![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-128_TCCTGAGC-CTAAGCCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1163287 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
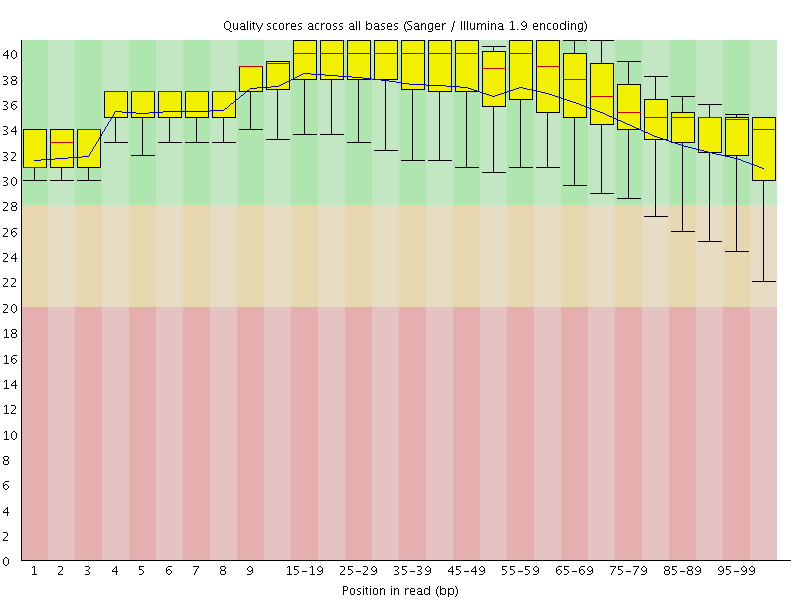
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
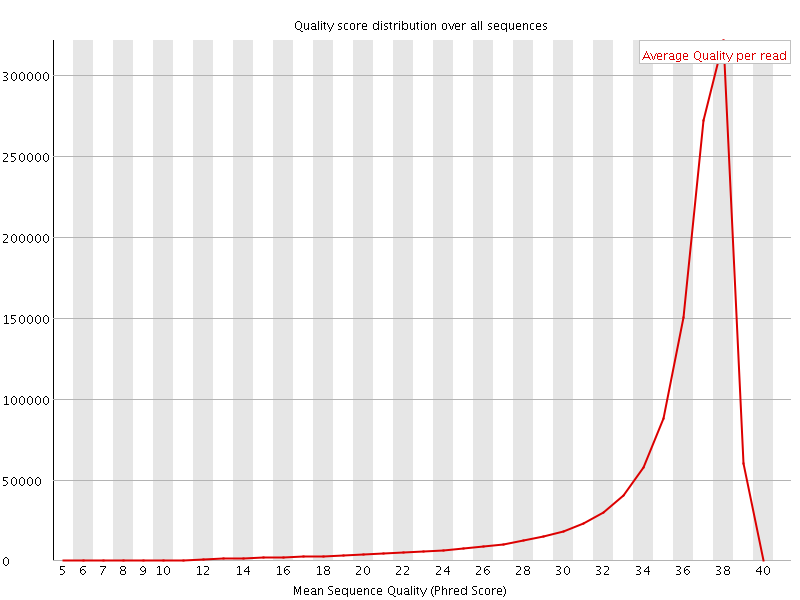
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
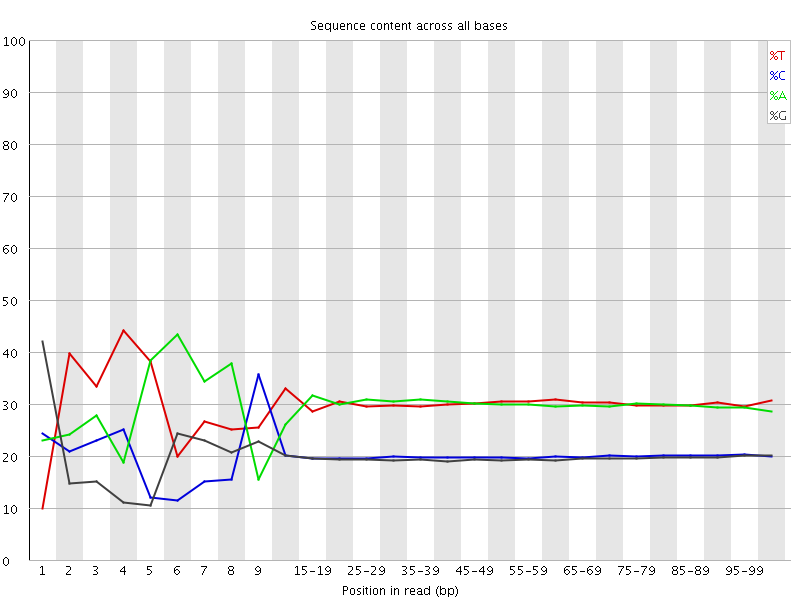
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
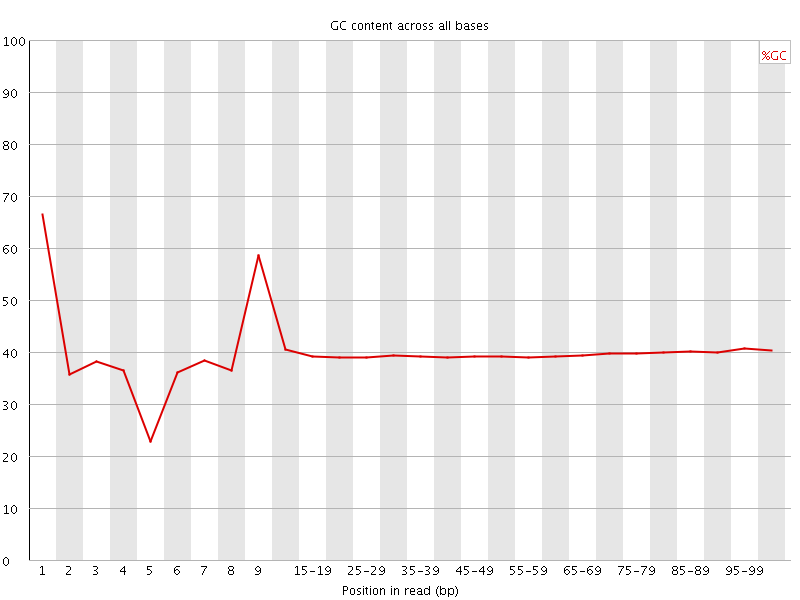
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
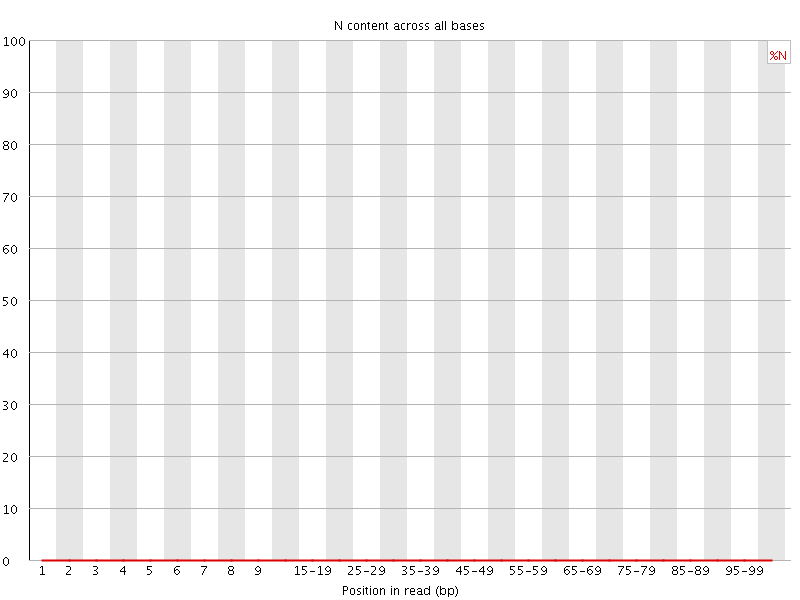
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
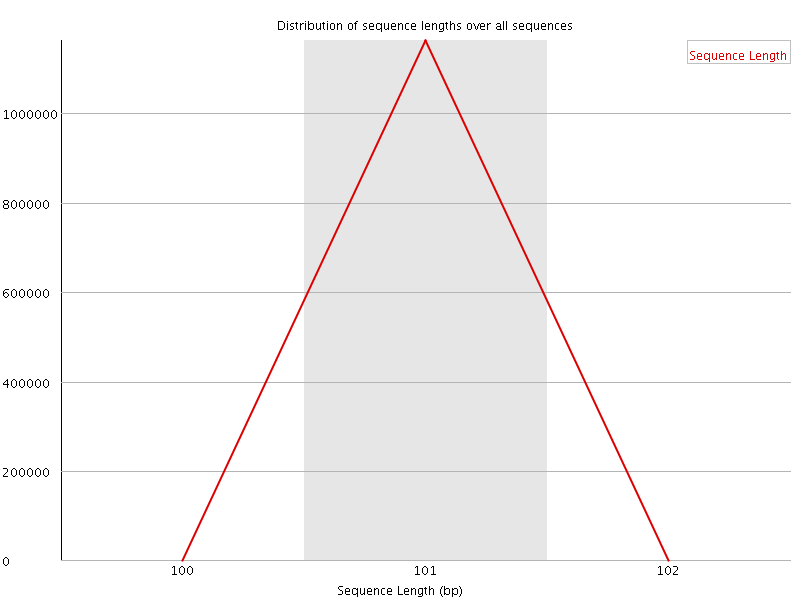
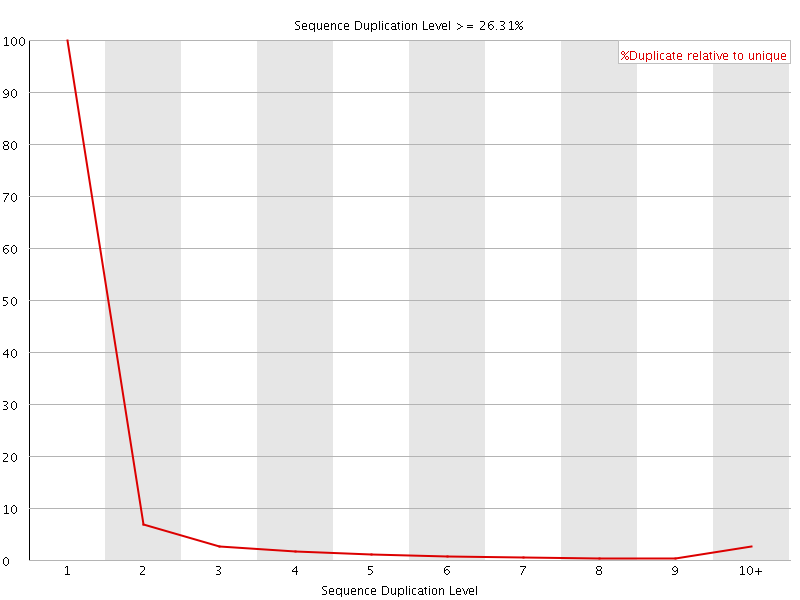
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 1225 | 0.10530505369698104 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
