![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-123_TCCTGAGC-TATCCTCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1279393 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
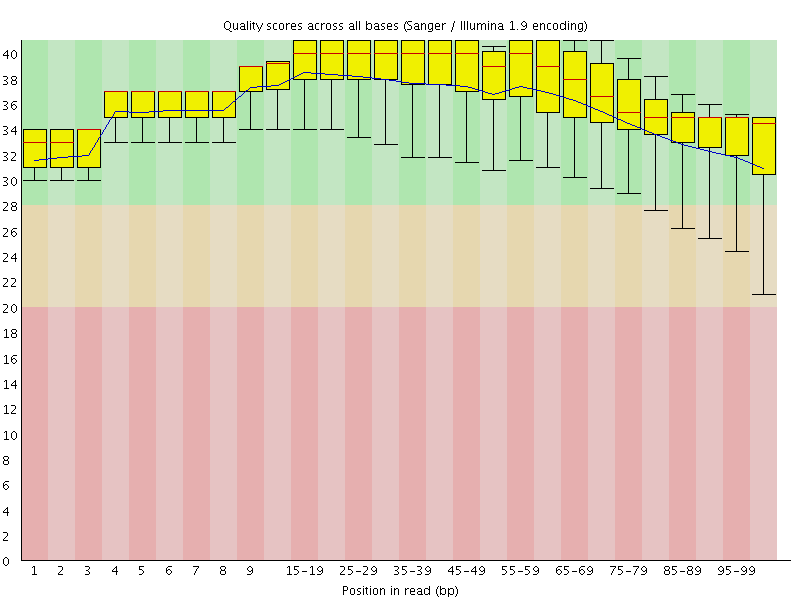
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
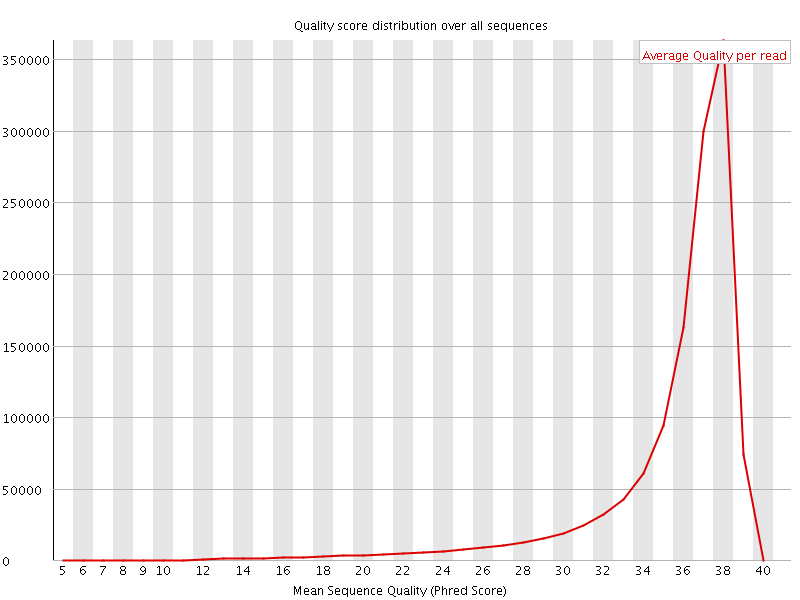
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
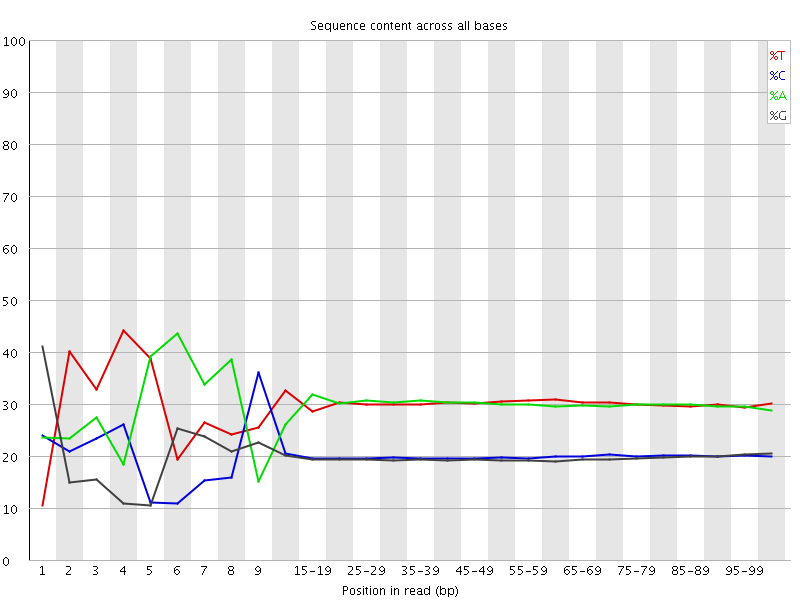
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
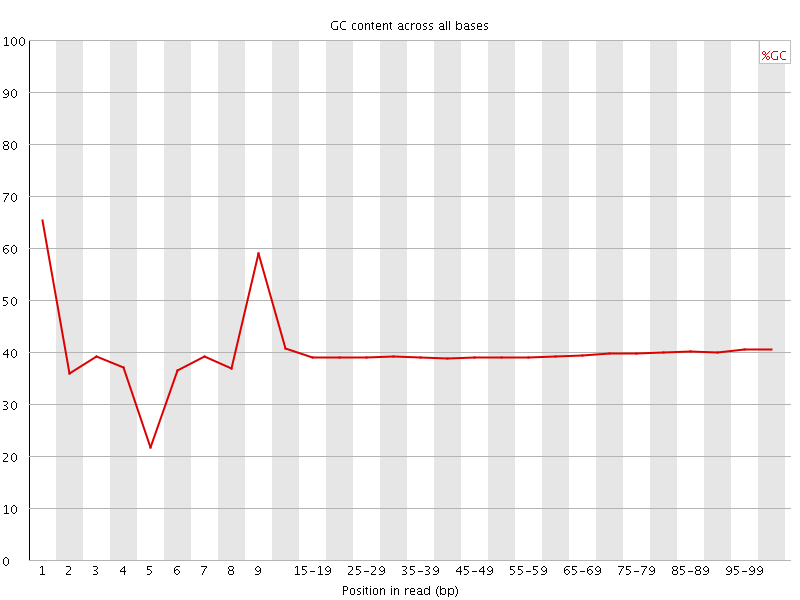
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
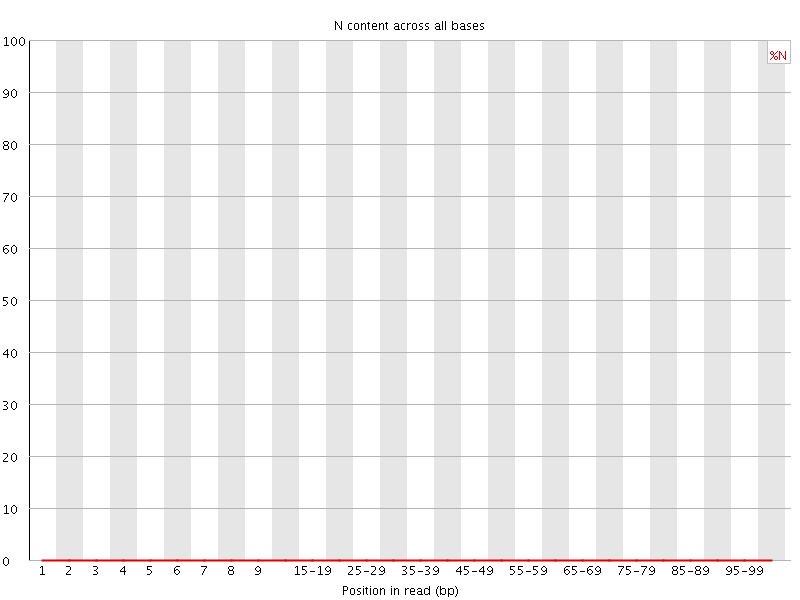
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
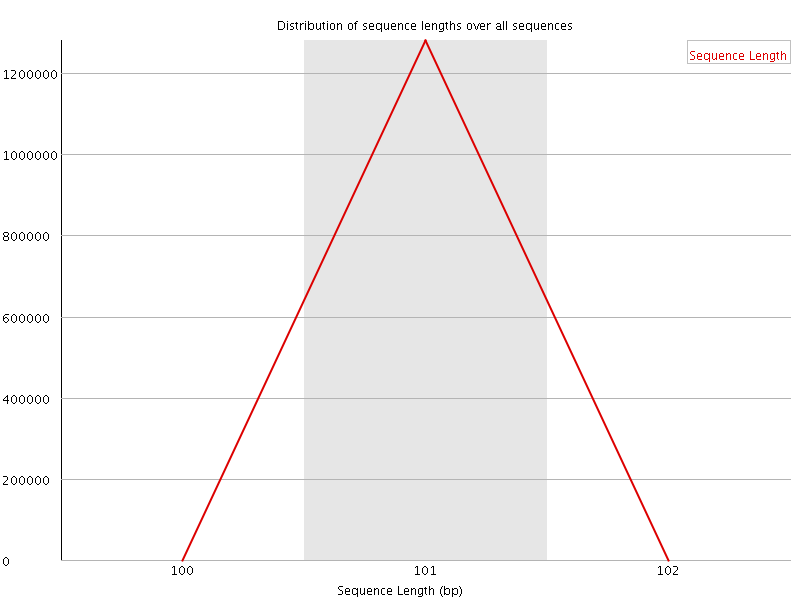
