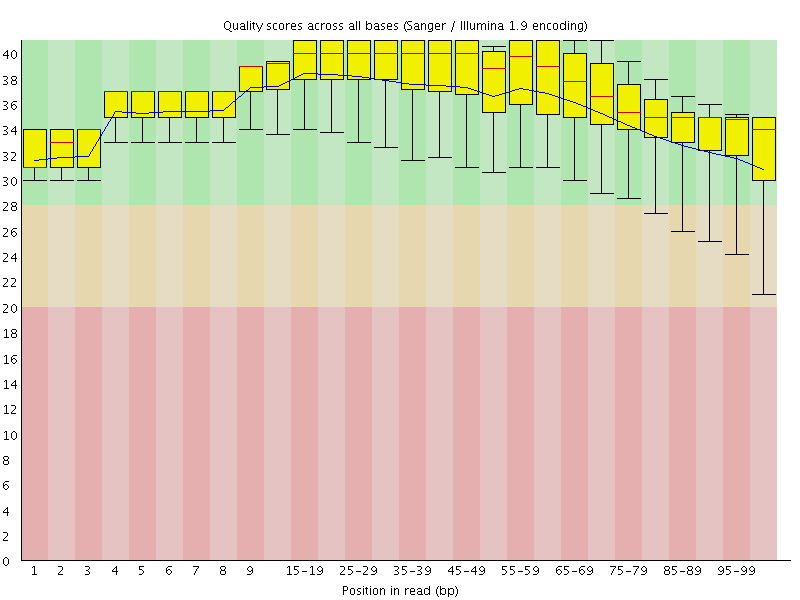
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
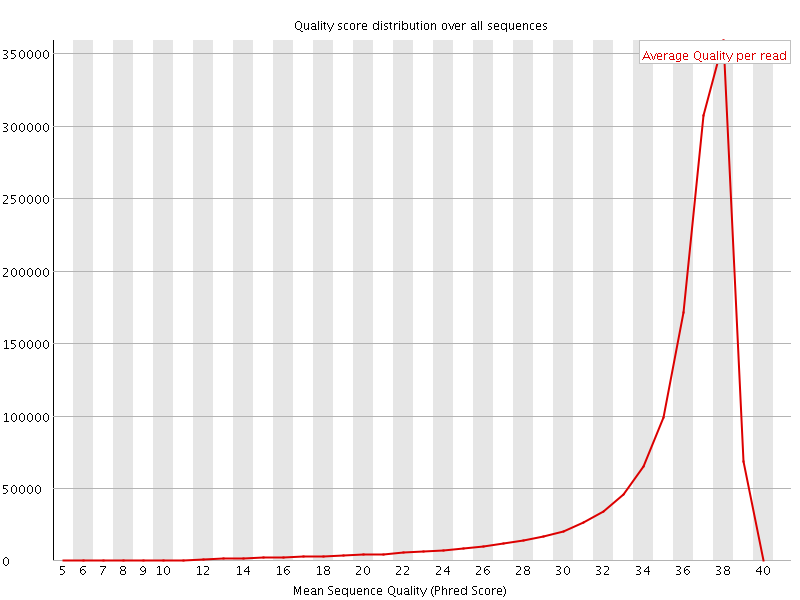
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
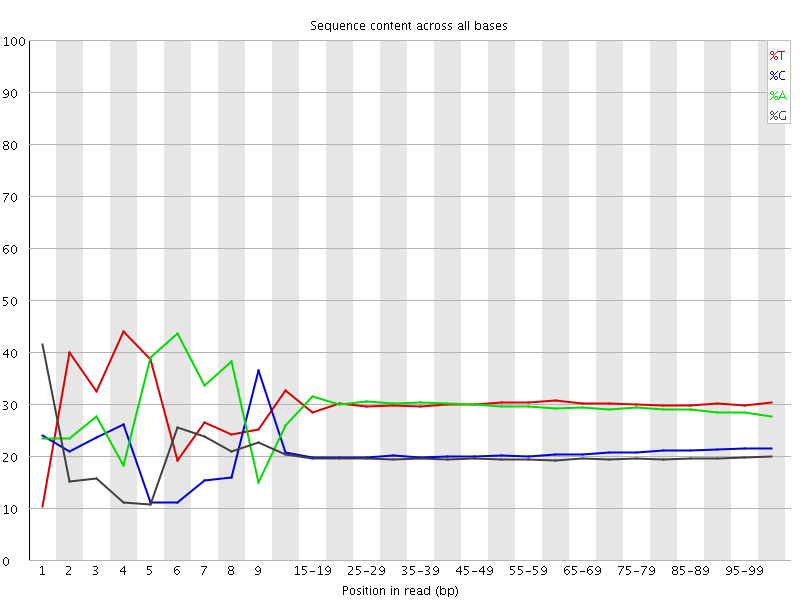
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
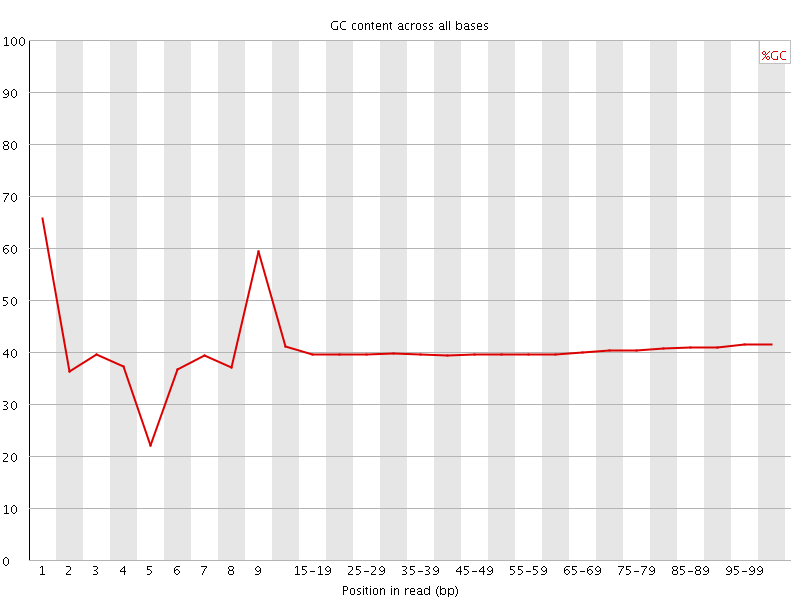
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
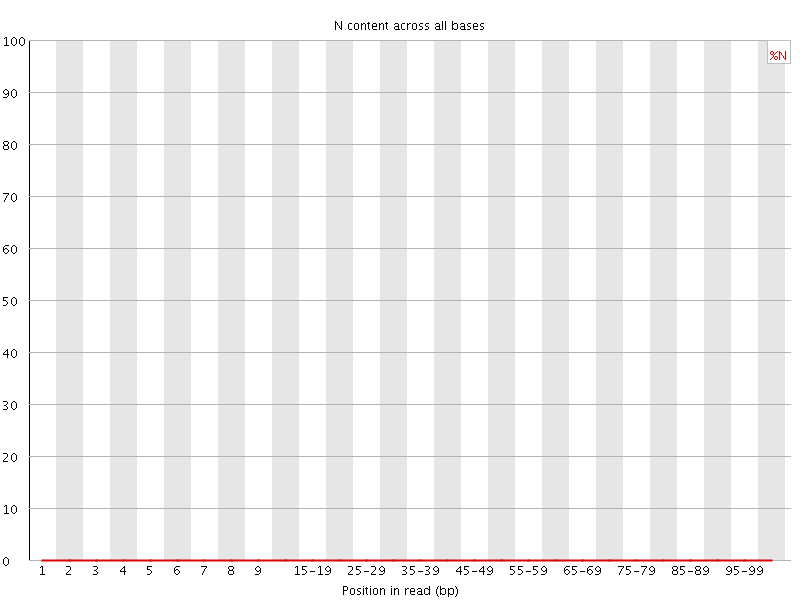
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
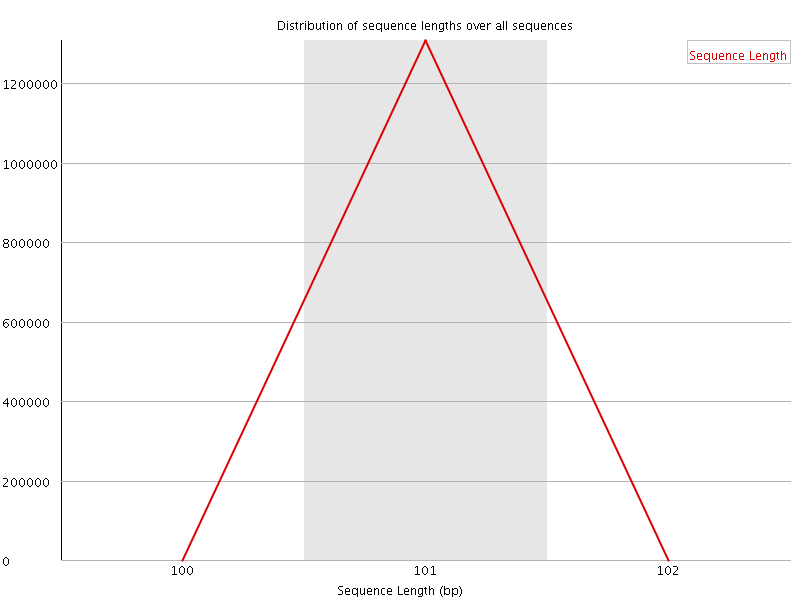
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
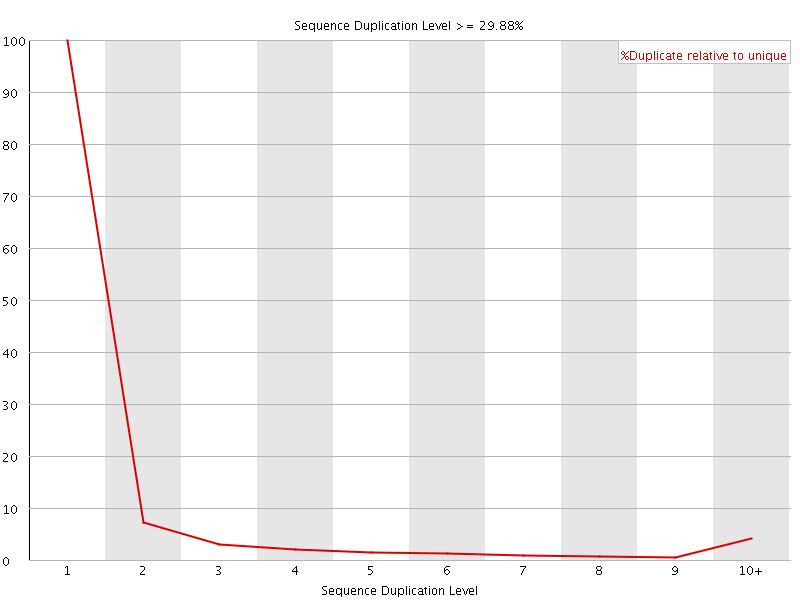
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
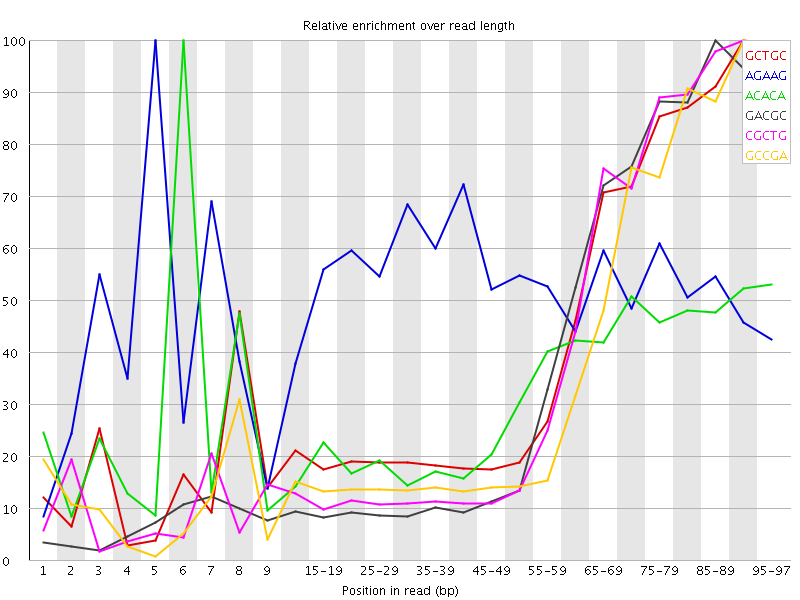