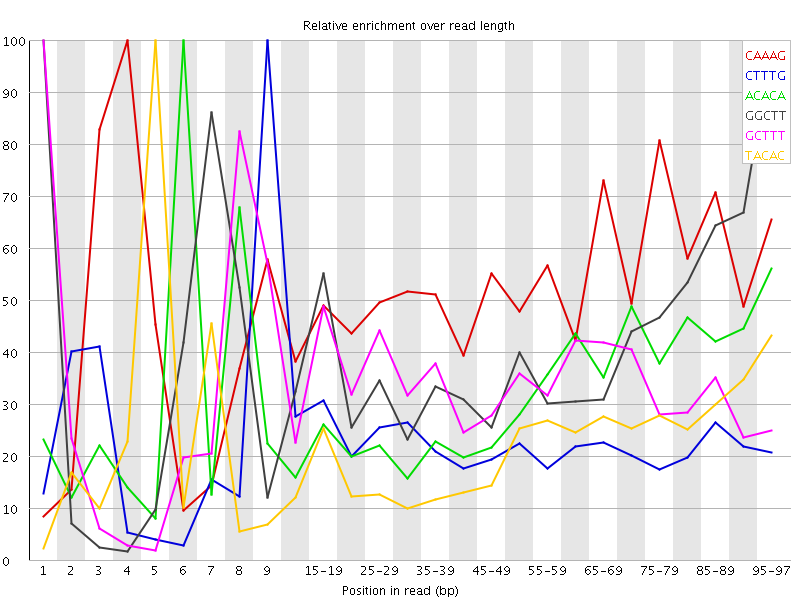
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CAAAG | 323190 | 2.6940432 | 5.1191893 | 4 |
| CTTTG | 315110 | 2.546868 | 11.213835 | 9 |
| ACACA | 283150 | 2.3231459 | 7.2732115 | 6 |
| GGCTT | 176860 | 2.1861842 | 5.3547215 | 1 |
| GCTTT | 261770 | 2.1157494 | 6.241884 | 1 |
| TACAC | 251010 | 2.0383723 | 9.16269 | 5 |
| GCCAA | 162965 | 2.0239496 | 5.5788493 | 1 |
| TTGGC | 156210 | 1.9309275 | 5.1207767 | 5 |
| TGGCT | 155785 | 1.9256741 | 5.688706 | 6 |
| CACAT | 226890 | 1.8425013 | 5.1982865 | 7 |
| GACTC | 148985 | 1.8313881 | 5.758493 | 7 |
| GAGTC | 141085 | 1.7619982 | 10.265646 | 9 |
| CACCA | 143870 | 1.7586882 | 5.115914 | 7 |
| GAGAG | 133065 | 1.705857 | 5.2221813 | 7 |
