![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-127_TCCTGAGC-AAGGAGTA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 905197 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
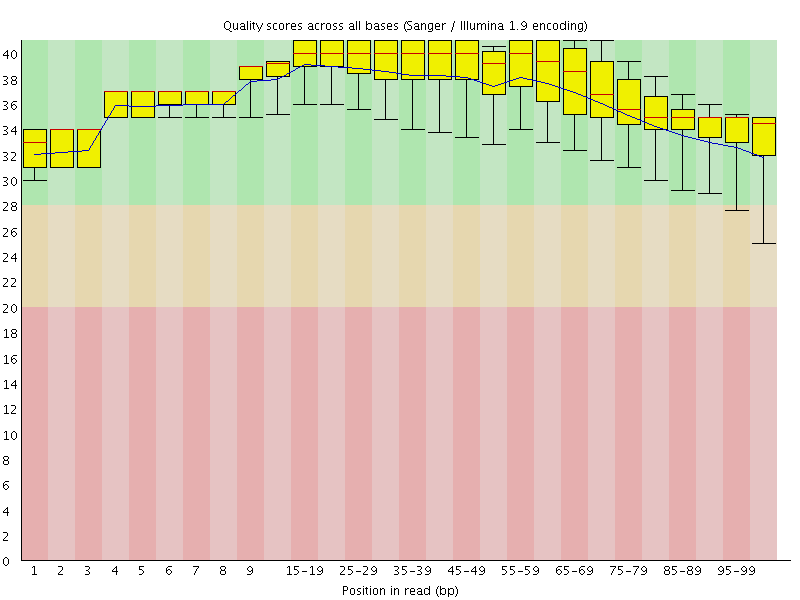
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
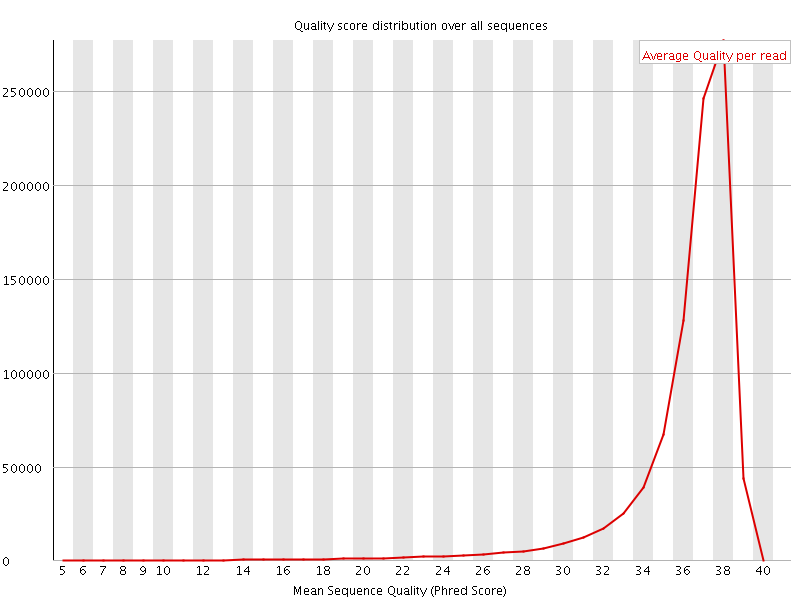
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
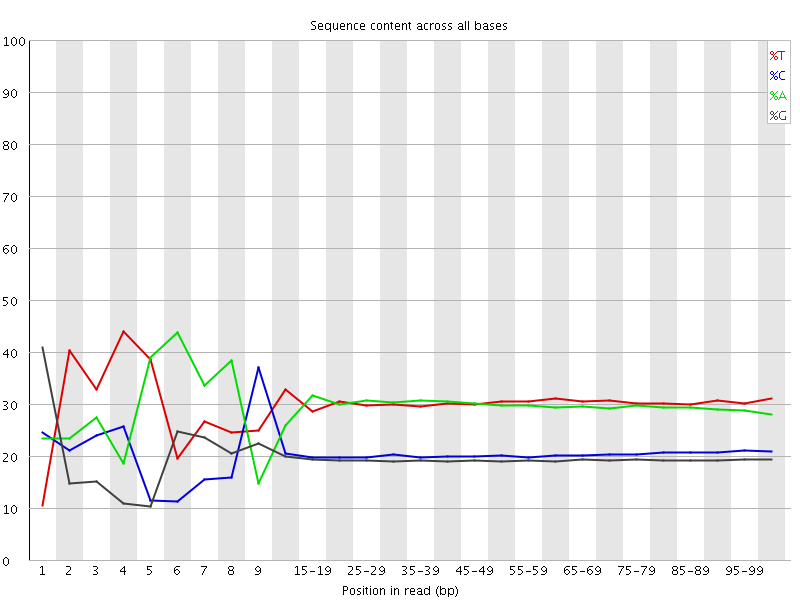
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
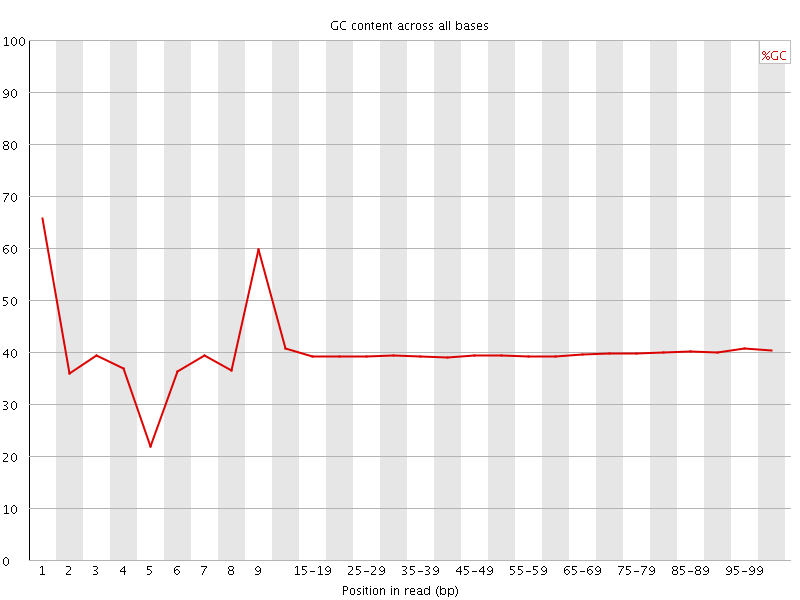
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
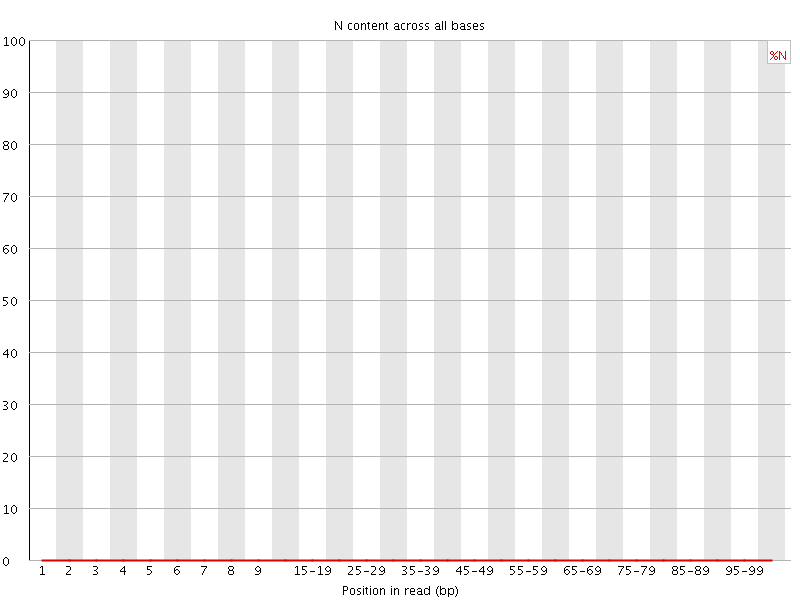
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
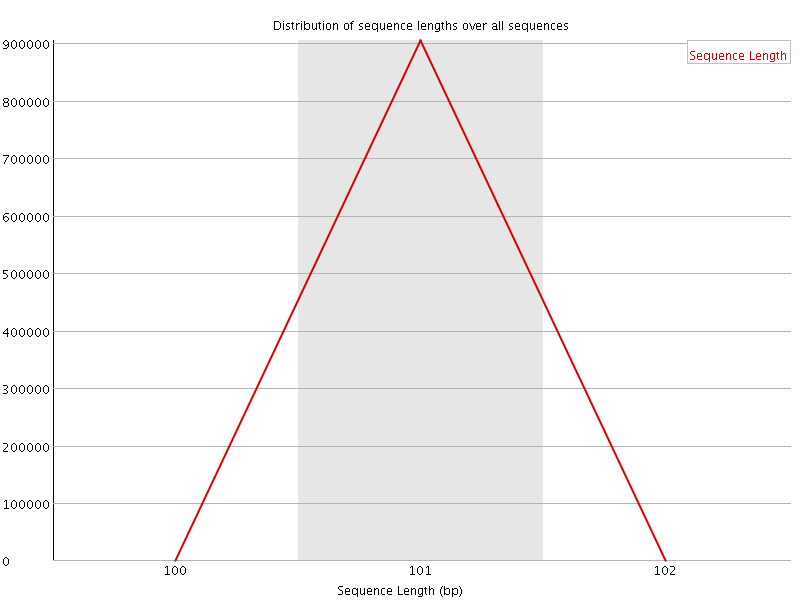
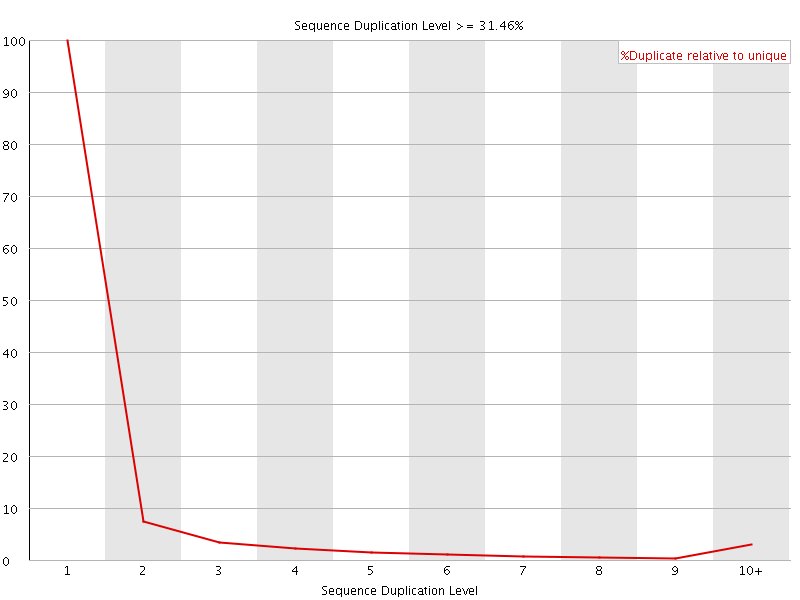
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
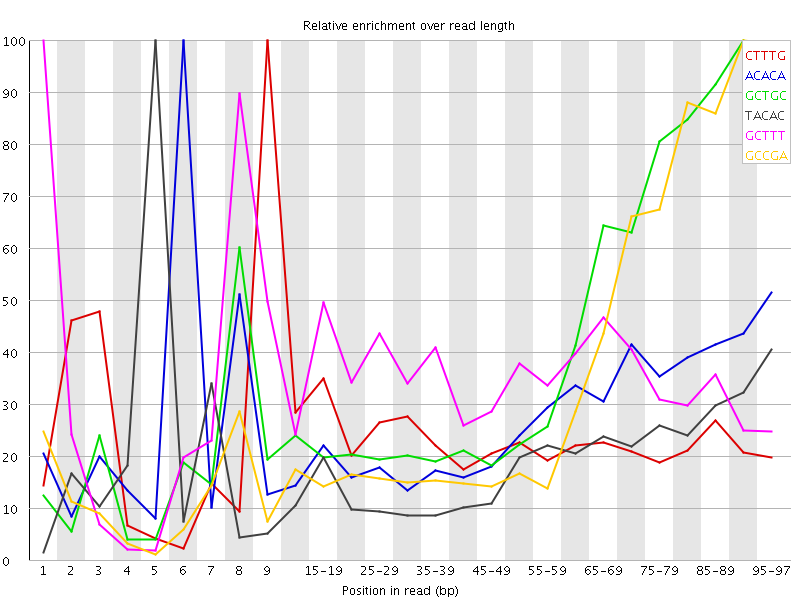
