![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-131_GGACTCCT-TATCCTCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1727814 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
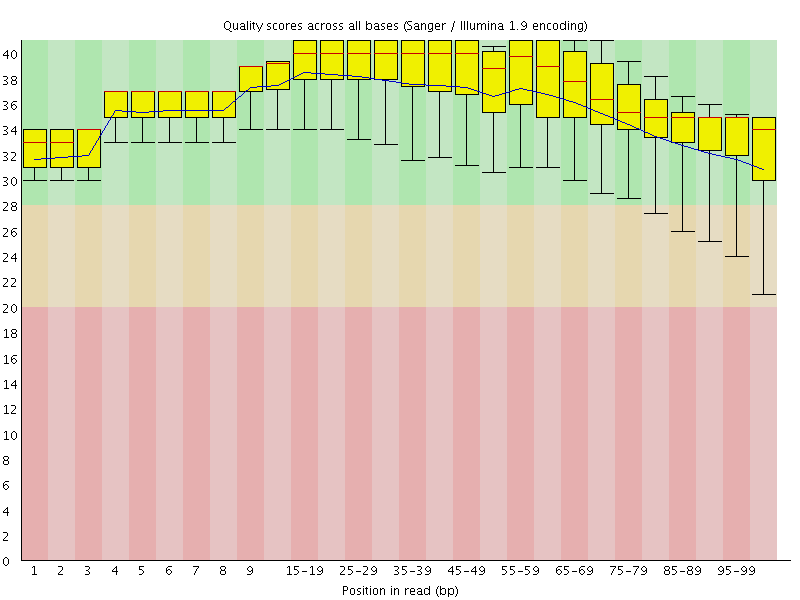
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
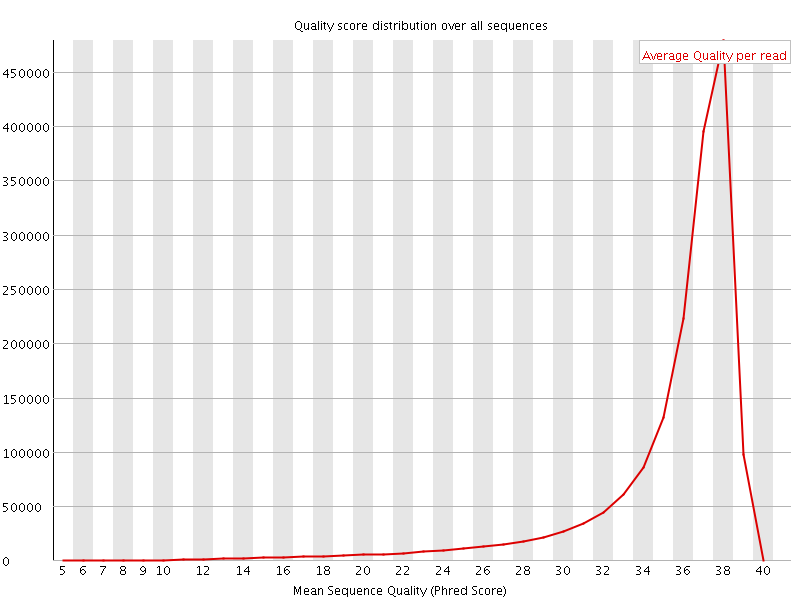
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
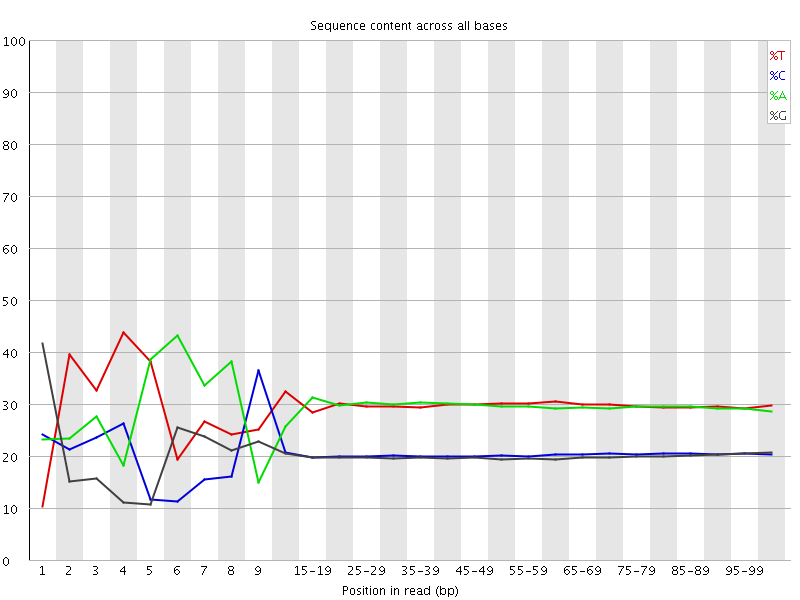
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
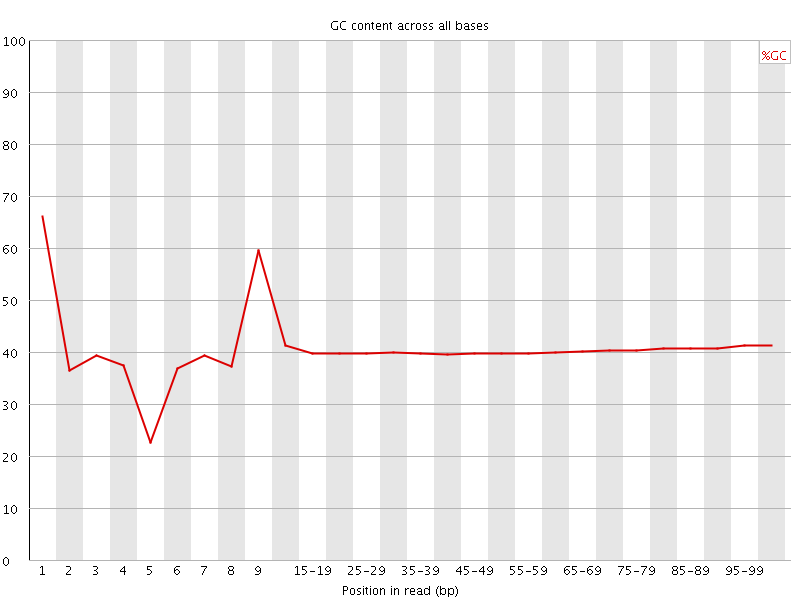
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
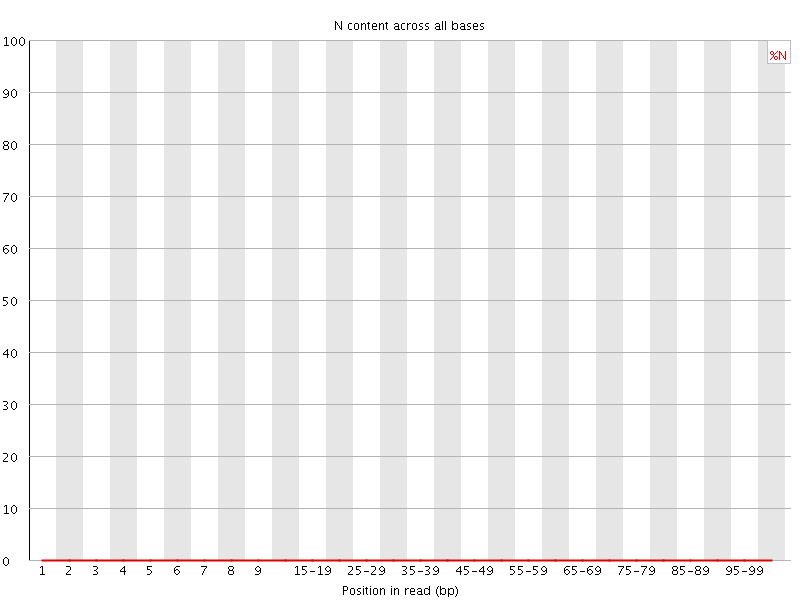
