![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-130_GGACTCCT-CTCTCTAT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1174790 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
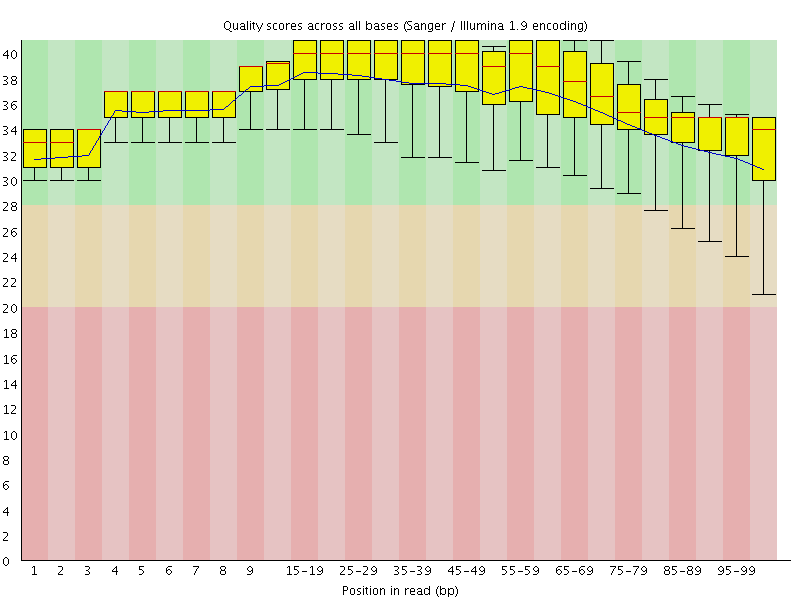
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
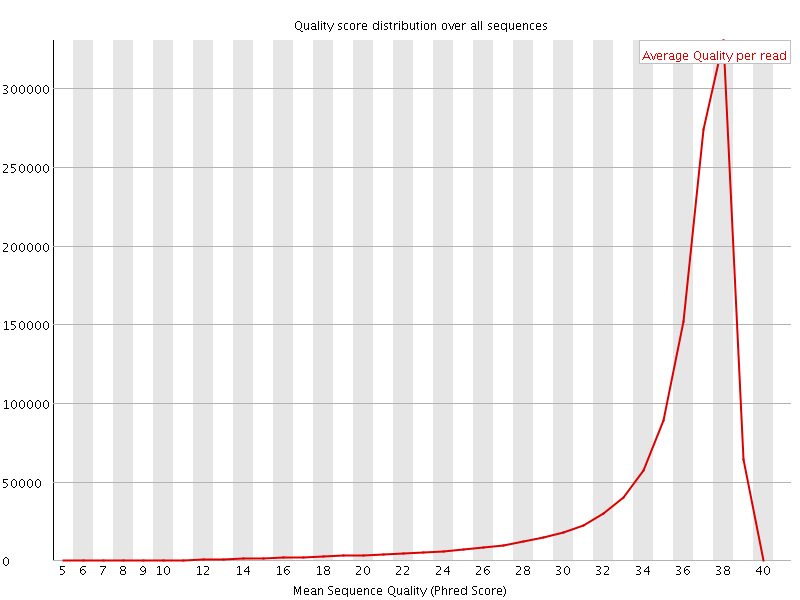
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
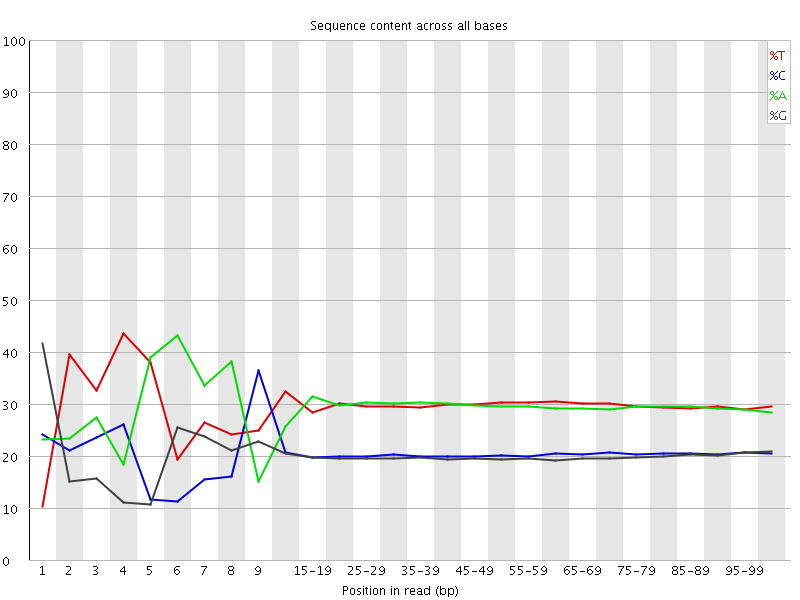
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
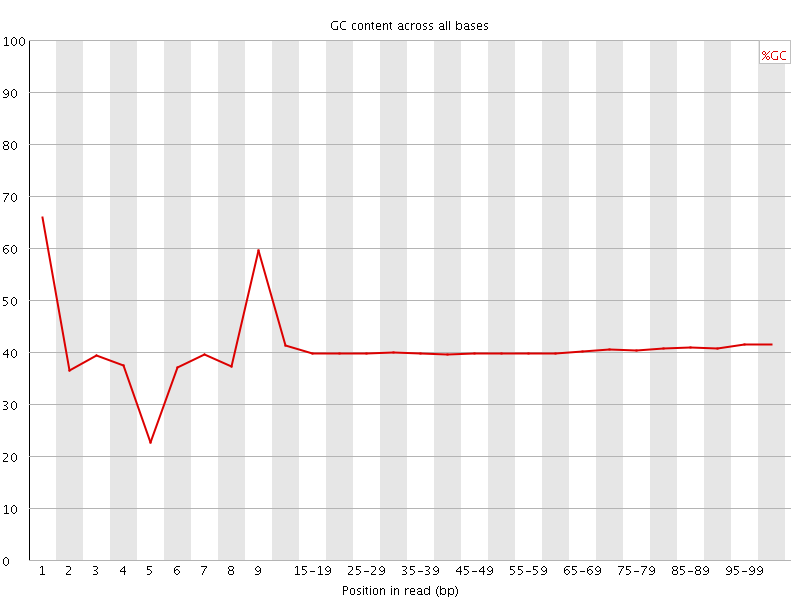
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
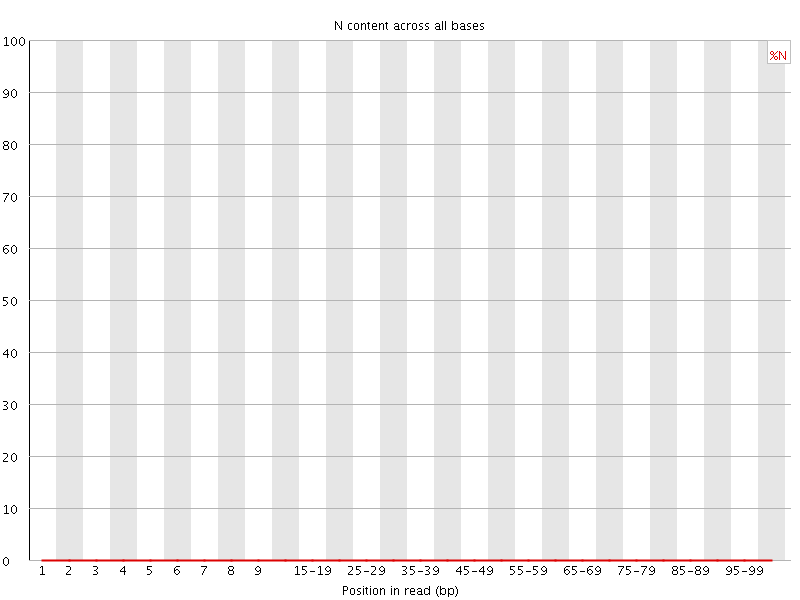
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
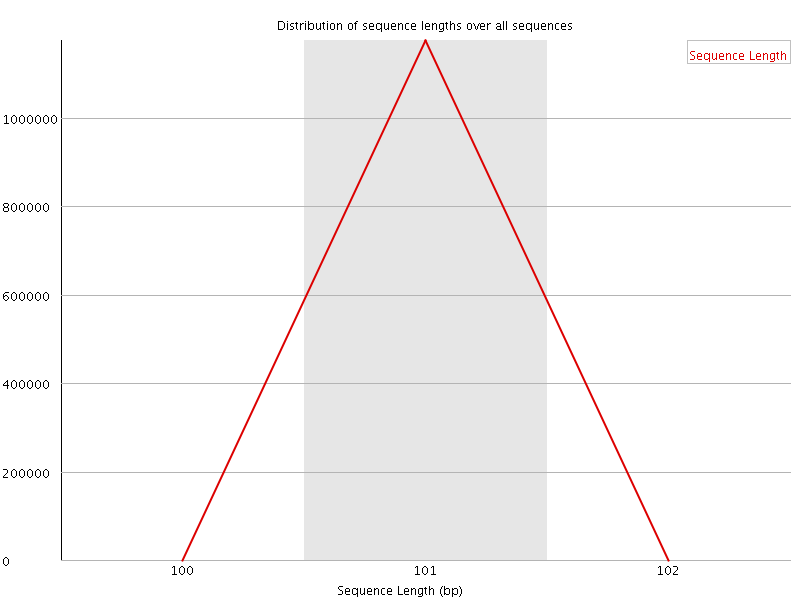