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CTTTG | 237040 | 2.4159472 | 10.271214 | 9 |
| ACACA | 223695 | 2.336601 | 8.459471 | 6 |
| GCTGC | 98585 | 2.3360736 | 5.472522 | 90-94 |
| TACAC | 200955 | 2.0506806 | 10.622708 | 5 |
| GCTTT | 201200 | 2.0506606 | 5.833932 | 1 |
| GCCGA | 81845 | 1.9851667 | 5.371099 | 90-94 |
| GACGC | 80520 | 1.9530288 | 5.3879247 | 95-97 |
| CACAT | 189120 | 1.929908 | 6.3642983 | 7 |
| CGCTG | 81405 | 1.9289756 | 5.2292356 | 95-97 |
| CTGCC | 85165 | 1.9284655 | 5.1768155 | 90-94 |
| GCCAA | 120730 | 1.9216831 | 6.0456347 | 1 |
| CGACG | 77920 | 1.889965 | 5.615362 | 95-97 |
| TGGCT | 115465 | 1.8356235 | 5.843874 | 6 |
| TTGGC | 115225 | 1.8318082 | 5.236118 | 5 |
| CCGAC | 76750 | 1.7789274 | 5.1486874 | 95-97 |
| GAGTC | 107385 | 1.7474551 | 10.401314 | 9 |
| GACTC | 112360 | 1.7472264 | 5.927484 | 7 |
| TGCCG | 73455 | 1.7405922 | 5.0840893 | 90-94 |
