![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-130_GGACTCCT-CTCTCTAT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1174790 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
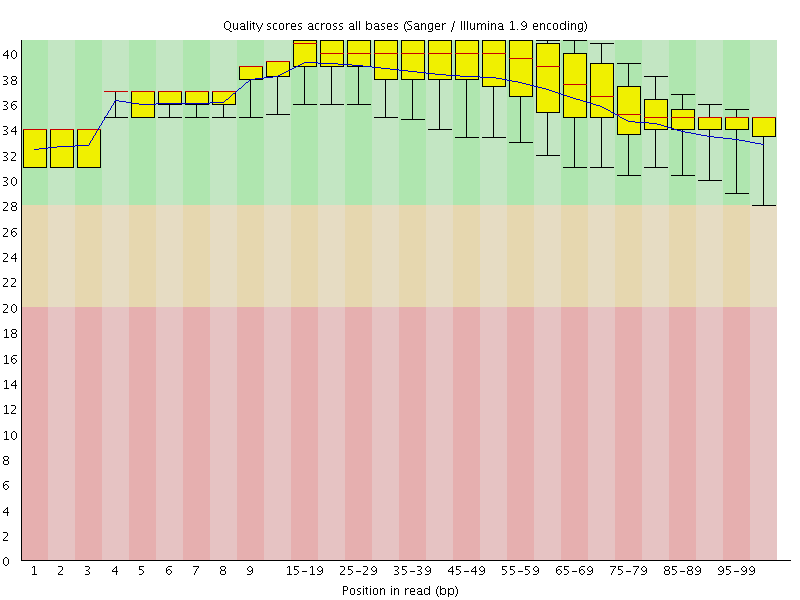
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
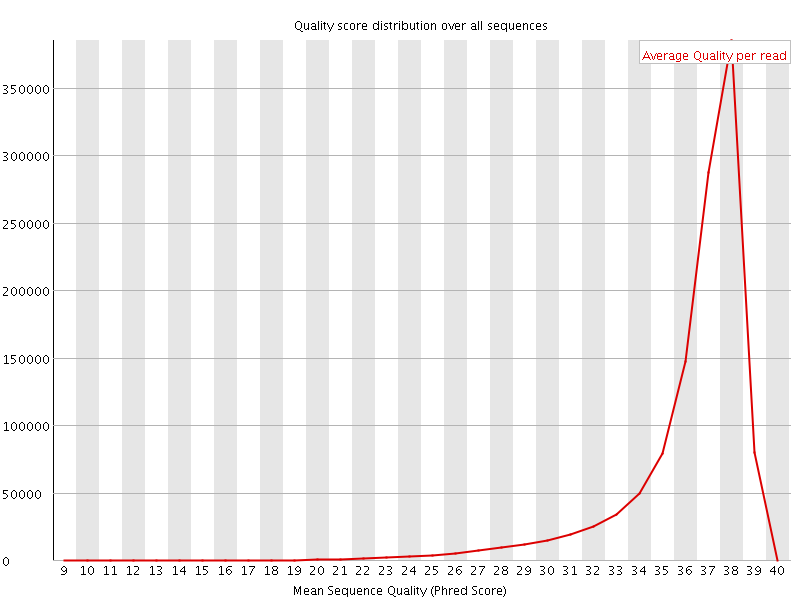
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
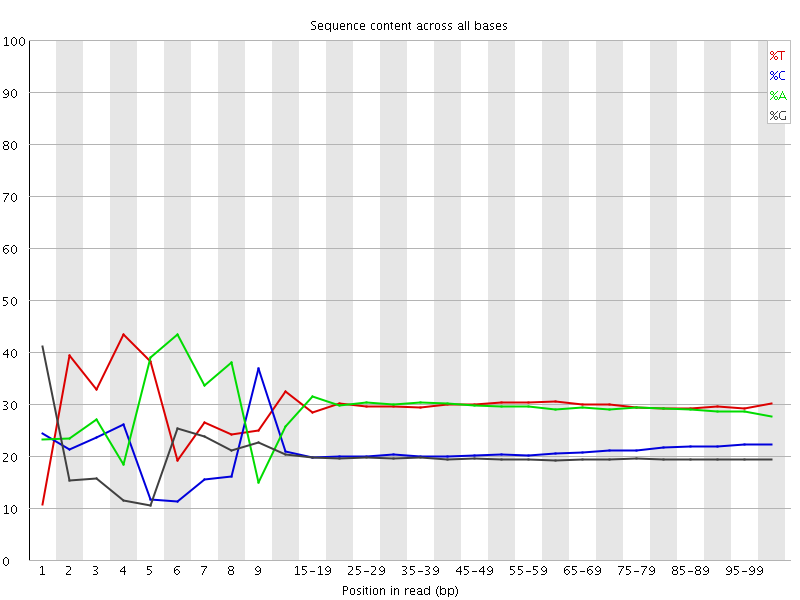
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
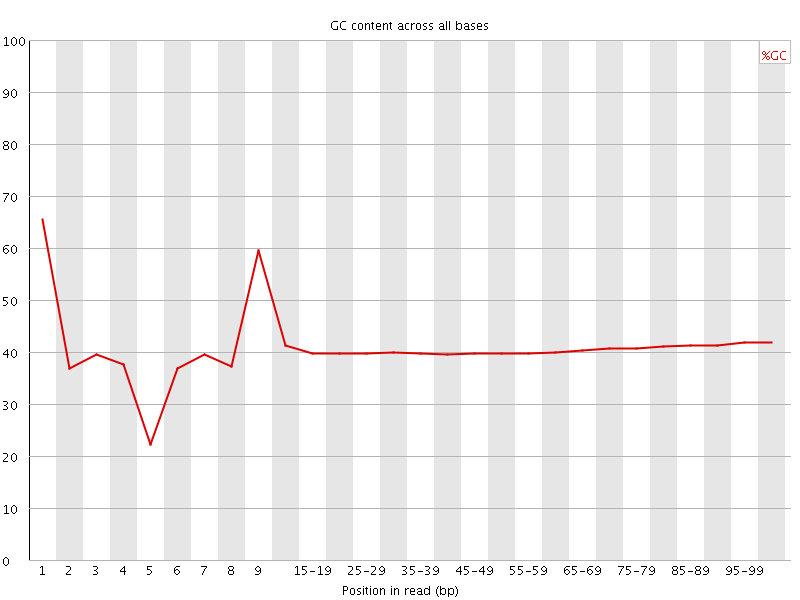
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
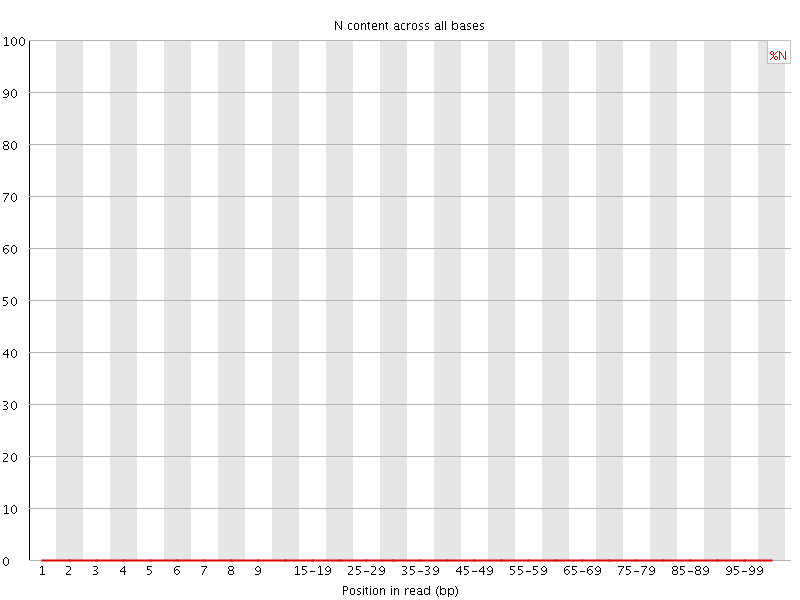
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
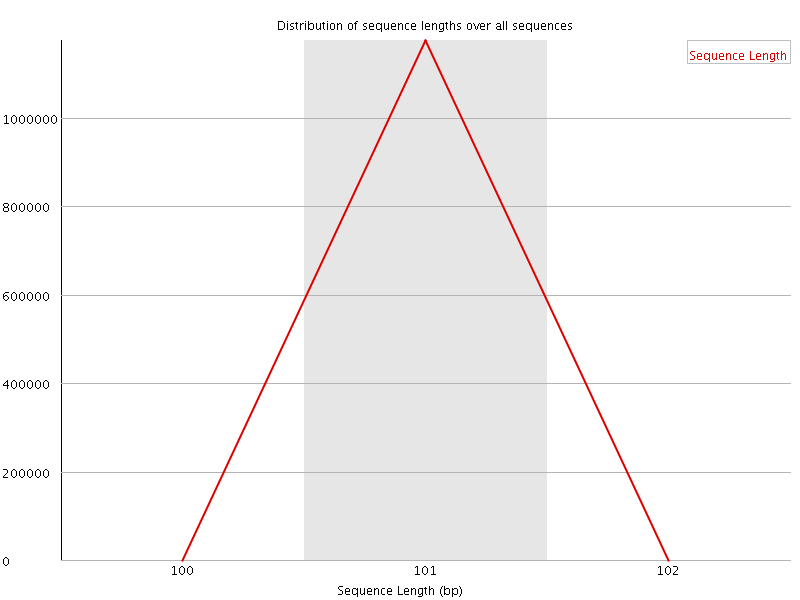
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
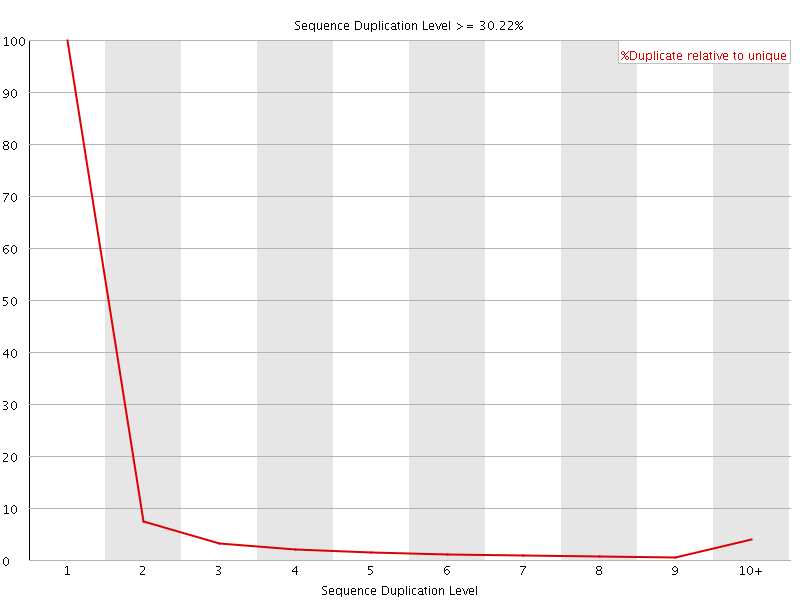
