![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-127_TCCTGAGC-AAGGAGTA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 905197 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
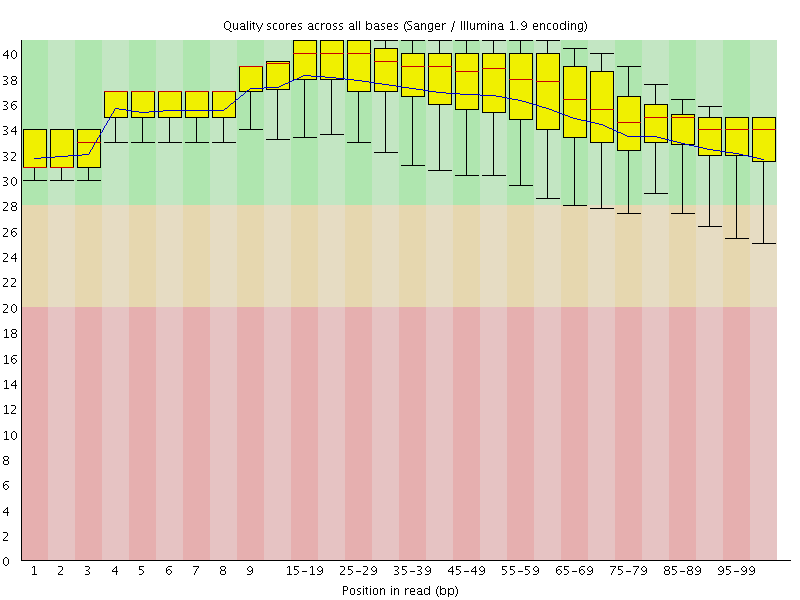
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
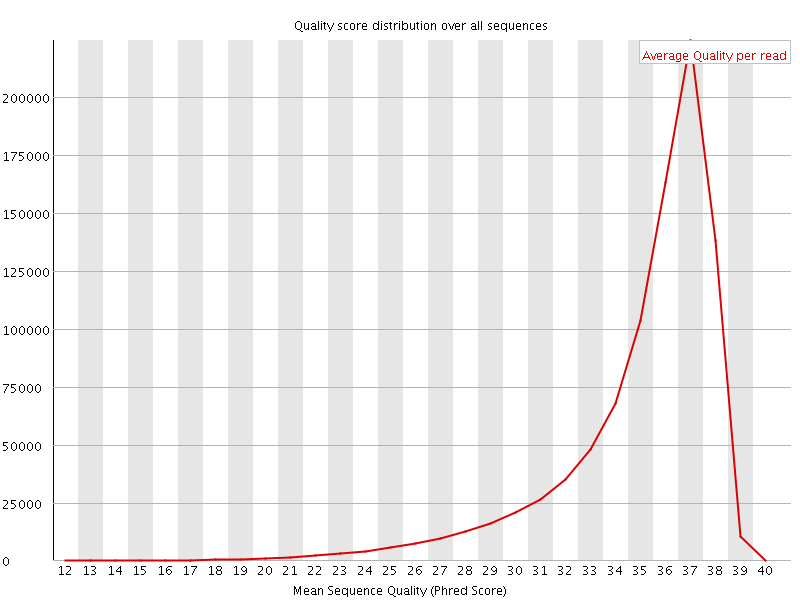
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
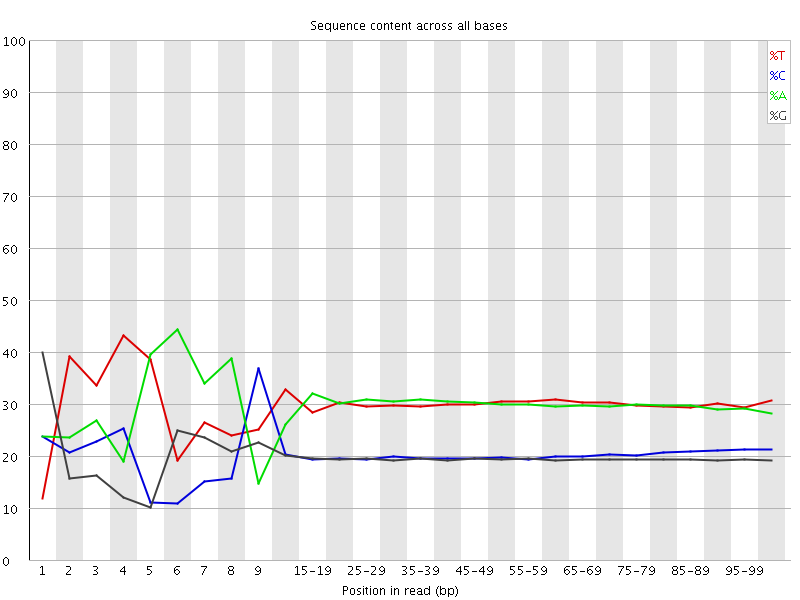
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
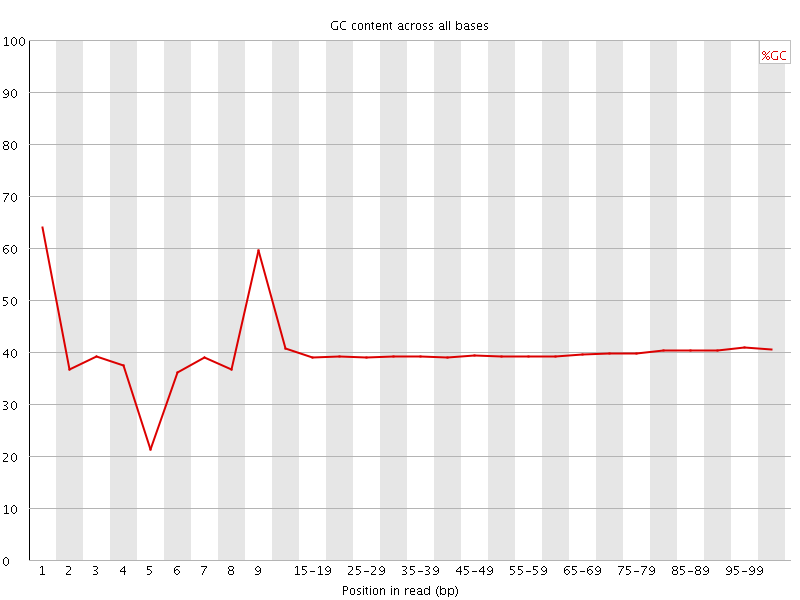
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
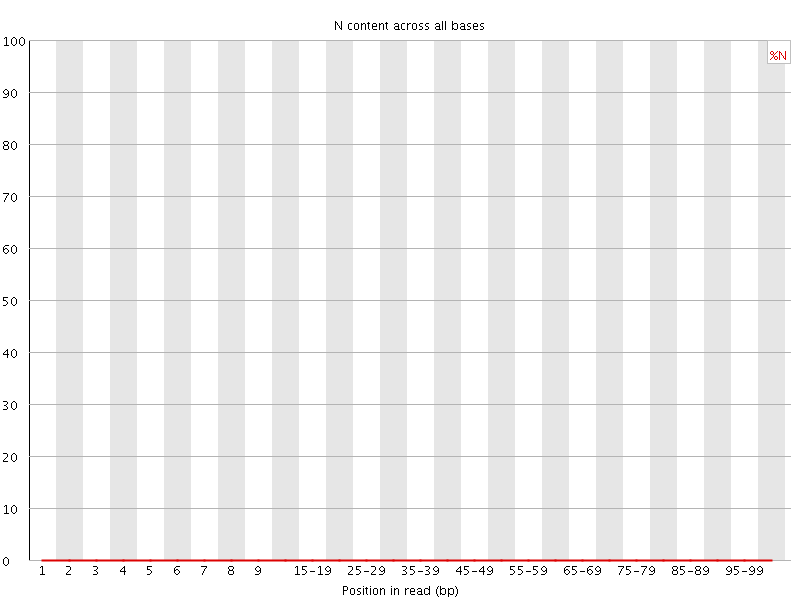
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
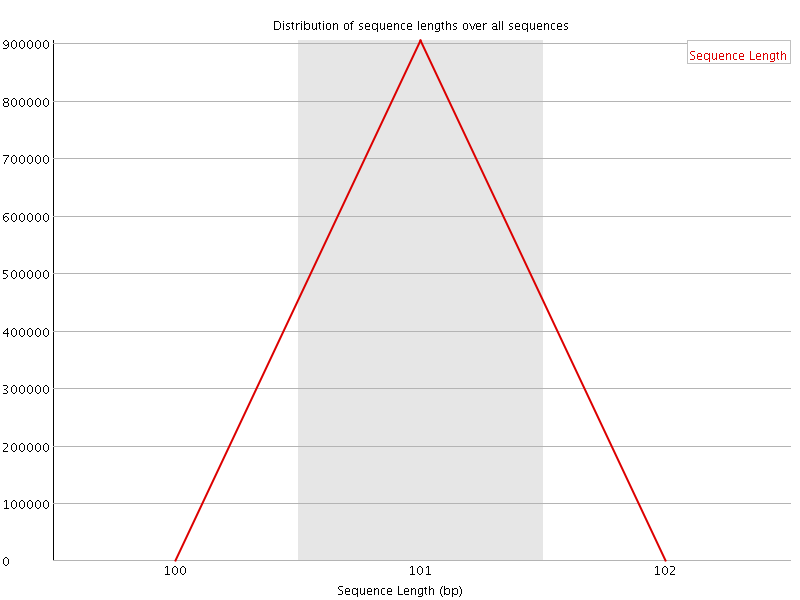
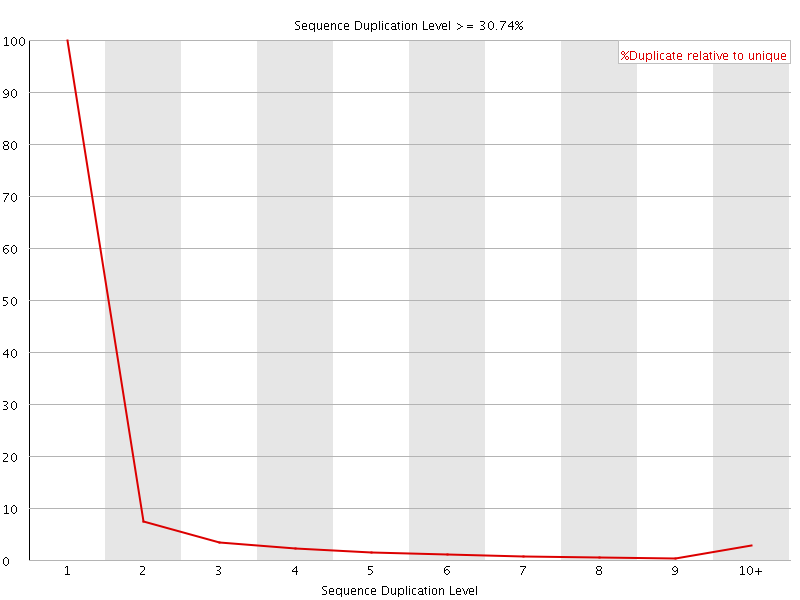
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 1017 | 0.11235123404076681 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