| GGCTT | 108690 | 1.7279168 | 5.390437 | 1 |
| ATACA | 238430 | 1.6708858 | 5.607486 | 6 |
| TGTGT | 153660 | 1.6388968 | 5.2863913 | 8 |
| GAGAG | 93665 | 1.632653 | 5.5200133 | 7 |
| TTGAG | 148910 | 1.6257129 | 6.3746576 | 9 |
| CACCA | 103530 | 1.5747364 | 5.1561933 | 7 |
| GTGTA | 141950 | 1.5497277 | 11.862646 | 1 |
| CATGA | 144030 | 1.5380735 | 6.188743 | 9 |
| ATGGA | 136240 | 1.5224875 | 5.350232 | 4 |
| GTCTT | 148085 | 1.5093045 | 6.96611 | 1 |
| AAGAC | 136970 | 1.4971962 | 5.890792 | 5 |
| TATGA | 204930 | 1.4682068 | 6.8845015 | 4 |
| TGAGT | 134425 | 1.4675741 | 7.6031227 | 8 |
| CCATG | 93730 | 1.457525 | 6.2442193 | 9 |
| TCCAT | 145935 | 1.4548877 | 5.8222833 | 2 |
| TCATA | 211360 | 1.4470366 | 7.6048207 | 2 |
| ATGAG | 126720 | 1.4161011 | 6.6280527 | 7 |
| GACTT | 134025 | 1.3982371 | 7.7258177 | 7 |
| ACTCA | 135105 | 1.3787025 | 5.39935 | 8 |
| AACAC | 131095 | 1.3693498 | 5.249216 | 5 |
