![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-126_TCCTGAGC-ACTGCATA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1173561 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
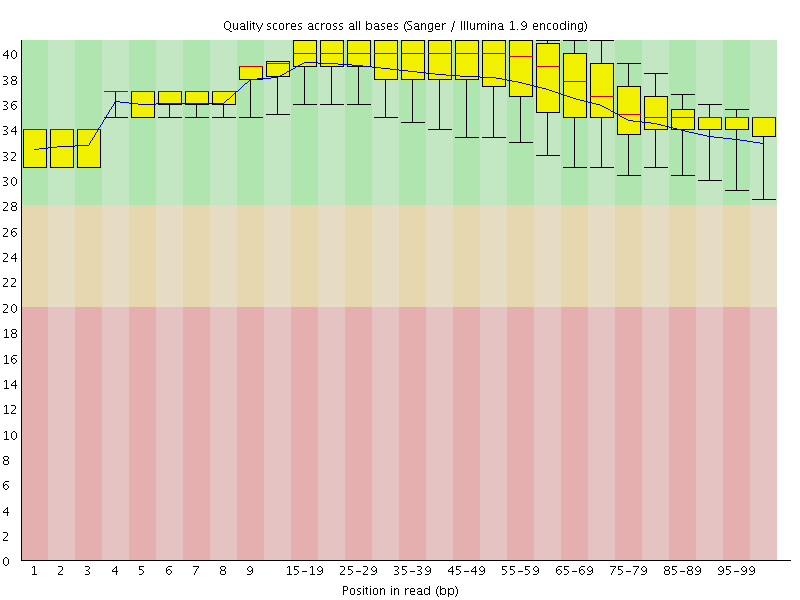
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
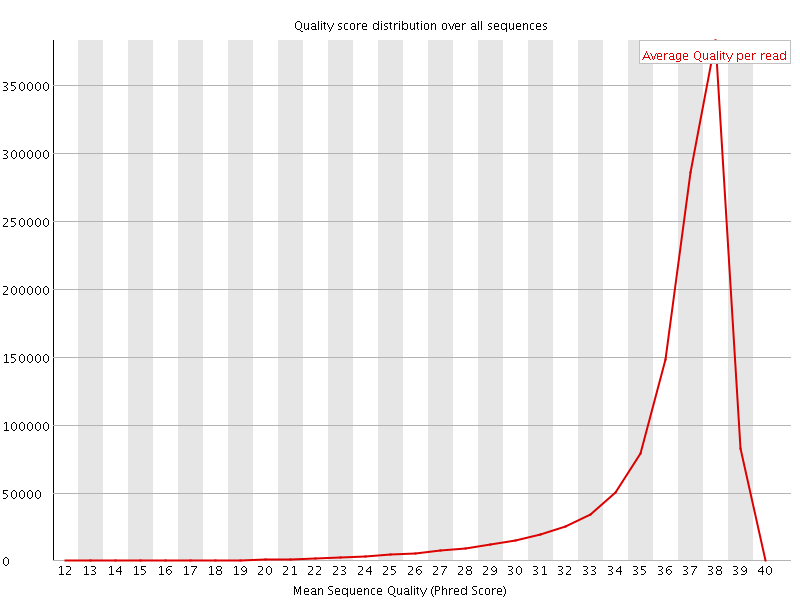
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
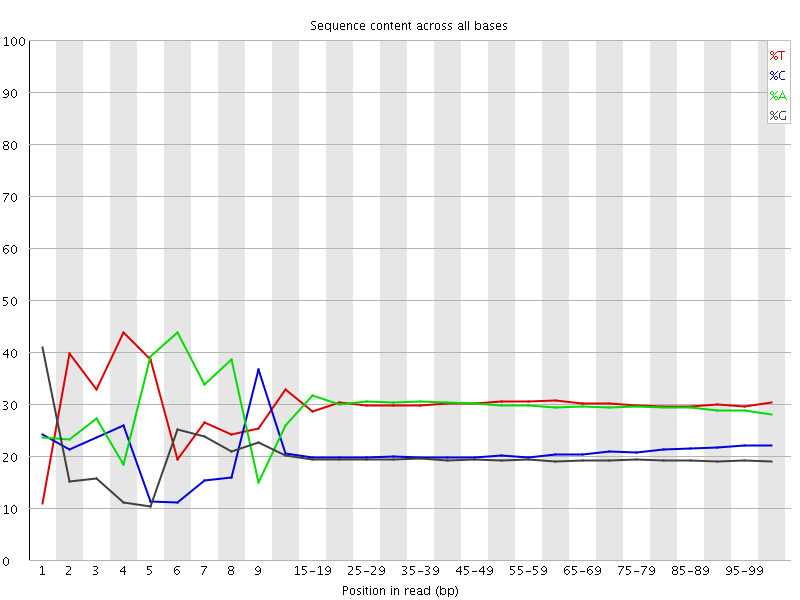
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
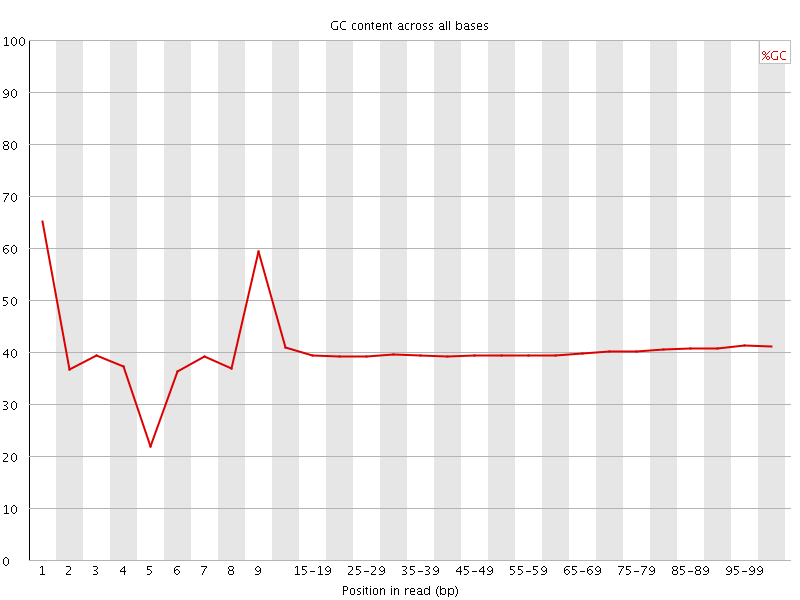
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
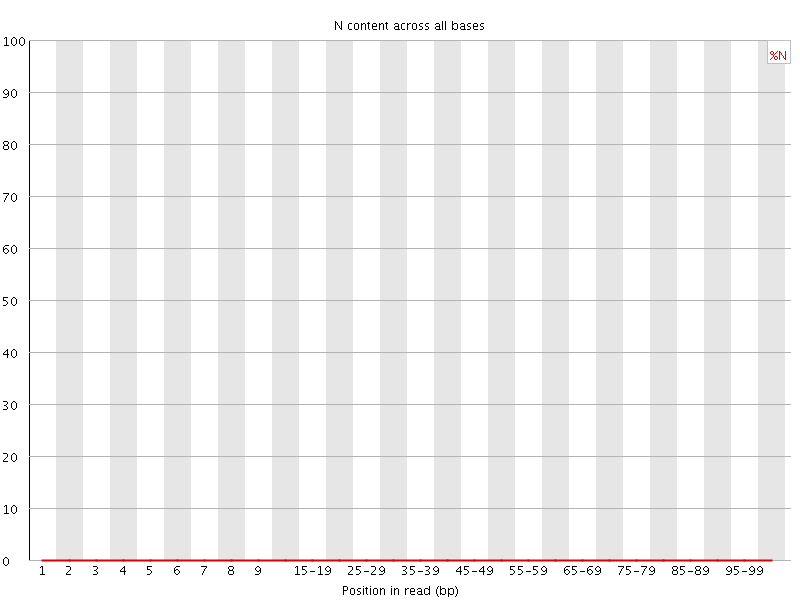
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
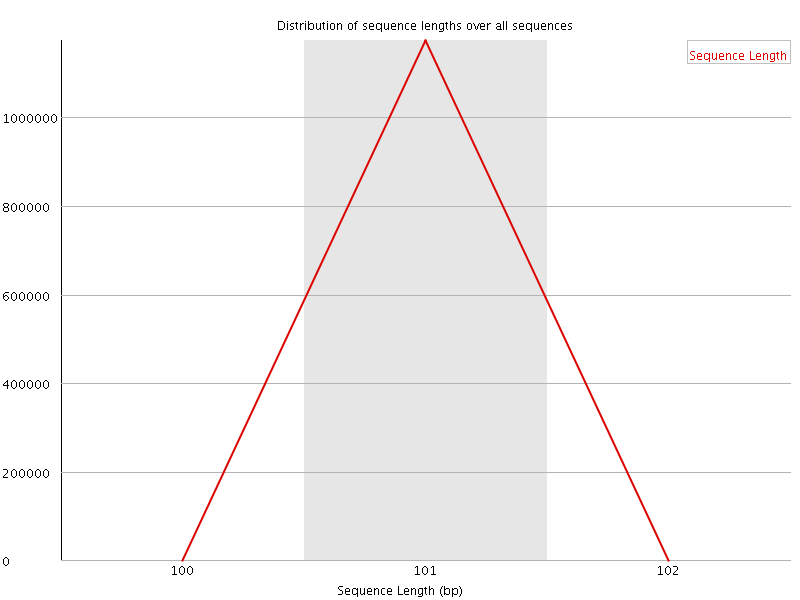
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
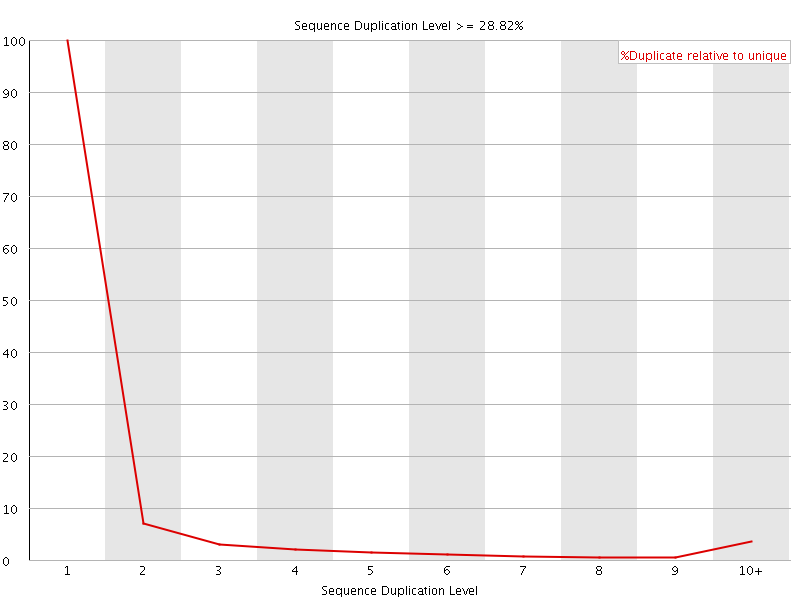
