![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-133_GGACTCCT-GTAAGGAG_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1307432 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
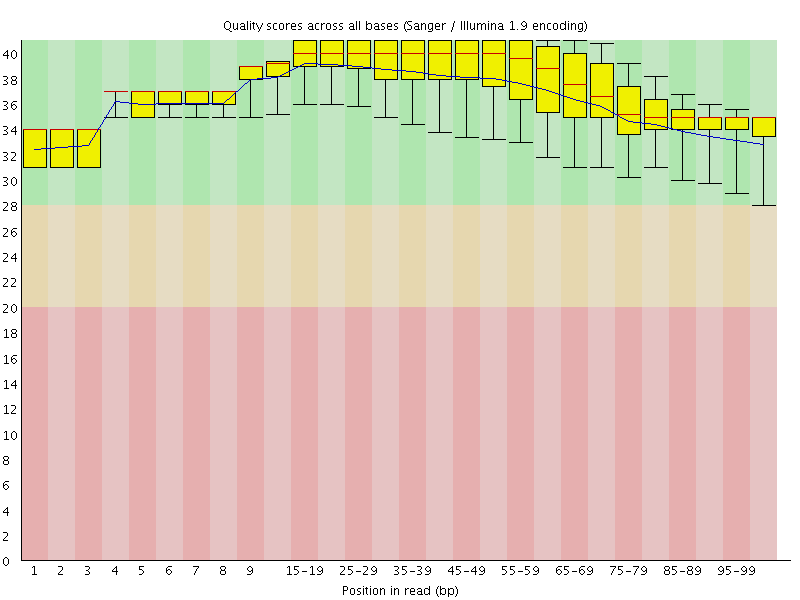
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
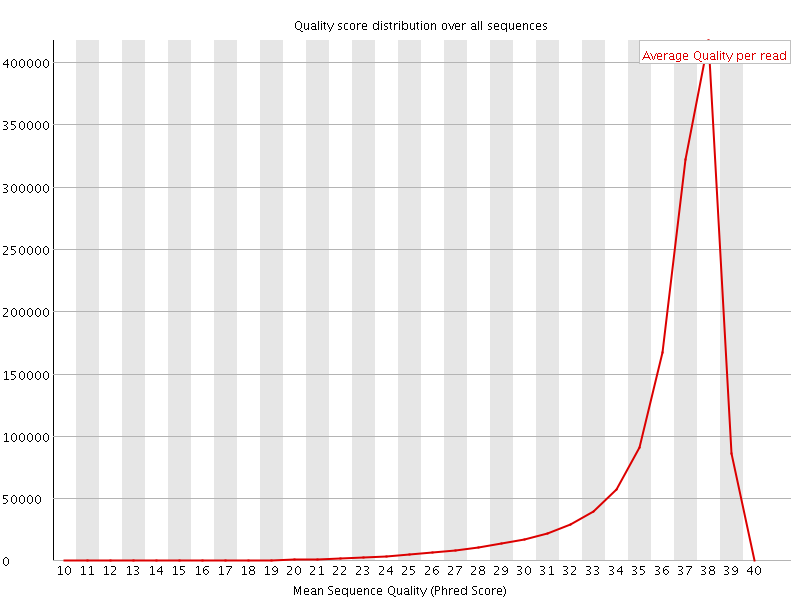
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
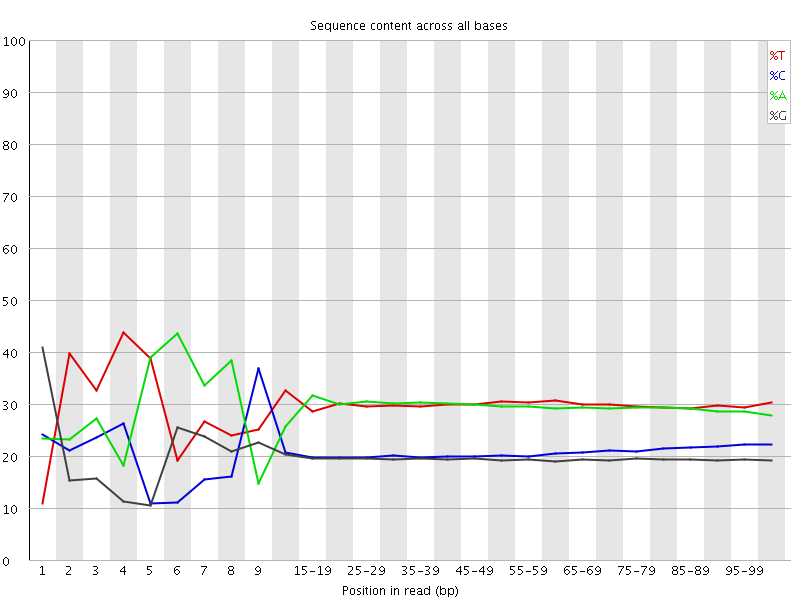
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
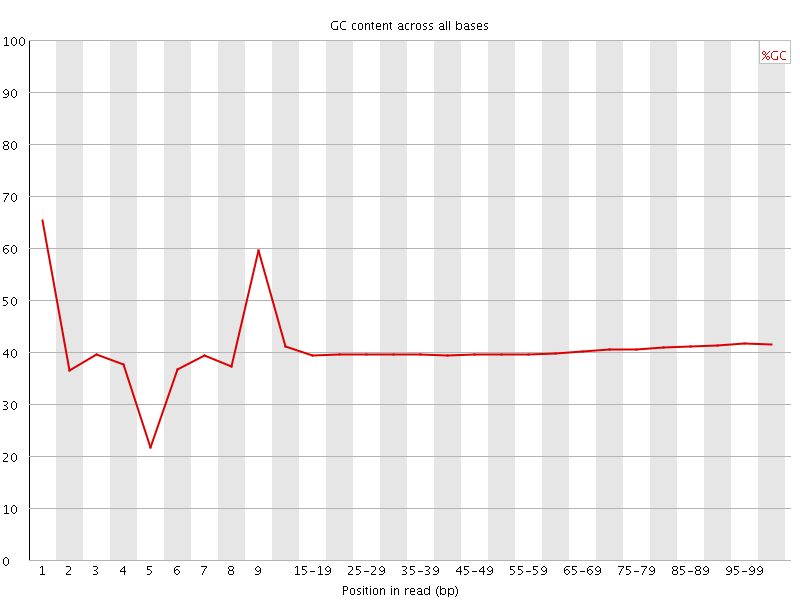
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
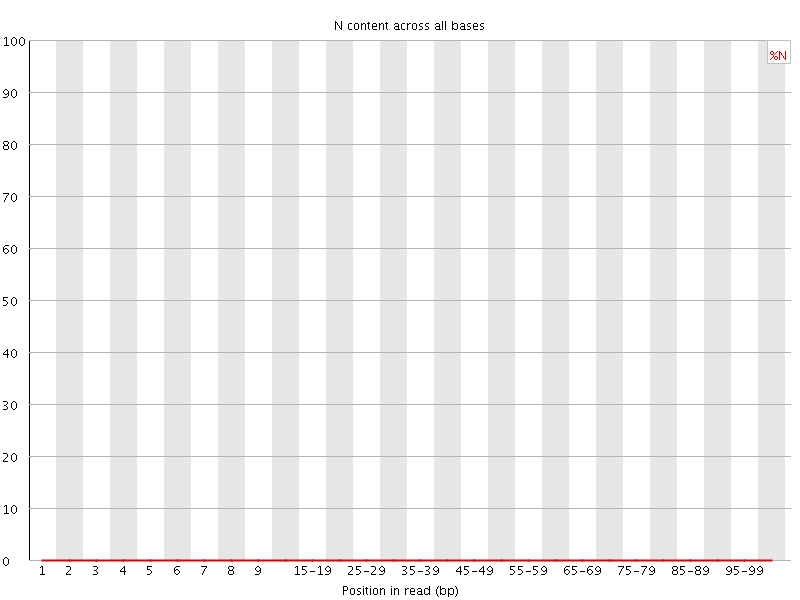
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
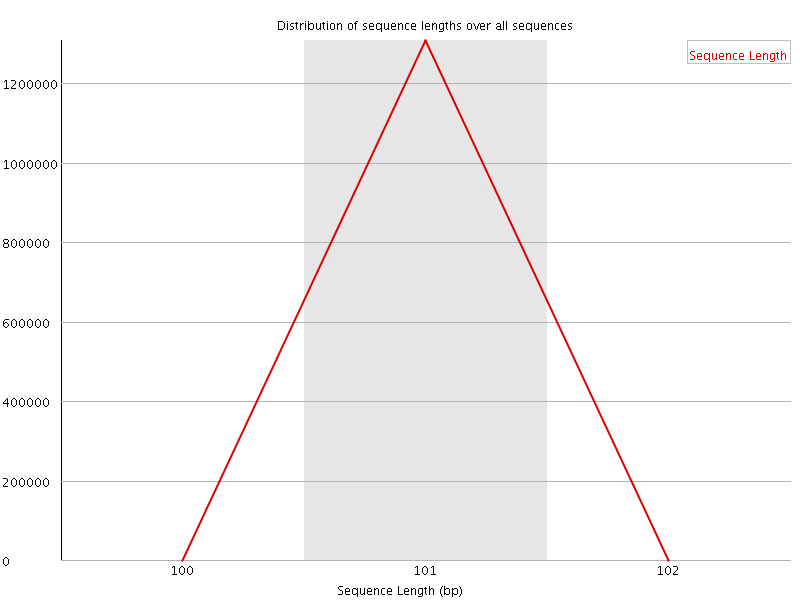
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