| TTGAG | 203300 | 1.6866924 | 5.322925 | 9 |
| AAGAC | 194270 | 1.6193935 | 5.7419024 | 5 |
| ATACA | 290835 | 1.5851868 | 5.6140776 | 6 |
| GTGTA | 188720 | 1.5657282 | 10.480155 | 1 |
| CATGA | 188990 | 1.559258 | 5.0973043 | 9 |
| CCATG | 124635 | 1.5320675 | 5.6690516 | 9 |
| GTCTT | 188920 | 1.5269408 | 7.0887723 | 1 |
| GACTT | 186610 | 1.5238657 | 8.656768 | 7 |
| TCATG | 185965 | 1.5185986 | 5.506141 | 5 |
| TGTGT | 184330 | 1.5136554 | 6.949087 | 8 |
| ATGGA | 178190 | 1.4936513 | 5.2575717 | 4 |
| TATGA | 271730 | 1.489329 | 6.0060663 | 4 |
| ACTCA | 180875 | 1.4688282 | 5.2180667 | 8 |
| TGAGT | 176525 | 1.4645516 | 8.087156 | 8 |
| TCATA | 267140 | 1.4411371 | 6.107766 | 2 |
| TCAAG | 174550 | 1.4401212 | 5.0053887 | 8 |
| TCCAT | 179120 | 1.4396905 | 5.6494823 | 2 |
| GTGTT | 173815 | 1.4273098 | 6.242065 | 1 |