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
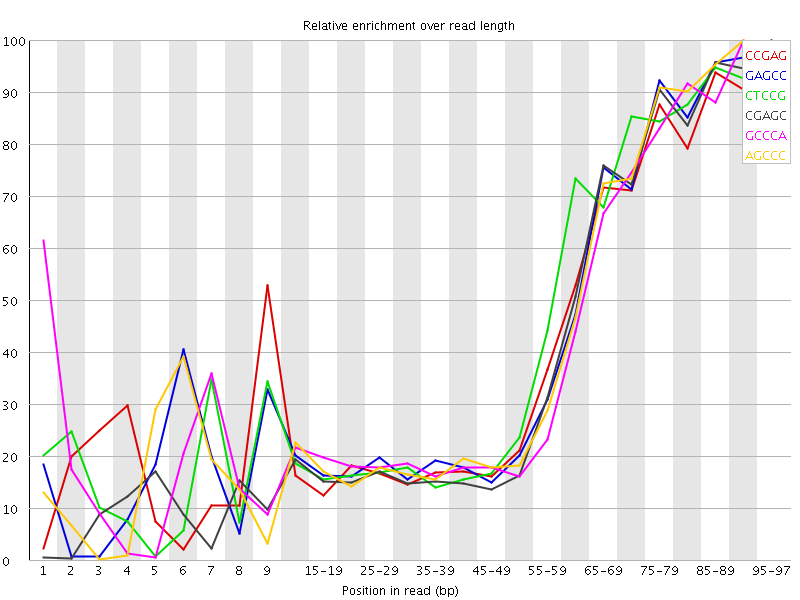
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 164470 | 2.874041 | 6.7331614 | 95-97 |
| GAGCC | 158465 | 2.7691064 | 6.3462305 | 95-97 |
| CTCCG | 166950 | 2.7336836 | 6.0472026 | 95-97 |
| CGAGC | 150065 | 2.62232 | 6.2530284 | 95-97 |
| GCCCA | 150520 | 2.4977124 | 5.854324 | 90-94 |
| AGCCC | 149340 | 2.4781313 | 5.7046666 | 90-94 |
| ACACA | 301085 | 2.3735182 | 7.14374 | 6 |
| CAAAG | 283505 | 2.3535433 | 5.091725 | 4 |
| ATCTC | 293590 | 2.2535748 | 5.1373887 | 95-97 |
| CCCAC | 140995 | 2.2217433 | 5.5195513 | 90-94 |
| CTTTG | 276510 | 2.2055316 | 8.914281 | 9 |
| TACAC | 274645 | 2.1364307 | 8.859578 | 5 |
| GACTC | 184110 | 2.132288 | 5.2054844 | 7 |
| CCACG | 126170 | 2.093651 | 5.538918 | 90-94 |
