![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-131_GGACTCCT-TATCCTCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1727814 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
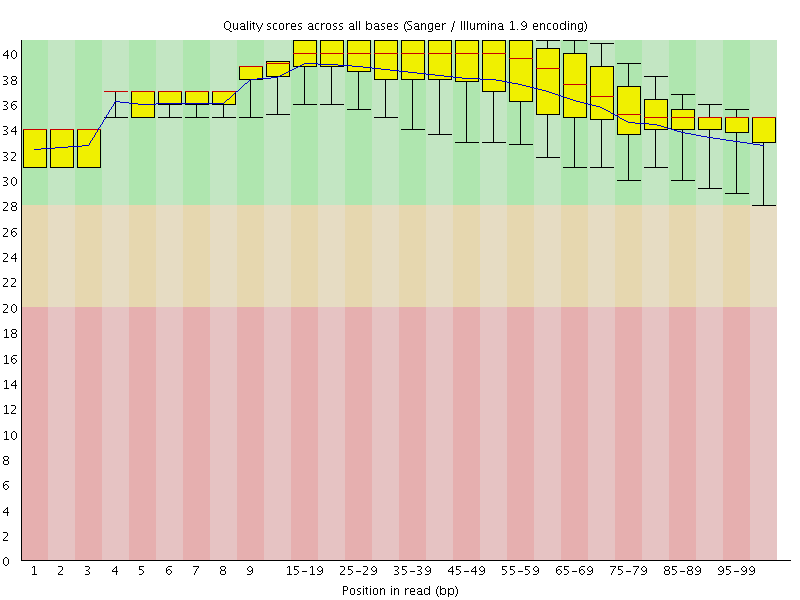
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
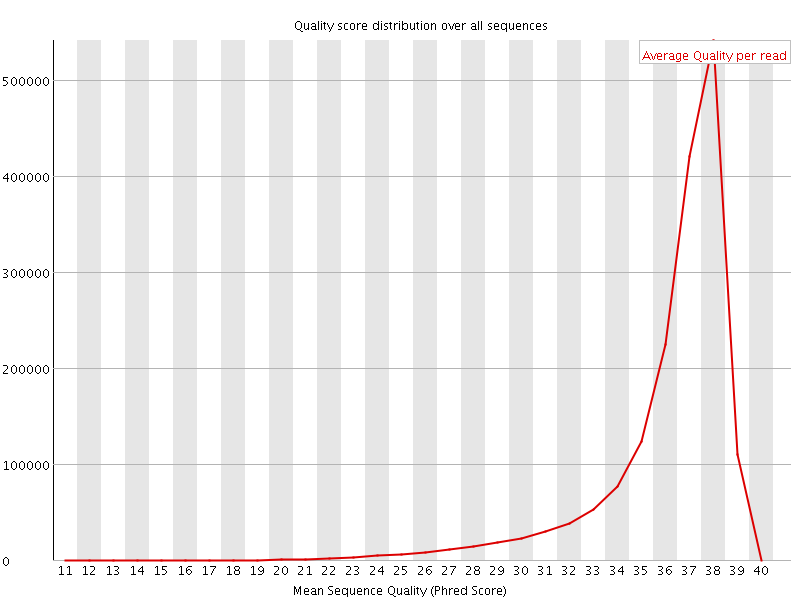
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
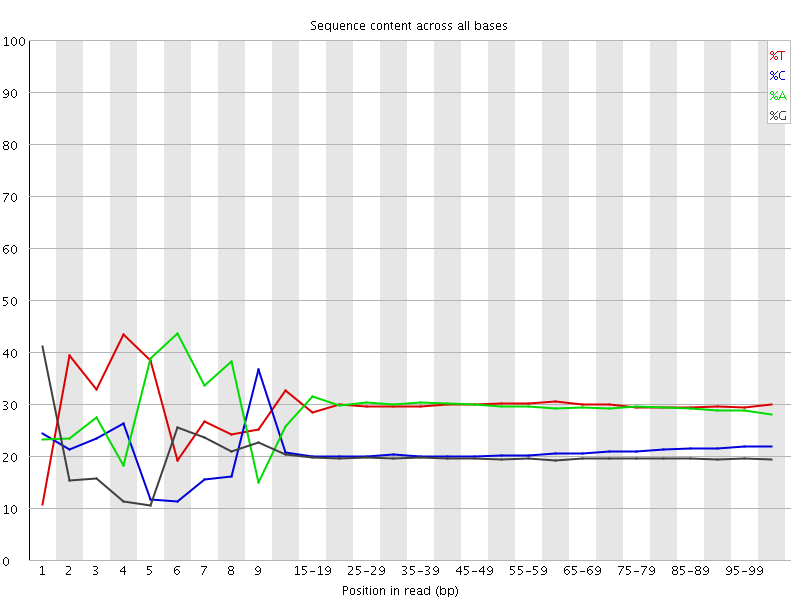
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
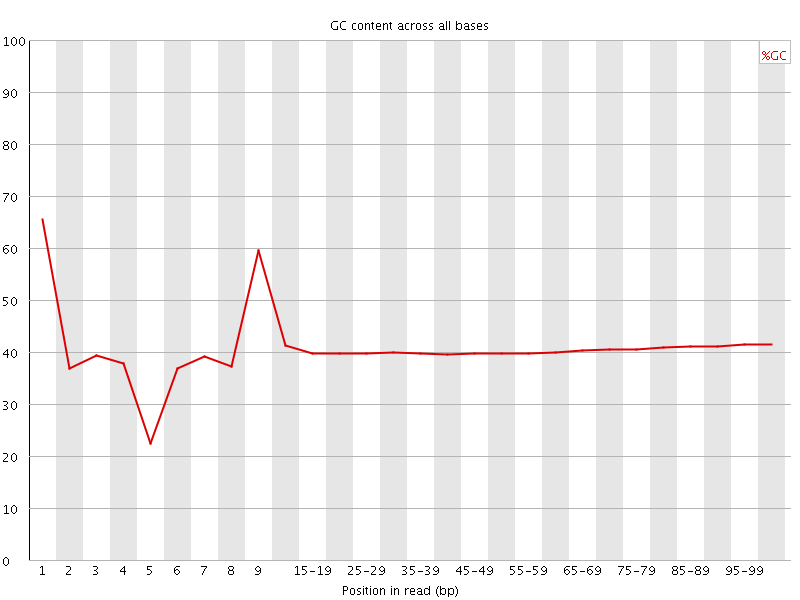
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
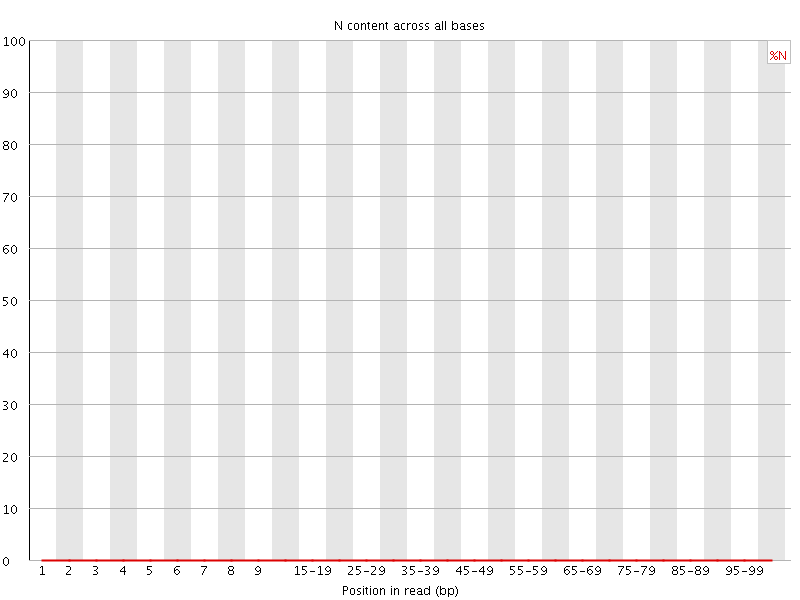