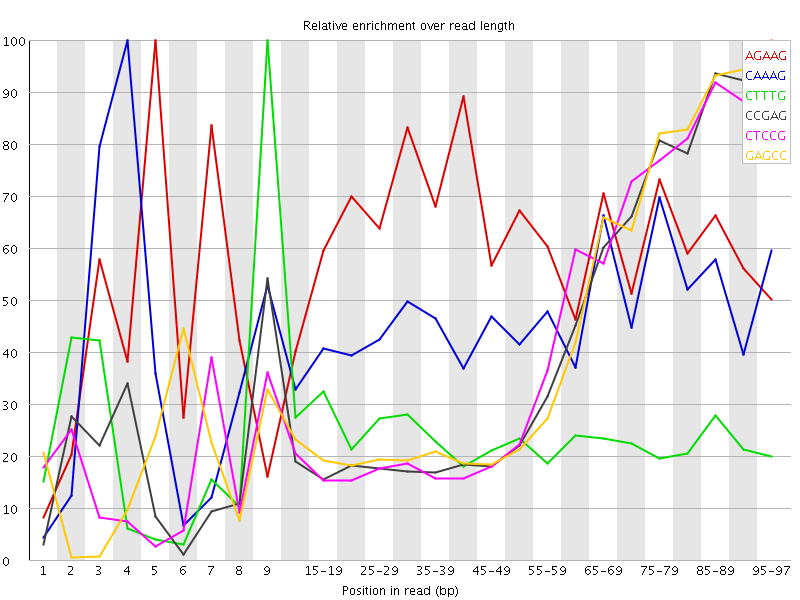
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 277840 | 3.0405533 | 4.945373 | 5 |
| CAAAG | 246570 | 2.619543 | 5.661198 | 4 |
| CTTTG | 234015 | 2.451995 | 10.303129 | 9 |
| CCGAG | 101125 | 2.4227962 | 5.82413 | 95-97 |
| CTCCG | 104320 | 2.4151845 | 5.736536 | 95-97 |
| GAGCC | 99645 | 2.387338 | 5.615019 | 95-97 |
| GACTC | 148690 | 2.3265598 | 6.4184194 | 7 |
| ACACA | 219070 | 2.2594135 | 8.226291 | 6 |
| CGAGC | 92210 | 2.2092066 | 5.448504 | 95-97 |
| AAAGC | 204400 | 2.1715317 | 5.29546 | 5 |
| AGCCC | 92150 | 2.1432915 | 5.2646832 | 90-94 |
| GCCCA | 90855 | 2.113171 | 5.3932548 | 90-94 |
| TACAC | 198400 | 2.036814 | 10.72212 | 5 |
| GCTTT | 194280 | 2.035654 | 5.4670954 | 1 |
| GCCAA | 121805 | 1.9146998 | 5.5263968 | 1 |
| CACAT | 180365 | 1.851663 | 6.0927925 | 7 |
| TGGCT | 114220 | 1.8324999 | 5.2586446 | 6 |
| TTGAG | 160730 | 1.7428035 | 6.3562813 | 9 |