| GACGG | 110230 | 2.0284493 | 5.978151 | 95-97 |
| GCTTT | 232560 | 1.8549725 | 5.2951694 | 1 |
| CGGAC | 105765 | 1.8481969 | 5.463453 | 90-94 |
| ATACA | 324515 | 1.7854944 | 5.332728 | 6 |
| GGACT | 144830 | 1.766384 | 6.9482813 | 6 |
| GAGAG | 132075 | 1.7190623 | 5.641849 | 7 |
| TGGCT | 137960 | 1.6603261 | 5.0299125 | 6 |
| TTGAG | 184705 | 1.572264 | 5.472721 | 9 |
| GAGTC | 124530 | 1.5187998 | 8.006784 | 9 |
| TATAC | 273900 | 1.487063 | 5.6491103 | 5 |
| GTCTT | 177085 | 1.4124862 | 6.416863 | 1 |
| AAGAC | 169800 | 1.4096106 | 5.647186 | 5 |
| CCATG | 121705 | 1.4095384 | 5.581717 | 9 |
| TATGA | 239530 | 1.369479 | 6.098613 | 4 |
| TGAGT | 158605 | 1.350093 | 6.240388 | 8 |
| GACTT | 164765 | 1.3318452 | 6.5843215 | 7 |
| TCATA | 243910 | 1.3242408 | 6.271235 | 2 |