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
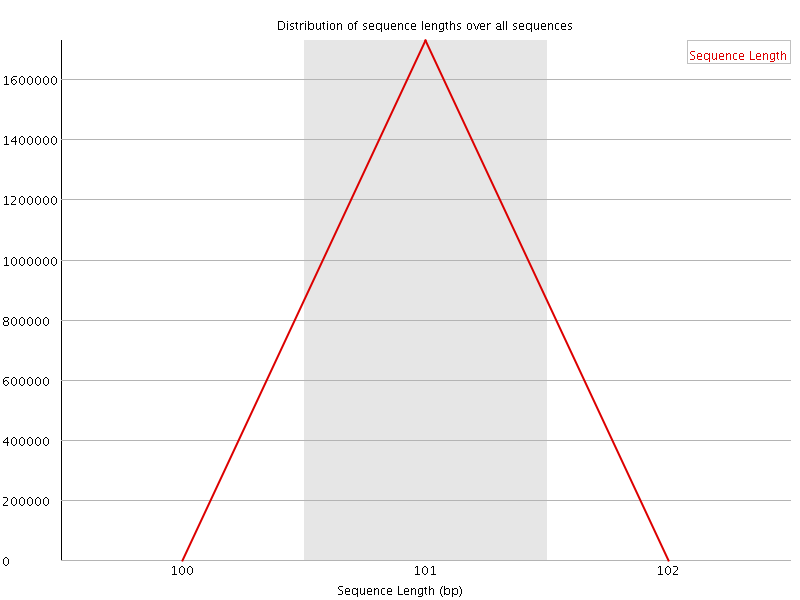
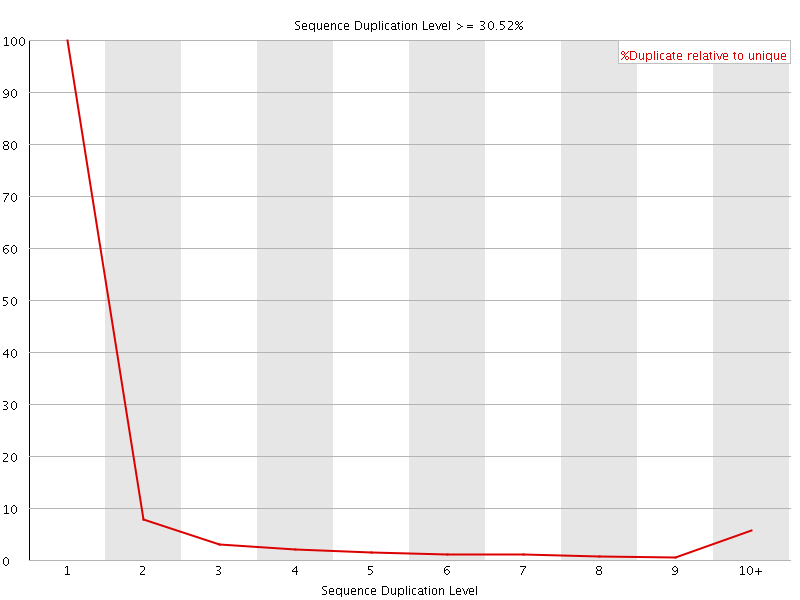
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
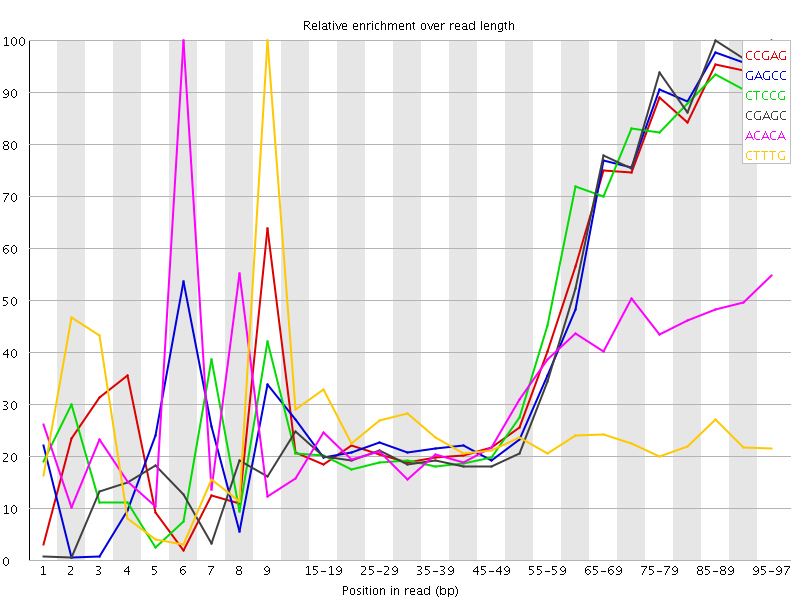
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 213495 | 2.5518527 | 5.5154796 | 95-97 |
| GAGCC | 206445 | 2.4675858 | 5.302974 | 95-97 |
| CTCCG | 215230 | 2.4330235 | 5.238181 | 95-97 |
| CGAGC | 196180 | 2.3448908 | 5.1836705 | 85-89 |
| ACACA | 406955 | 2.1892195 | 6.6719995 | 6 |
| CTTTG | 396380 | 2.1559308 | 8.802822 | 9 |
| TACAC | 364205 | 1.9374378 | 8.717884 | 5 |
| GCTTT | 338040 | 1.8386166 | 5.343556 | 1 |