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 176410 | 2.779683 | 6.4880786 | 90-94 |
| AGAAG | 343610 | 2.690567 | 5.0479236 | 5 |
| ACACA | 334155 | 2.4218616 | 7.535515 | 6 |
| GACGC | 148500 | 2.3879006 | 6.0533633 | 85-89 |
| CGCTG | 149155 | 2.3502276 | 5.9026995 | 90-94 |
| GCCGA | 145845 | 2.345208 | 6.242135 | 90-94 |
| CTGCC | 152450 | 2.3110597 | 5.9641423 | 90-94 |
| CAAAG | 304410 | 2.2932353 | 5.0282617 | 4 |
| CGACG | 140015 | 2.2514603 | 6.460504 | 95-97 |
| GACTC | 209070 | 2.2116663 | 5.1659174 | 7 |
| TACAC | 299835 | 2.129443 | 9.411831 | 5 |
| TGCCG | 134200 | 2.114582 | 5.919511 | 90-94 |
| CTTTG | 296560 | 2.1020794 | 8.638185 | 9 |
| CCGAC | 135770 | 2.1004157 | 5.840346 | 95-97 |
| CACAT | 289485 | 2.0559366 | 5.3038898 | 7 |
| GCTTT | 254360 | 1.8029572 | 5.4464245 | 1 |
| ATACA | 357670 | 1.7725981 | 5.037393 | 6 |
| GAGAG | 144250 | 1.682434 | 5.508981 | 7 |
| TTGAG | 200530 | 1.5077244 | 5.290582 | 9 |
| TATAC | 305975 | 1.4859222 | 5.747906 | 5 |
| GAGTC | 132120 | 1.4527293 | 7.321164 | 9 |
| GTGTA | 191465 | 1.4395674 | 10.668127 | 1 |
| AAGAC | 186180 | 1.4025643 | 5.6641035 | 5 |
| CCATG | 130715 | 1.3827807 | 5.925204 | 9 |
| TATGA | 262485 | 1.3249605 | 6.1556435 | 4 |
| GTCTT | 182645 | 1.2946261 | 6.4951363 | 1 |
| GACTT | 177800 | 1.2861334 | 6.51062 | 7 |
| TCATA | 263375 | 1.2790416 | 6.405146 | 2 |
| TGAGT | 169900 | 1.2774267 | 5.498519 | 8 |
| GATTG | 164090 | 1.233743 | 5.943199 | 7 |
| AACAC | 169040 | 1.2251544 | 5.1786175 | 5 |
| GTGTT | 166045 | 1.2233499 | 5.8111906 | 1 |
| GTTCT | 164075 | 1.1629981 | 5.432671 | 1 |
| ACACC | 109060 | 1.1327205 | 5.479904 | 6 |
| ACCAT | 159350 | 1.1317115 | 5.152488 | 8 |
| TGGAC | 102215 | 1.123908 | 7.5151362 | 5 |
| ACATG | 151375 | 1.1174449 | 5.4307437 | 8 |
| GTCCA | 103785 | 1.0978992 | 7.748623 | 1 |
| GGACT | 96365 | 1.0595843 | 7.129095 | 6 |
| ACACT | 147730 | 1.0491858 | 5.2280803 | 6 |
| AGACT | 134955 | 0.99623317 | 5.298105 | 6 |
| GTATG | 131100 | 0.9857011 | 5.4596624 | 3 |
| GAGTA | 128150 | 0.9832838 | 5.136403 | 1 |
| CATAC | 130380 | 0.9259652 | 5.136446 | 3 |
| GTATA | 153305 | 0.7738463 | 7.5000663 | 1 |