![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-125_TCCTGAGC-GTAAGGAG_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 843328 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
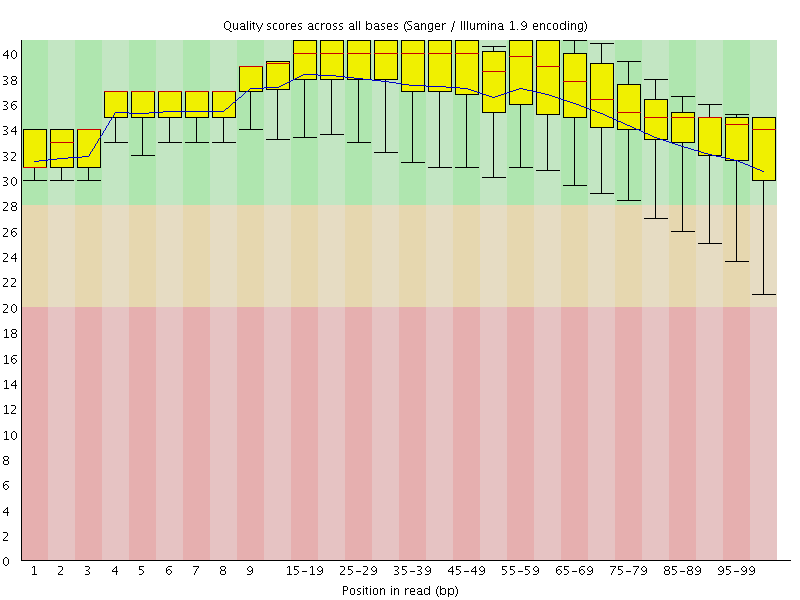
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
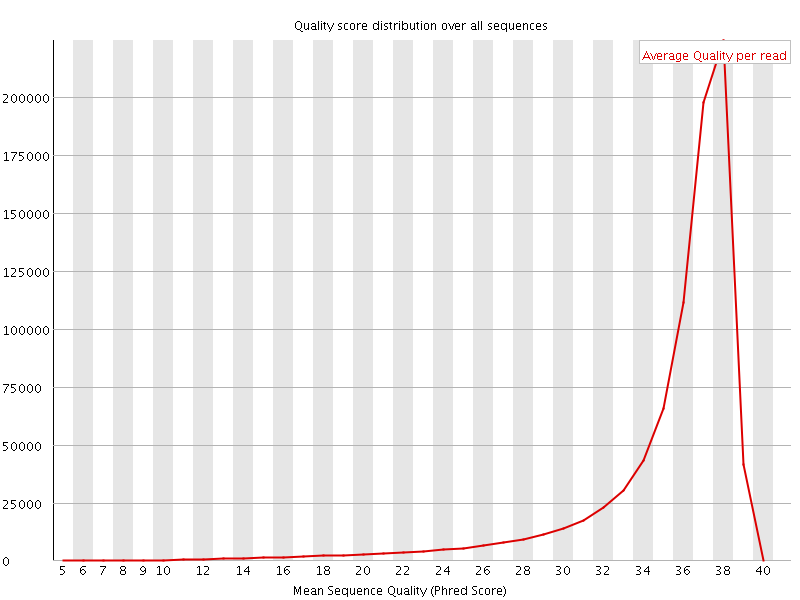
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
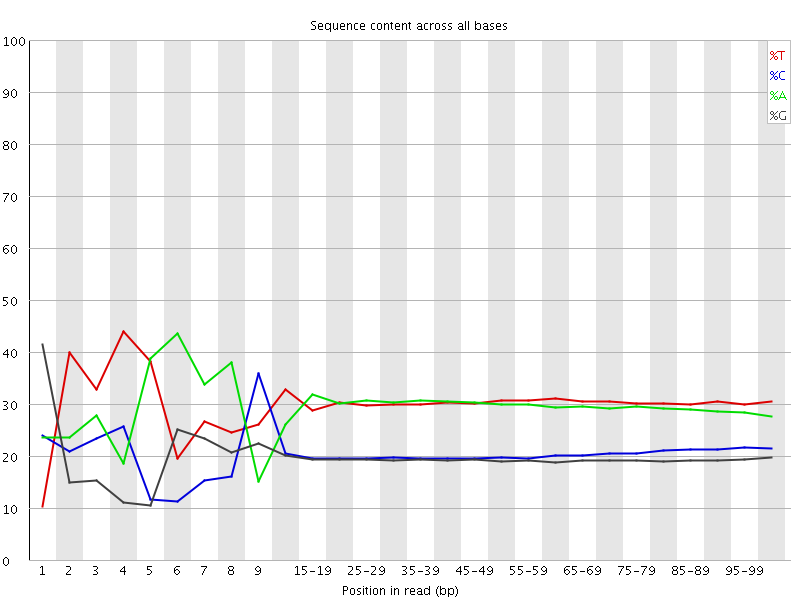
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
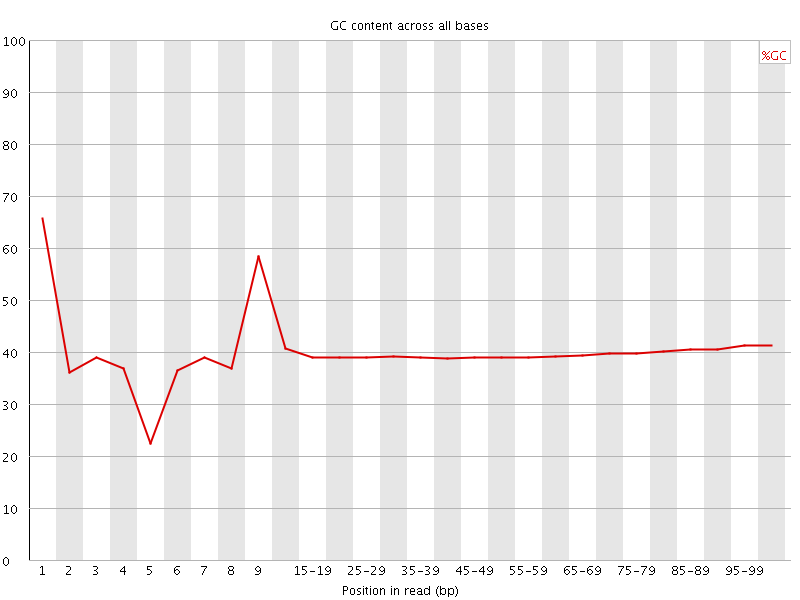
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
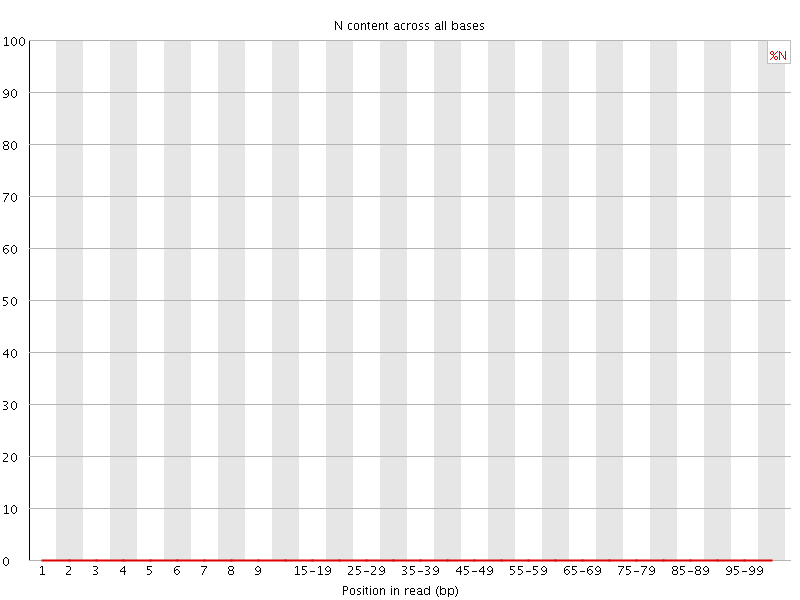
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
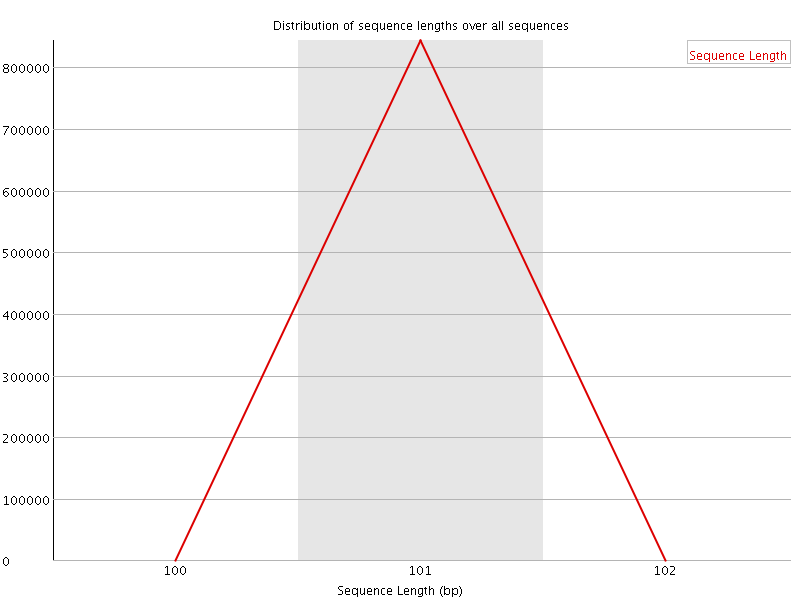
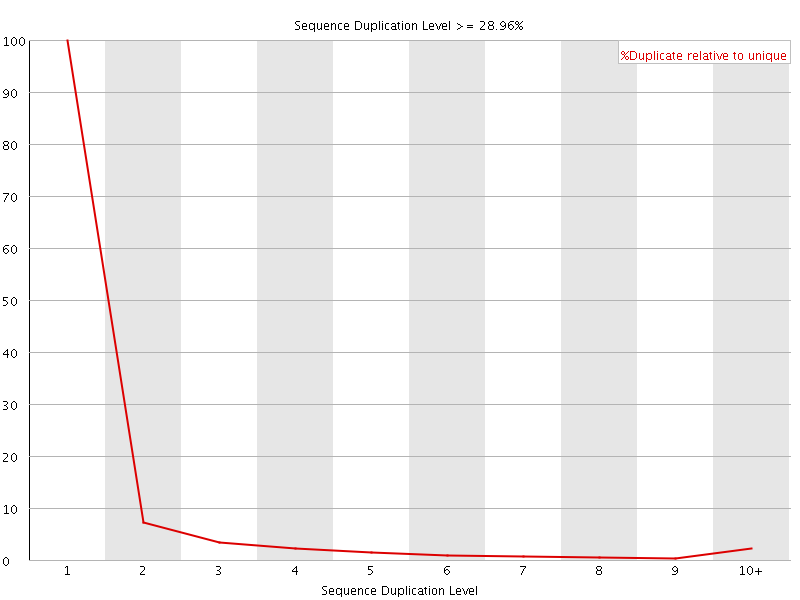
