![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-126_TCCTGAGC-ACTGCATA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1173561 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
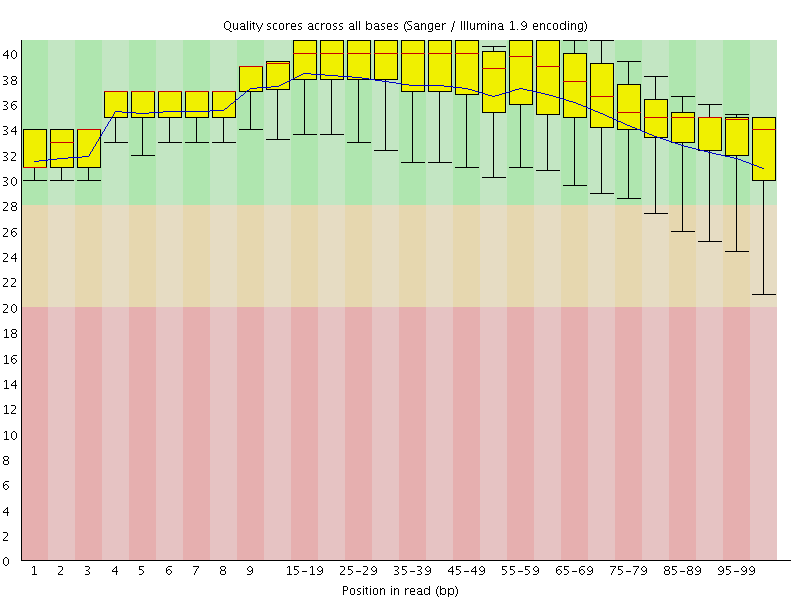
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
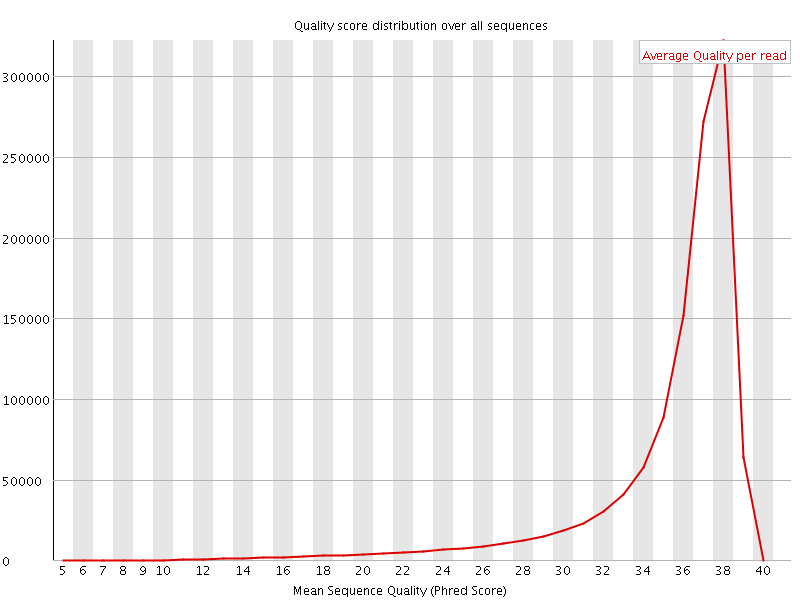
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
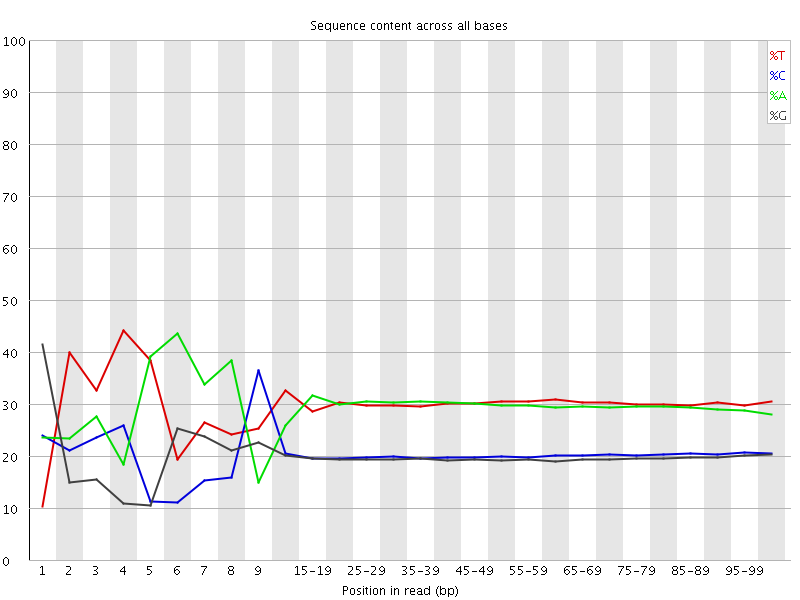
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
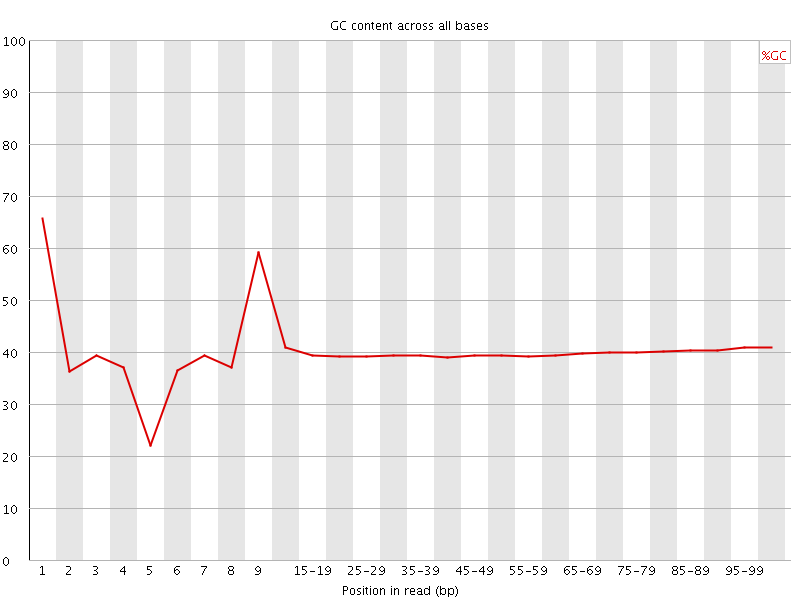
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
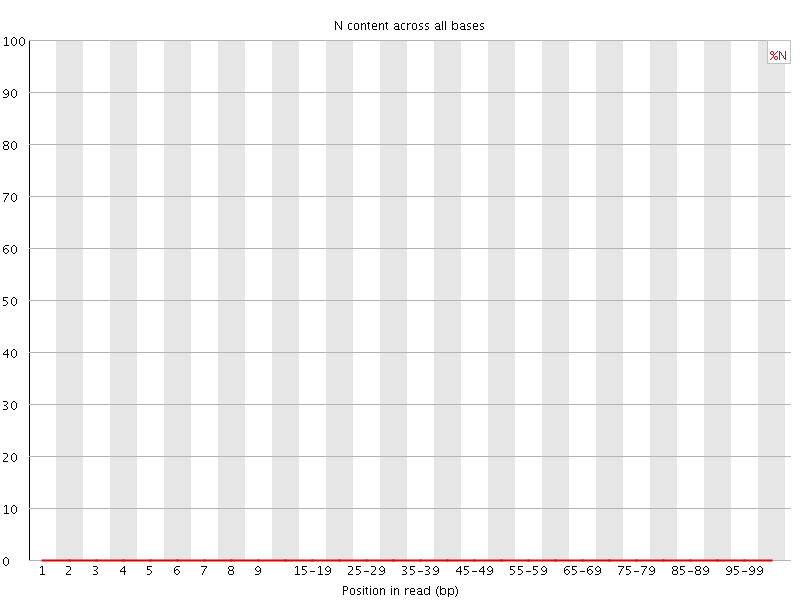
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
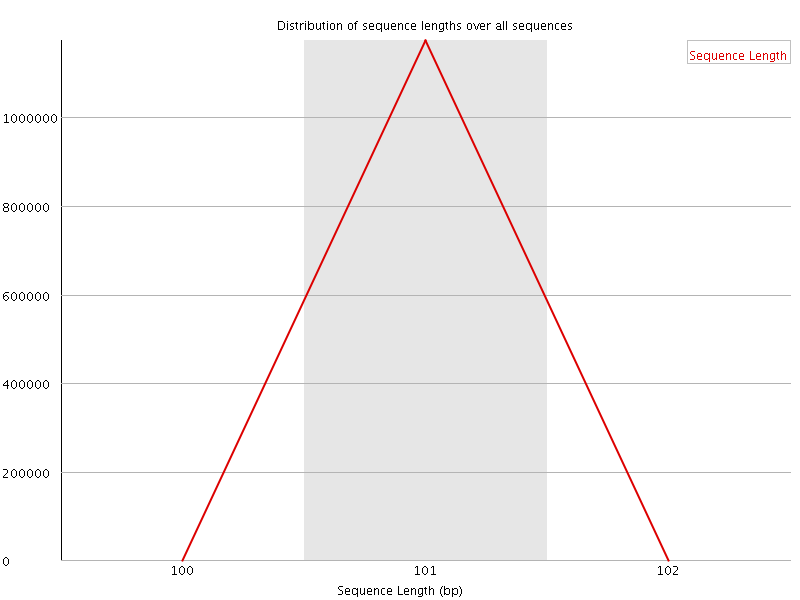
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