| AACAC | 172670 | 1.4166964 | 5.4247108 | 5 |
| ATGAG | 162980 | 1.3661557 | 6.443013 | 7 |
| ACACC | 109865 | 1.3430061 | 5.2877827 | 6 |
| CAAGT | 159810 | 1.3185091 | 5.7494717 | 9 |
| GATTG | 157605 | 1.3075807 | 5.9465737 | 7 |
| TATAC | 236960 | 1.2783253 | 5.2939897 | 5 |
| GTTCT | 152490 | 1.2324964 | 5.516539 | 1 |
| GTCCA | 100060 | 1.229981 | 9.165159 | 1 |
| ACACT | 151110 | 1.2271162 | 5.729881 | 6 |
| TGGAC | 98160 | 1.2259116 | 8.348234 | 5 |
| ATGTG | 143010 | 1.1864923 | 6.976562 | 7 |
| GGACT | 94720 | 1.1829498 | 9.145168 | 6 |
| ACATG | 143320 | 1.1824588 | 5.4375086 | 8 |
| AGACT | 137645 | 1.1356373 | 5.9294925 | 6 |
| ACTTT | 204745 | 1.0932316 | 5.429985 | 8 |
| CCATA | 134555 | 1.0926783 | 5.21149 | 3 |
| GTATG | 130710 | 1.0844445 | 5.022532 | 3 |
| GTATA | 152275 | 0.83460635 | 7.380755 | 1 |