| GTGTT | 155395 | 1.3052615 | 6.073123 | 1 |
| ATGAG | 148990 | 1.2852582 | 5.065127 | 7 |
| GATTG | 146670 | 1.2484988 | 6.0092626 | 7 |
| GTATG | 144965 | 1.2339853 | 5.4273214 | 3 |
| GTGTA | 144020 | 1.2259412 | 10.66216 | 1 |
| AACAC | 152725 | 1.2039642 | 5.1982274 | 5 |
| GTTCT | 149130 | 1.1895082 | 5.360923 | 1 |
| ATGTG | 133090 | 1.1329019 | 5.1673055 | 7 |
| TGGAC | 92685 | 1.1304101 | 6.9896746 | 5 |
| ACATG | 137045 | 1.1226344 | 5.361939 | 8 |
| GTCCA | 94775 | 1.0976459 | 7.3516035 | 1 |
| ACACT | 135335 | 1.0527548 | 5.4123006 | 6 |
| TGTAT | 181440 | 1.0236279 | 5.0365925 | 2 |
| GAGTA | 118575 | 1.0228839 | 5.270814 | 1 |
| AGACT | 124540 | 1.0201969 | 5.508896 | 6 |
| CATAC | 121695 | 0.94665086 | 5.076625 | 3 |
| GGATA | 103080 | 0.8892167 | 5.0198226 | 1 |
| GTATA | 137295 | 0.7849648 | 7.222326 | 1 |