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
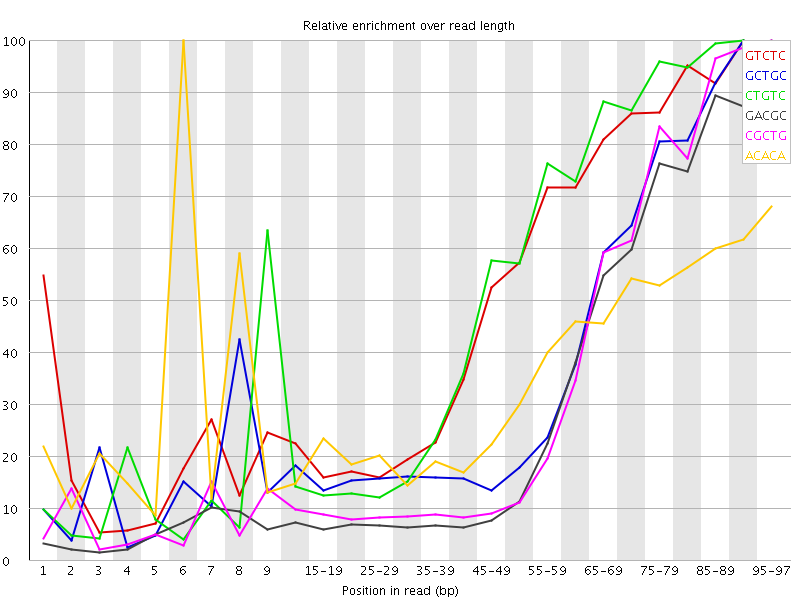
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 191085 | 3.1178272 | 5.8463984 | 90-94 |
| GCTGC | 115020 | 2.9191873 | 7.4125133 | 90-94 |
| CTGTC | 177755 | 2.9003289 | 5.4079976 | 90-94 |
| GACGC | 101875 | 2.643483 | 8.013169 | 95-97 |
| CGCTG | 100460 | 2.549657 | 7.20158 | 95-97 |
| ACACA | 227355 | 2.5487518 | 7.1541133 | 6 |
| GCCGA | 97460 | 2.528921 | 7.3394322 | 90-94 |
| CTGCC | 101000 | 2.4536924 | 6.8762264 | 90-94 |
| CGACG | 91915 | 2.3850377 | 7.3880577 | 95-97 |
| GACTC | 138760 | 2.3147814 | 5.5873156 | 85-89 |
| TACAC | 209975 | 2.3023448 | 8.749009 | 5 |
| TGCCG | 89720 | 2.2770777 | 6.954616 | 90-94 |
| CCGAC | 91320 | 2.2682188 | 6.754716 | 95-97 |