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
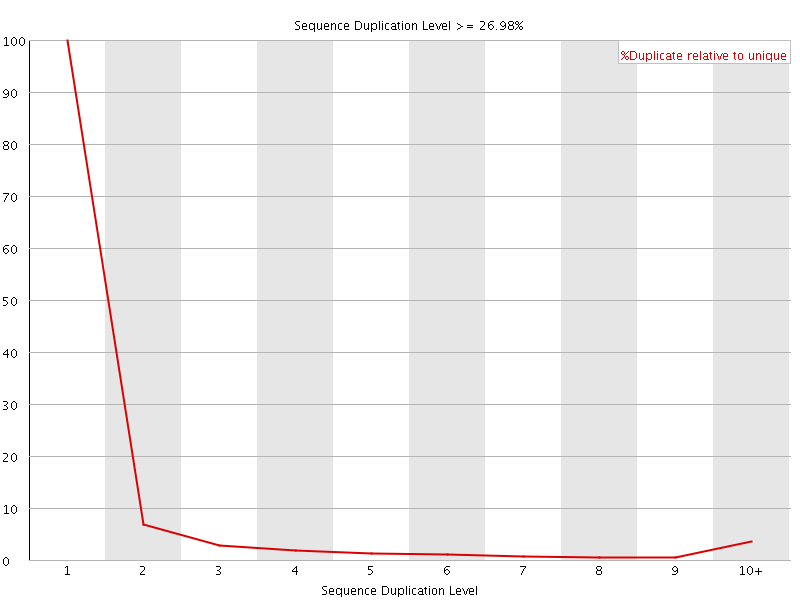
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
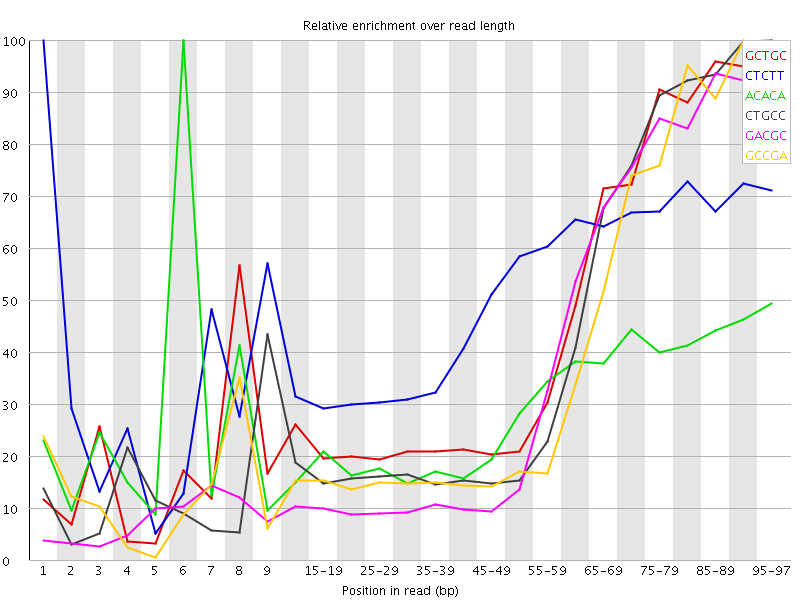
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 154070 | 2.617562 | 5.801302 | 95-97 |
| CTCTT | 350380 | 2.5388358 | 5.026409 | 1 |
| ACACA | 306460 | 2.2848916 | 7.7375126 | 6 |
| CTGCC | 131330 | 2.1974099 | 5.261618 | 95-97 |
| GACGC | 125310 | 2.1492991 | 5.557269 | 95-97 |
| GCCGA | 124850 | 2.1414092 | 5.5152893 | 90-94 |
| CGCTG | 125585 | 2.133618 | 5.397518 | 95-97 |
| CTTTG | 288570 | 2.1231384 | 8.117259 | 9 |
| CGACG | 119685 | 2.0528197 | 5.3742456 | 95-97 |
| TACAC | 269315 | 1.9889332 | 9.550906 | 5 |
| CCGAC | 117250 | 1.9805789 | 5.223616 | 90-94 |