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
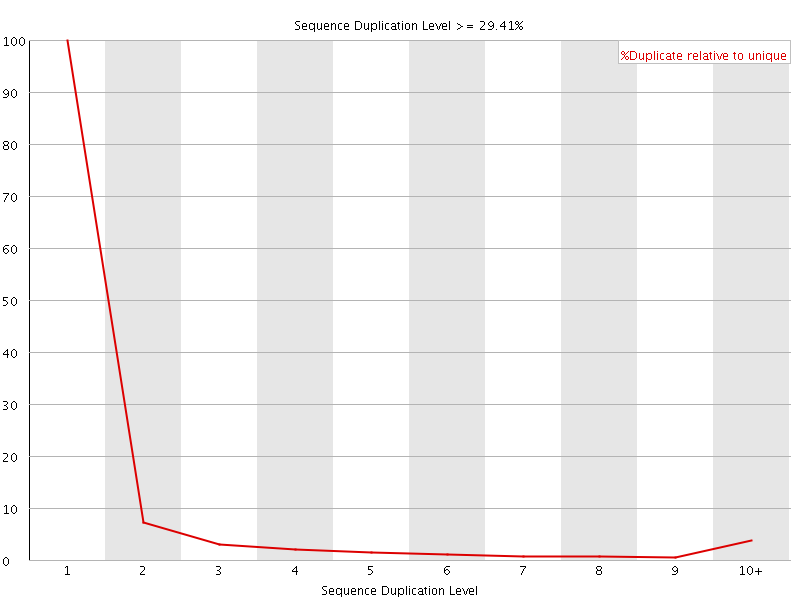
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
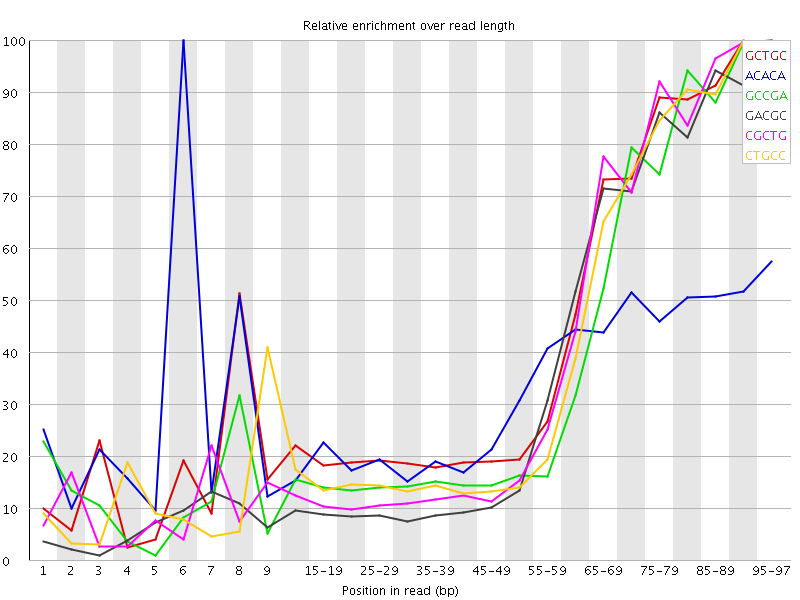
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 156945 | 2.7390816 | 6.2598114 | 90-94 |
| ACACA | 302080 | 2.4447677 | 7.347153 | 6 |
| GCCGA | 133535 | 2.356468 | 6.0880027 | 95-97 |
| GACGC | 133455 | 2.355056 | 6.2049685 | 95-97 |
| CGCTG | 134045 | 2.3394198 | 5.8686047 | 95-97 |
| CTGCC | 136140 | 2.326165 | 5.8484836 | 90-94 |
| GAGAG | 182130 | 2.270795 | 5.0367045 | 7 |
| CGACG | 128095 | 2.2604692 | 6.2648787 | 95-97 |
| CTTTG | 275835 | 2.2056727 | 8.581795 | 9 |
| TACAC | 272915 | 2.18441 | 8.871099 | 5 |