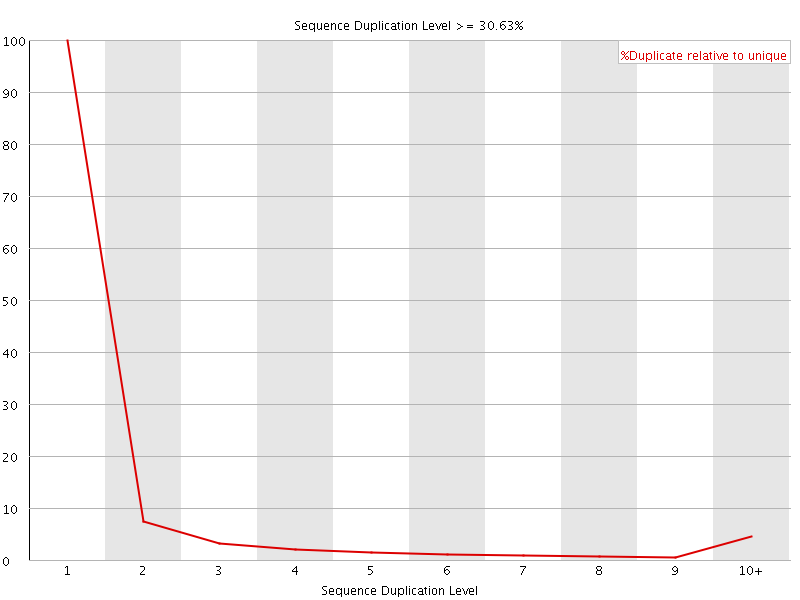
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
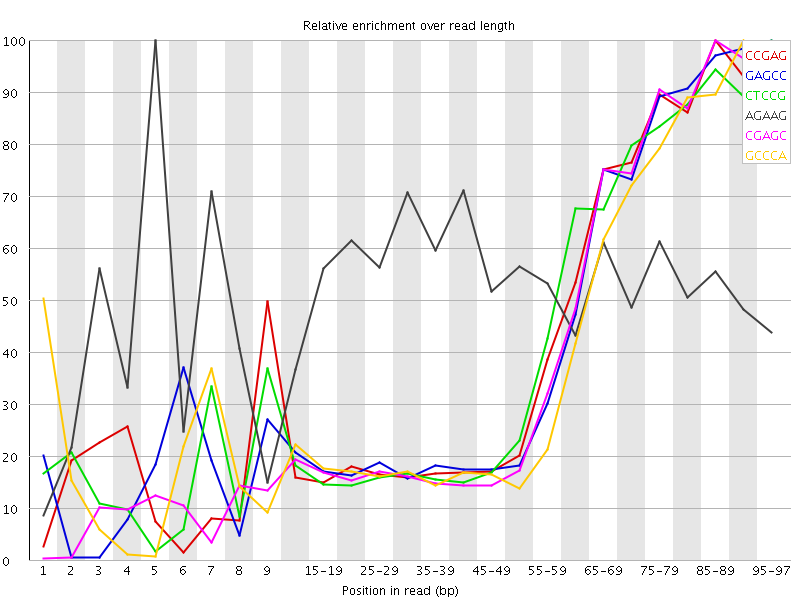
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 179405 | 2.8656812 | 6.5153575 | 85-89 |
| GAGCC | 175280 | 2.799791 | 6.3818736 | 95-97 |
| CTCCG | 184900 | 2.7581954 | 6.2661514 | 95-97 |
| AGAAG | 346105 | 2.7307184 | 5.068577 | 5 |
| CGAGC | 167185 | 2.6704879 | 6.2690563 | 85-89 |
| GCCCA | 164675 | 2.4952838 | 6.0850973 | 90-94 |
| AGCCC | 164100 | 2.486571 | 5.988112 | 90-94 |
| ACACA | 332580 | 2.3613648 | 6.898677 | 6 |
| ATCTC | 329745 | 2.2690125 | 5.1303687 | 95-97 |
| CCCAC | 155115 | 2.2296932 | 5.633135 | 90-94 |
| CTTTG | 297350 | 2.1233592 | 8.617505 | 9 |