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
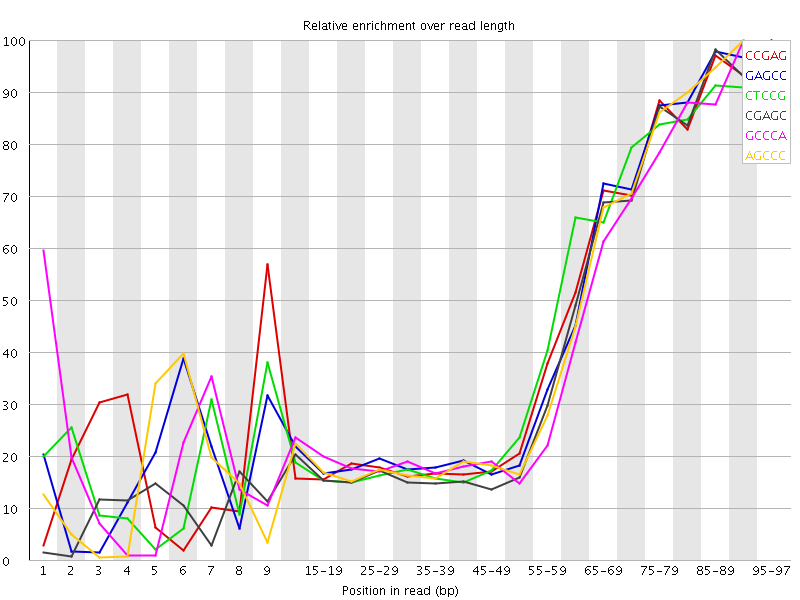
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 152875 | 2.7875147 | 6.4125266 | 95-97 |
| GAGCC | 147125 | 2.6826696 | 6.1237273 | 95-97 |
| CTCCG | 156405 | 2.6653082 | 6.097671 | 95-97 |
| CGAGC | 138925 | 2.5331514 | 6.138462 | 95-97 |
| GCCCA | 140485 | 2.4320428 | 5.8287334 | 90-94 |
| AGCCC | 138470 | 2.3971598 | 5.620564 | 90-94 |
| CAAAG | 284195 | 2.3695896 | 5.206765 | 4 |
| ACACA | 294370 | 2.3302944 | 7.78744 | 6 |
| GACTC | 189360 | 2.2394571 | 5.280919 | 7 |
| CTTTG | 278020 | 2.2110534 | 9.142188 | 9 |
| CCCAC | 132950 | 2.185195 | 5.5148168 | 90-94 |
| CATCT | 279200 | 2.1416261 | 5.030717 | 95-97 |
| TACAC | 268225 | 2.090124 | 10.004287 | 5 |
| CCACG | 117905 | 2.0411434 | 5.511443 | 90-94 |