| CACAT | 199430 | 2.1867206 | 5.0513906 | 7 |
| CTTTG | 197605 | 2.1654592 | 8.736624 | 9 |
| CTGAC | 123135 | 2.0541265 | 5.3376923 | 95-97 |
| TGACG | 114280 | 1.9916164 | 5.4466486 | 95-97 |
| GACGA | 105830 | 1.8856649 | 5.358346 | 95-97 |
| ATACA | 249420 | 1.8779365 | 5.505894 | 6 |
| GCTTT | 167860 | 1.8394977 | 5.4645047 | 1 |
| GCCAA | 101040 | 1.7232935 | 5.3444633 | 1 |
| GAGAG | 88475 | 1.6468959 | 5.036912 | 7 |
| TATAC | 213010 | 1.5686619 | 5.2259336 | 5 |
| GAGTC | 87770 | 1.5296131 | 8.0625725 | 9 |
| TTGAG | 129735 | 1.5185157 | 5.2447386 | 9 |
| GTGTA | 123765 | 1.4486386 | 10.180008 | 1 |
| AAGAC | 120515 | 1.4114124 | 5.442125 | 5 |
| CCATG | 81970 | 1.3674159 | 5.4120474 | 9 |
| TATGA | 174085 | 1.3393081 | 5.616243 | 4 |