| GAGTC | 105435 | 1.6993771 | 8.088549 | 9 |
| GAGAG | 97675 | 1.6291611 | 5.596442 | 7 |
| CACCA | 106735 | 1.6288083 | 5.682633 | 7 |
| ATACA | 233730 | 1.6217126 | 5.493776 | 6 |
| GGCTT | 100660 | 1.6149486 | 5.687058 | 1 |
| AAGAC | 149530 | 1.5885966 | 6.243286 | 5 |
| ATGGA | 143675 | 1.5650777 | 5.3134675 | 4 |
| GACTT | 145545 | 1.5320619 | 8.370439 | 7 |
| CATGA | 143830 | 1.5210086 | 5.9274154 | 9 |
| AGACT | 143415 | 1.5166199 | 6.070986 | 6 |
| CCATG | 96735 | 1.5136173 | 6.4411798 | 9 |
| TATGA | 209360 | 1.4894383 | 7.1956234 | 4 |
| TGAGT | 134040 | 1.4534026 | 6.1249533 | 8 |
| GTCTT | 138260 | 1.4486799 | 5.406124 | 1 |
| TGTGT | 133980 | 1.446067 | 5.594373 | 8 |
| TCATA | 206320 | 1.4249433 | 6.9794703 | 2 |
| GTGTT | 131015 | 1.4140652 | 6.0136333 | 1 |
| ACTCA | 135270 | 1.3887088 | 5.7941265 | 8 |
| GTGTA | 124765 | 1.3528333 | 12.040805 | 1 |