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
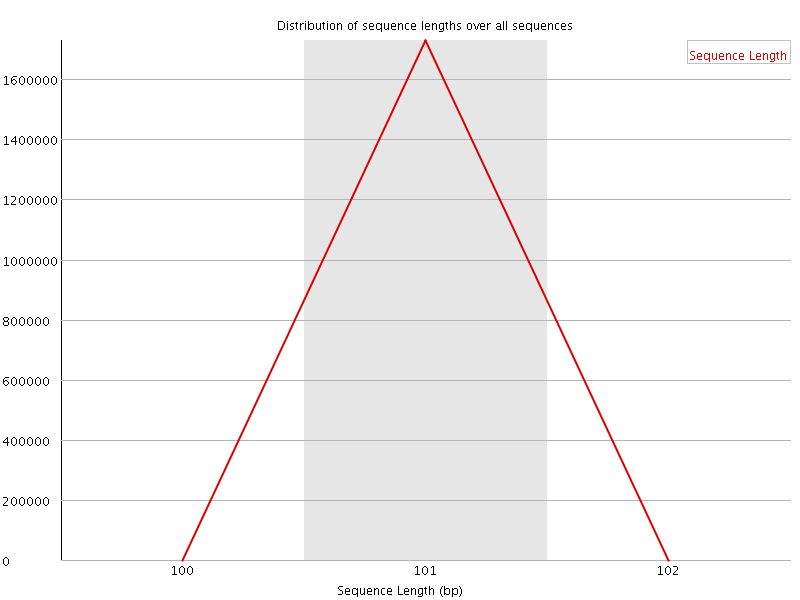
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
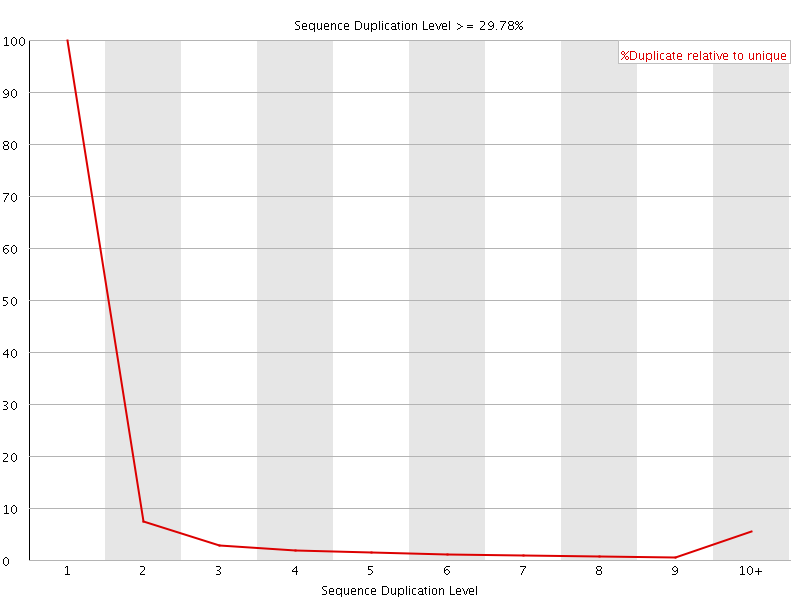
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
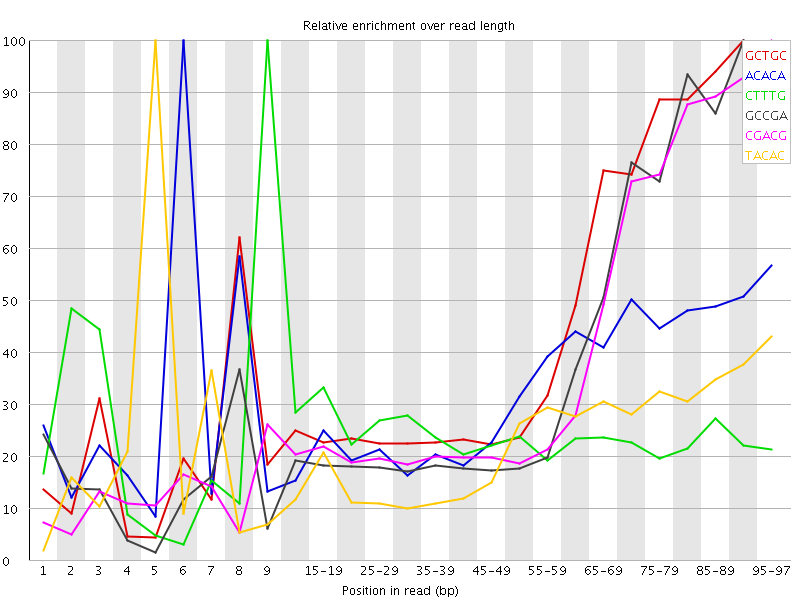
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 208675 | 2.4857242 | 5.3346105 | 90-94 |
| ACACA | 406875 | 2.2430923 | 6.7318196 | 6 |
| CTTTG | 397085 | 2.1647546 | 8.856801 | 9 |
| GCCGA | 177390 | 2.1326017 | 5.280163 | 90-94 |
| CGACG | 172370 | 2.0722508 | 5.1779003 | 95-97 |