| CCACG | 139285 | 2.1105547 | 5.716259 | 90-94 |
| TACAC | 301805 | 2.1095471 | 9.058638 | 5 |
| GACTC | 200585 | 2.1029282 | 5.4083996 | 7 |
| GACGG | 121300 | 2.0424676 | 5.9490957 | 95-97 |
| CGGAC | 114290 | 1.8255829 | 5.509944 | 90-94 |
| GGACT | 158870 | 1.7557752 | 7.260515 | 6 |
| GAGAG | 143810 | 1.7018498 | 5.531085 | 7 |
| TGGCT | 147455 | 1.6042882 | 5.1695185 | 6 |
| TTGAG | 202935 | 1.5517354 | 5.460905 | 9 |
| TATAC | 305670 | 1.4782549 | 5.826719 | 5 |
| GAGTC | 132790 | 1.4675481 | 7.399831 | 9 |
| AAGAC | 185955 | 1.3917968 | 5.733585 | 5 |
| CCATG | 131820 | 1.3819977 | 5.865877 | 9 |
| GTCTT | 192120 | 1.3719178 | 6.2395697 | 1 |
| TATGA | 261940 | 1.3353629 | 6.3251224 | 4 |
| TGAGT | 171530 | 1.3115982 | 5.3533926 | 8 |
| GACTT | 178780 | 1.2968168 | 6.7137747 | 7 |
| TCATA | 265350 | 1.283263 | 6.216567 | 2 |
| GATTG | 166185 | 1.2707278 | 6.1690063 | 7 |
| GTGTT | 168590 | 1.2690783 | 5.975161 | 1 |
| GTATG | 158310 | 1.2105119 | 5.735248 | 3 |
| GTGTA | 156250 | 1.1947602 | 10.896941 | 1 |
| GTTCT | 164985 | 1.1781484 | 5.270047 | 1 |
| TGGAC | 101565 | 1.1224606 | 7.3462486 | 5 |
| ACATG | 152035 | 1.1202306 | 5.315791 | 8 |
| ACACC | 109905 | 1.1103159 | 5.231202 | 6 |
| GTCCA | 104960 | 1.100398 | 7.803307 | 1 |
| ACACT | 147135 | 1.0284396 | 5.0529103 | 6 |
| TGTAT | 203695 | 1.0222893 | 5.16883 | 2 |
| GAGTA | 129690 | 1.007329 | 5.408335 | 1 |
| AGACT | 135700 | 0.99987036 | 5.5051303 | 6 |
| GTATA | 155225 | 0.7913329 | 7.2008266 | 1 |