| CACAT | 264525 | 1.9535583 | 5.3148293 | 7 |
| TGCCG | 114035 | 1.9373901 | 5.1664248 | 90-94 |
| GCTTT | 242700 | 1.7856523 | 5.096615 | 1 |
| GAGAG | 149105 | 1.7384305 | 5.840651 | 7 |
| TTGAG | 199405 | 1.5039285 | 5.500977 | 9 |
| GACTC | 132605 | 1.4853513 | 5.2691045 | 7 |
| GAGTC | 125200 | 1.4239851 | 7.3248725 | 9 |
| CCATG | 123740 | 1.3860517 | 5.709169 | 9 |
| AAGAC | 182070 | 1.3783579 | 5.7362685 | 5 |
| TATAC | 274835 | 1.3459275 | 5.7632203 | 5 |
| GTGTA | 174010 | 1.3123974 | 10.555126 | 1 |
| GTCTT | 178150 | 1.3107291 | 6.4278736 | 1 |
| GACTT | 172450 | 1.2809215 | 6.0911417 | 7 |
| TATGA | 254670 | 1.2663661 | 5.9290915 | 4 |
| TCATA | 258400 | 1.2654417 | 6.1337433 | 2 |
| TGAGT | 166865 | 1.2585093 | 5.076759 | 8 |
| GTGTT | 164915 | 1.2320238 | 5.483078 | 1 |
| GATTG | 162745 | 1.2274358 | 5.9800477 | 7 |
| GTTCT | 165875 | 1.2204164 | 5.5106254 | 1 |
| ACATG | 152705 | 1.1451036 | 5.516746 | 8 |
| TGGAC | 99810 | 1.1352073 | 7.227652 | 5 |
| GTCCA | 101105 | 1.1325097 | 8.150553 | 1 |
| ACACC | 101060 | 1.125509 | 5.109243 | 6 |
| ACACT | 142950 | 1.055708 | 5.210968 | 6 |
| GGACT | 91070 | 1.0358013 | 6.67394 | 6 |
| AGACT | 133125 | 0.99827695 | 5.287515 | 6 |
| TGTAT | 198280 | 0.97662586 | 5.058129 | 2 |
| GTATG | 127875 | 0.9644435 | 5.5939465 | 3 |
| GAGTA | 126155 | 0.96056724 | 5.152571 | 1 |
| CATAC | 127585 | 0.9422351 | 5.352171 | 3 |
| GTATA | 149865 | 0.7452152 | 6.852976 | 1 |