| TACAC | 363205 | 1.9839917 | 8.569256 | 5 |
| GCTTT | 339945 | 1.853249 | 5.484729 | 1 |
| ATACA | 435230 | 1.6366012 | 5.3484344 | 6 |
| GAGAG | 193110 | 1.6246302 | 5.0754495 | 7 |
| GACTC | 183050 | 1.4765735 | 5.1715555 | 7 |
| GAGTC | 174095 | 1.4275943 | 7.8381224 | 9 |
| CCATG | 172815 | 1.3940129 | 5.433686 | 9 |
| AAGAC | 248240 | 1.391205 | 5.4044404 | 5 |
| TATAC | 365750 | 1.3627313 | 5.92809 | 5 |
| GTGTA | 240670 | 1.3461032 | 9.756575 | 1 |
| GTCTT | 244805 | 1.3345826 | 6.2860184 | 1 |
| TCATA | 351745 | 1.310551 | 5.6533422 | 2 |
| GACTT | 235460 | 1.2955089 | 6.473191 | 7 |
| TATGA | 341590 | 1.2937914 | 5.6716676 | 4 |
| TGAGT | 227055 | 1.2699524 | 6.02719 | 8 |
| AACAC | 225530 | 1.2433416 | 5.016884 | 5 |
| GTGTT | 223005 | 1.2358704 | 5.9572988 | 1 |
| ATGAG | 216545 | 1.2223698 | 5.048005 | 7 |
| GATTG | 217455 | 1.2162583 | 5.4764247 | 7 |
| GTTCT | 219585 | 1.1970927 | 5.289035 | 1 |
| TGGAC | 138025 | 1.1318172 | 7.120108 | 5 |
| GTCCA | 138890 | 1.1203567 | 8.201588 | 1 |
| ACACT | 200535 | 1.095414 | 5.5072293 | 6 |
| GGACT | 127600 | 1.0463312 | 7.14988 | 6 |
| AGACT | 179575 | 0.9971653 | 5.2968326 | 6 |
| GTATA | 203140 | 0.7694043 | 7.240806 | 1 |