| GTCTT | 120560 | 1.3211597 | 6.4744816 | 1 |
| TGAGT | 112310 | 1.3145604 | 6.6581726 | 8 |
| GACTT | 116810 | 1.3087369 | 6.7643533 | 7 |
| TCATA | 175730 | 1.2941222 | 5.7832108 | 2 |
| ATGAG | 106055 | 1.2691516 | 5.135825 | 7 |
| GTGTT | 110030 | 1.259659 | 5.680979 | 1 |
| GATTG | 105205 | 1.2313982 | 5.891747 | 7 |
| TGGAC | 66720 | 1.162764 | 7.48116 | 5 |
| GTTCT | 105700 | 1.158316 | 5.0445676 | 1 |
| ACATG | 98800 | 1.1317475 | 5.349507 | 8 |
| GTCCA | 66325 | 1.1064274 | 8.156651 | 1 |
| GGACT | 63255 | 1.1023775 | 7.631328 | 6 |
| ACACT | 98245 | 1.0772419 | 5.056618 | 6 |
| GAGTA | 84135 | 1.0068368 | 5.2127194 | 1 |
| GTATG | 85715 | 1.0032727 | 5.1266646 | 3 |
| AGACT | 87320 | 1.0002449 | 5.3992786 | 6 |
| GTATA | 100990 | 0.77695805 | 7.2136774 | 1 |