| CCGAC | 124935 | 2.1584787 | 5.8542175 | 95-97 |
| TGCCG | 120765 | 2.1076503 | 5.7875323 | 90-94 |
| CACAT | 260105 | 2.081879 | 5.197474 | 7 |
| GCTTT | 233380 | 1.8661878 | 5.3758917 | 1 |
| ATACA | 322150 | 1.7835786 | 5.222174 | 6 |
| GACTC | 131245 | 1.5511895 | 5.0840607 | 7 |
| TTGAG | 184835 | 1.5264673 | 5.1344786 | 9 |
| TATAC | 273660 | 1.4984304 | 5.5083117 | 5 |
| GAGTC | 123900 | 1.4957402 | 8.658807 | 9 |
| GTGTA | 171915 | 1.419767 | 10.102852 | 1 |
| CCATG | 119010 | 1.4065837 | 5.9037004 | 9 |
| AAGAC | 170070 | 1.4058728 | 5.2846746 | 5 |
| GTCTT | 168555 | 1.3478246 | 6.376599 | 1 |
| TATGA | 240270 | 1.3437777 | 6.0006394 | 4 |
| TCATA | 241375 | 1.3216531 | 6.0767226 | 2 |
| GACTT | 162705 | 1.3155322 | 6.814835 | 7 |
| TGAGT | 158695 | 1.3105891 | 6.5723486 | 8 |
| GTGTT | 153415 | 1.2530321 | 5.7802324 | 1 |
| ATGAG | 148740 | 1.2420526 | 5.260492 | 7 |
| GATTG | 147605 | 1.2190019 | 5.538989 | 7 |
| GTTCT | 149275 | 1.1936548 | 5.313832 | 1 |
| ACATG | 138030 | 1.1284516 | 5.4079485 | 8 |
| TGGAC | 92810 | 1.1204168 | 7.0969534 | 5 |
| GTCCA | 93800 | 1.1086258 | 8.060519 | 1 |
| ATGTG | 132495 | 1.0942154 | 5.066393 | 7 |
| ACACT | 135580 | 1.0851815 | 5.1973853 | 6 |
| GGACT | 87045 | 1.0508208 | 7.2828746 | 6 |
| AGACT | 123430 | 1.0090907 | 5.0668488 | 6 |
| GTATG | 121990 | 1.0074594 | 5.3556914 | 3 |
| GAGTA | 118160 | 0.9866944 | 5.046913 | 1 |
| GTATA | 139400 | 0.7796338 | 7.498278 | 1 |