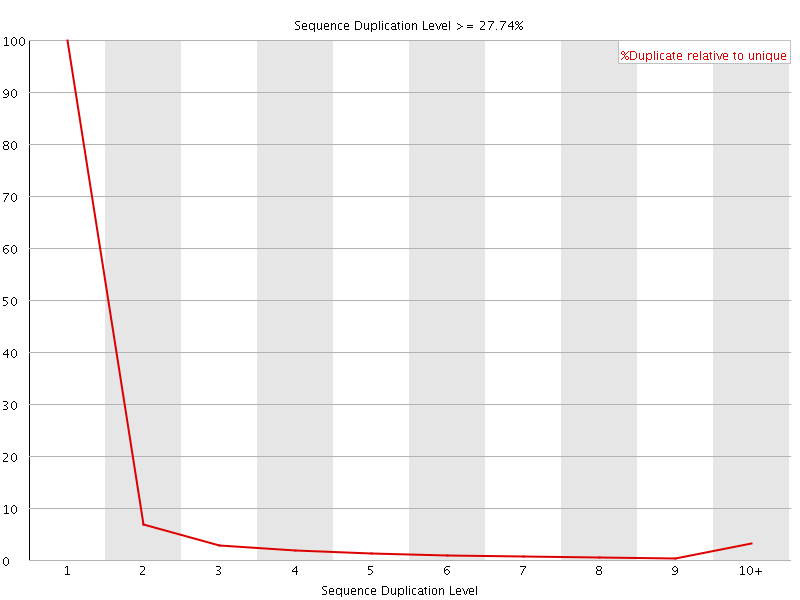
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
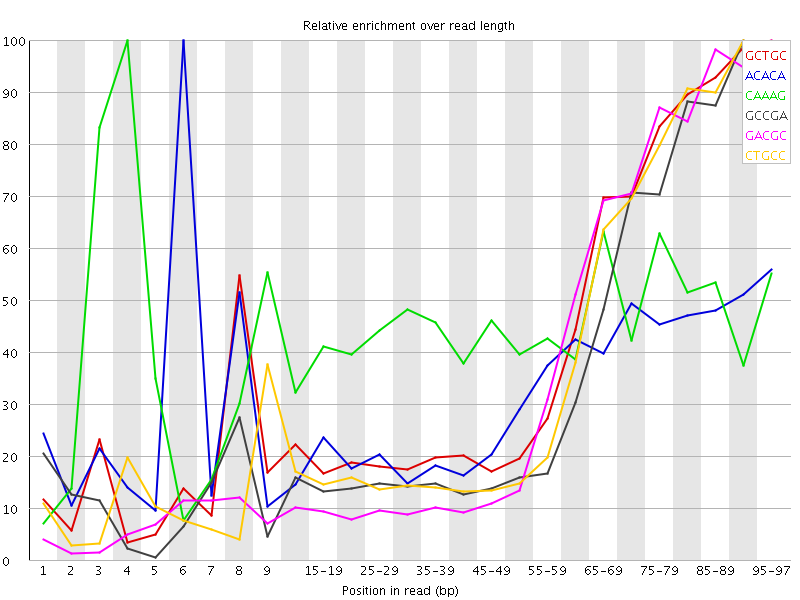
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 144555 | 2.6132085 | 6.05869 | 95-97 |
| ACACA | 294950 | 2.4113867 | 7.513011 | 6 |
| CAAAG | 280610 | 2.343529 | 5.2140207 | 4 |
| GCCGA | 123410 | 2.2699625 | 6.1069994 | 90-94 |
| GACGC | 123130 | 2.2648122 | 5.8553452 | 95-97 |
| CTGCC | 126430 | 2.2373936 | 5.6833453 | 90-94 |
| CGCTG | 122660 | 2.2173994 | 5.681666 | 95-97 |
| CTTTG | 278820 | 2.2105942 | 9.17384 | 9 |
| CGACG | 120160 | 2.2101831 | 6.140827 | 95-97 |
| TACAC | 266775 | 2.1435628 | 9.257242 | 5 |
| CCGAC | 114930 | 2.0694408 | 5.653369 | 95-97 |
| CACAT | 253915 | 2.0402315 | 5.4201355 | 7 |
| TGCCG | 111640 | 2.018184 | 5.7934012 | 90-94 |