| TATAC | 195960 | 1.3416033 | 5.8282776 | 5 |
| GTGTT | 123940 | 1.3219111 | 5.556594 | 1 |
| GATTG | 119670 | 1.3064874 | 7.089423 | 7 |
| GTTCT | 122640 | 1.2499653 | 5.171435 | 1 |
| GTCCA | 80060 | 1.2449532 | 9.4515295 | 1 |
| ACACC | 79115 | 1.2033736 | 5.731404 | 6 |
| ACATG | 111950 | 1.1954963 | 6.831035 | 8 |
| TGGAC | 73440 | 1.1950748 | 8.272372 | 5 |
| ACACT | 116315 | 1.1869568 | 5.498087 | 6 |
| ACCAT | 115130 | 1.1748642 | 5.1568494 | 8 |
| GGACT | 70700 | 1.1504874 | 8.428147 | 6 |
| ATGTG | 104875 | 1.1449642 | 5.606945 | 7 |
| CCATA | 108470 | 1.106901 | 5.2319803 | 3 |
| GTATG | 100655 | 1.0988927 | 6.201101 | 3 |
| AGACT | 99000 | 1.0572053 | 6.0176744 | 6 |
| TGTAT | 149630 | 1.0473002 | 5.4728594 | 2 |
| GAGTA | 93305 | 1.0426872 | 5.2798505 | 1 |
| CATAC | 101060 | 1.0312845 | 6.434792 | 3 |
| GTATA | 110645 | 0.7927084 | 7.107037 | 1 |