| ATGAG | 123750 | 1.3480309 | 5.044096 | 7 |
| GATTG | 123595 | 1.3401469 | 7.102843 | 7 |
| TATAC | 190455 | 1.3153722 | 6.1348944 | 5 |
| AACAC | 124370 | 1.28271 | 5.270827 | 5 |
| ACACC | 83475 | 1.2738537 | 6.1561856 | 6 |
| TGGAC | 78405 | 1.2637137 | 8.54182 | 5 |
| GTATG | 114515 | 1.2416921 | 6.8767715 | 3 |
| ACCAT | 119385 | 1.2256302 | 5.371016 | 8 |
| GGACT | 75765 | 1.2211628 | 8.854421 | 6 |
| ATGTG | 110770 | 1.2010849 | 5.6465235 | 7 |
| ATTGA | 166225 | 1.1825653 | 5.1500797 | 8 |
| GTCCA | 75030 | 1.1739981 | 7.071586 | 1 |
| ACATG | 107650 | 1.1384034 | 6.2350664 | 8 |
| ACACT | 108590 | 1.1148065 | 5.385949 | 6 |
| TGTAT | 153860 | 1.0895606 | 5.9407363 | 2 |
| GAGTA | 98295 | 1.0707451 | 5.292866 | 1 |
| ACTTT | 154785 | 1.064099 | 5.0066633 | 8 |
| CATAC | 96870 | 0.99448675 | 6.003193 | 3 |
| GTATA | 114520 | 0.8147233 | 6.647816 | 1 |
| ATACT | 114240 | 0.7889954 | 5.0499015 | 6 |