| GCTTT | 232220 | 1.841131 | 5.288351 | 1 |
| GCCAA | 142430 | 1.7466056 | 5.2765436 | 1 |
| ATACA | 317120 | 1.7353557 | 5.29147 | 6 |
| GAGAG | 130555 | 1.6706456 | 5.361625 | 7 |
| TGGCT | 136625 | 1.6531701 | 5.363149 | 6 |
| GACTC | 128735 | 1.5515387 | 5.6225452 | 7 |
| TTGAG | 188005 | 1.5492823 | 5.738669 | 9 |
| GAGTC | 121945 | 1.501339 | 7.851201 | 9 |
| TATAC | 270050 | 1.4523847 | 5.8022876 | 5 |
| AAGAC | 173570 | 1.4495788 | 6.0323825 | 5 |
| GTGTA | 173930 | 1.4332952 | 10.801375 | 1 |
| CCATG | 117695 | 1.4184827 | 5.5290785 | 9 |
| GACTT | 168655 | 1.360541 | 7.3311806 | 7 |
| GTCTT | 168405 | 1.3351808 | 6.3537135 | 1 |
| TCATA | 246075 | 1.3234425 | 6.352694 | 2 |
| TATGA | 240690 | 1.3223433 | 6.0444427 | 4 |
| TGAGT | 159165 | 1.3116221 | 6.002501 | 8 |
| ACTCA | 161995 | 1.3016454 | 5.0266886 | 8 |
| AACAC | 157250 | 1.2856097 | 5.151748 | 5 |
| GTGTT | 156330 | 1.2661235 | 6.0543694 | 1 |
| ATGAG | 147815 | 1.2393872 | 5.0257263 | 7 |
| GATTG | 149090 | 1.2285976 | 6.166219 | 7 |
| GTTCT | 150545 | 1.1935797 | 5.0383563 | 1 |
| ACATG | 140580 | 1.1538873 | 5.493132 | 8 |
| TGGAC | 93345 | 1.1492269 | 7.8117704 | 5 |
| GTCCA | 92535 | 1.1152495 | 8.0156 | 1 |
| ACACT | 138285 | 1.1111333 | 5.427859 | 6 |
| GGACT | 89135 | 1.0973951 | 7.833122 | 6 |
| AGACT | 127040 | 1.0427501 | 5.7636113 | 6 |
| GAGTA | 119965 | 1.0058728 | 5.19412 | 1 |
| TGTAT | 185040 | 0.9991357 | 5.1940126 | 2 |
| GTATG | 119215 | 0.9824084 | 5.4844956 | 3 |
| GTATA | 143225 | 0.7868737 | 7.5103426 | 1 |