| CACAT | 253965 | 1.9790038 | 5.3459463 | 7 |
| GCTTT | 234045 | 1.8613266 | 5.479774 | 1 |
| GAGAG | 128350 | 1.7106996 | 5.706029 | 7 |
| ATACA | 316015 | 1.7089847 | 5.1206856 | 6 |
| GCCAA | 141940 | 1.7053117 | 5.0916176 | 1 |
| TGGCT | 136490 | 1.6735946 | 5.487146 | 6 |
| TTGAG | 184455 | 1.569631 | 5.5656323 | 9 |
| AGACT | 188145 | 1.5442046 | 5.797852 | 6 |
| GAGTC | 123075 | 1.5330764 | 8.050438 | 9 |
| TATAC | 270170 | 1.4382125 | 5.8897524 | 5 |
| AAGAC | 171360 | 1.4287827 | 5.6918674 | 5 |
| CATGA | 172960 | 1.4195735 | 5.0497427 | 9 |
| GTCTT | 177585 | 1.4123082 | 6.3667183 | 1 |
| CCATG | 118540 | 1.4019078 | 5.9689875 | 9 |
| TATGA | 243175 | 1.3634659 | 6.271456 | 4 |
| TGAGT | 160095 | 1.3623383 | 6.291764 | 8 |
| GACTT | 167360 | 1.3521328 | 7.113428 | 7 |
| GTGTT | 157580 | 1.3199689 | 6.3240695 | 1 |