| GACGG | 146545 | 1.8314804 | 5.080174 | 95-97 |
| GAGAG | 191415 | 1.6780623 | 5.189916 | 7 |
| GGACT | 197680 | 1.6389679 | 7.2037435 | 6 |
| ATACA | 434680 | 1.6220056 | 5.433136 | 6 |
| TTGAG | 270055 | 1.5531011 | 5.28129 | 9 |
| GAGTC | 174625 | 1.4478185 | 7.8630147 | 9 |
| CCATG | 177790 | 1.409781 | 5.4440246 | 9 |
| AAGAC | 250450 | 1.4087285 | 5.664418 | 5 |
| GTCTT | 253835 | 1.3806213 | 6.2350273 | 1 |
| TATAC | 368085 | 1.3582188 | 5.927282 | 5 |
| TATGA | 344745 | 1.330096 | 5.7541857 | 4 |
| GACTT | 238840 | 1.3136847 | 6.47775 | 7 |
| TGAGT | 228330 | 1.3131381 | 6.198683 | 8 |
| TCATA | 351730 | 1.2978694 | 5.550489 | 2 |
| GTGTT | 225930 | 1.2848734 | 6.0780725 | 1 |
| GATTG | 219980 | 1.2651172 | 5.8501306 | 7 |
| ATGAG | 216870 | 1.2612698 | 5.27306 | 7 |
| GTGTA | 207565 | 1.1937176 | 10.1902275 | 1 |
| GTTCT | 219280 | 1.192675 | 5.187944 | 1 |
| TGGAC | 140450 | 1.164473 | 7.2278633 | 5 |
| GTATG | 201785 | 1.1604766 | 5.295232 | 3 |
| GTCCA | 141010 | 1.118135 | 7.7940993 | 1 |
| ACACT | 198890 | 1.0580223 | 5.50929 | 6 |
| AGACT | 178880 | 0.99496317 | 5.272353 | 6 |
| GAGTA | 170190 | 0.9897889 | 5.039656 | 1 |
| GGATA | 152015 | 0.8840869 | 5.0283756 | 1 |
| GTATA | 201595 | 0.7777943 | 7.229195 | 1 |