| TCATA | 246025 | 1.3096801 | 6.213087 | 2 |
| ATGAG | 149240 | 1.2901402 | 5.205574 | 7 |
| ACTCA | 163175 | 1.2715293 | 5.1305976 | 8 |
| GATTG | 147875 | 1.2583512 | 6.1473627 | 7 |
| GTGTA | 147460 | 1.2548198 | 11.070651 | 1 |
| AACAC | 157130 | 1.243874 | 5.0201936 | 5 |
| GTATG | 140830 | 1.1984015 | 5.697656 | 3 |
| GTTCT | 149500 | 1.1889522 | 5.132709 | 1 |
| TGGAC | 92525 | 1.1525321 | 7.603527 | 5 |
| ACATG | 139785 | 1.1472889 | 5.571032 | 8 |
| ACACC | 99155 | 1.1310301 | 5.3257556 | 6 |
| GTCCA | 94050 | 1.112278 | 7.6671033 | 1 |
| ACACT | 140075 | 1.0915242 | 5.463066 | 6 |
| GGACT | 87280 | 1.087198 | 7.3740315 | 6 |
| GAGTA | 119130 | 1.0298473 | 5.3654604 | 1 |
| TGTAT | 186035 | 1.0267756 | 5.189244 | 2 |
| CATAC | 118720 | 0.92511696 | 5.168378 | 3 |
| GTATA | 143665 | 0.8055202 | 7.0307126 | 1 |