![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-123_TCCTGAGC-TATCCTCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1279393 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
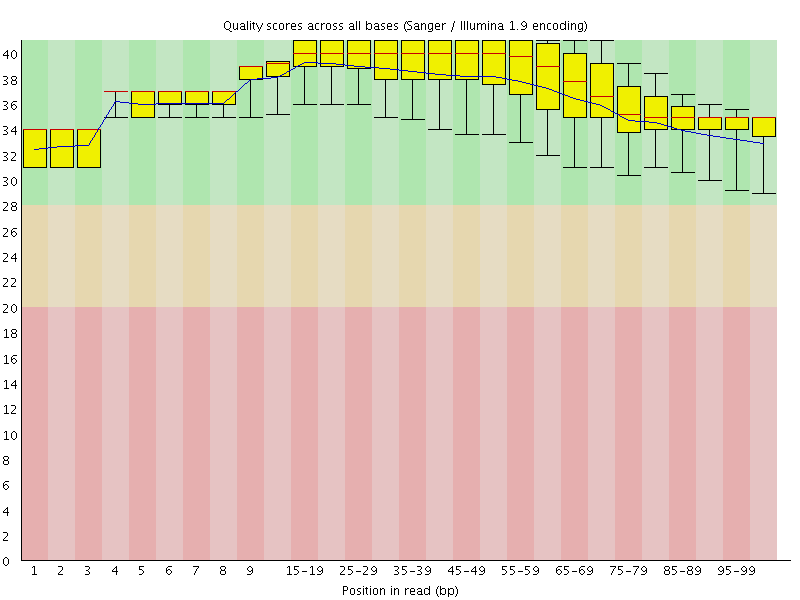
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
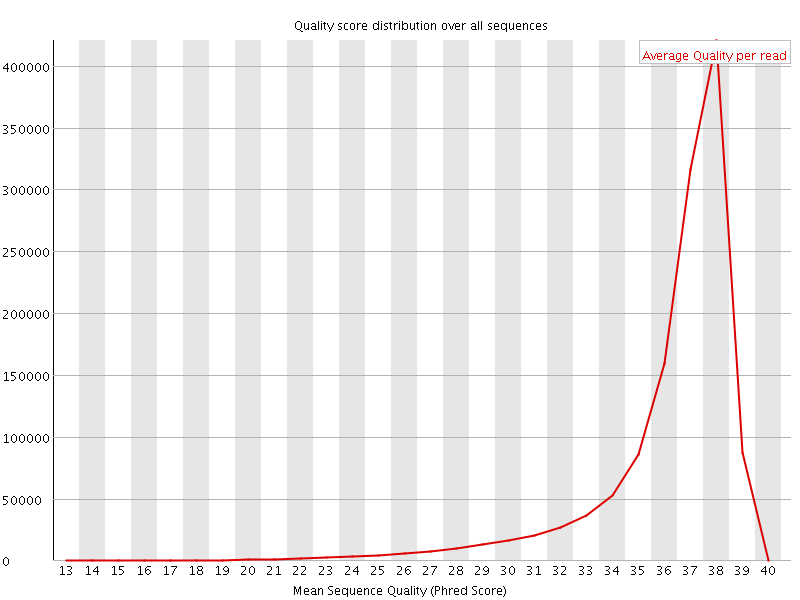
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
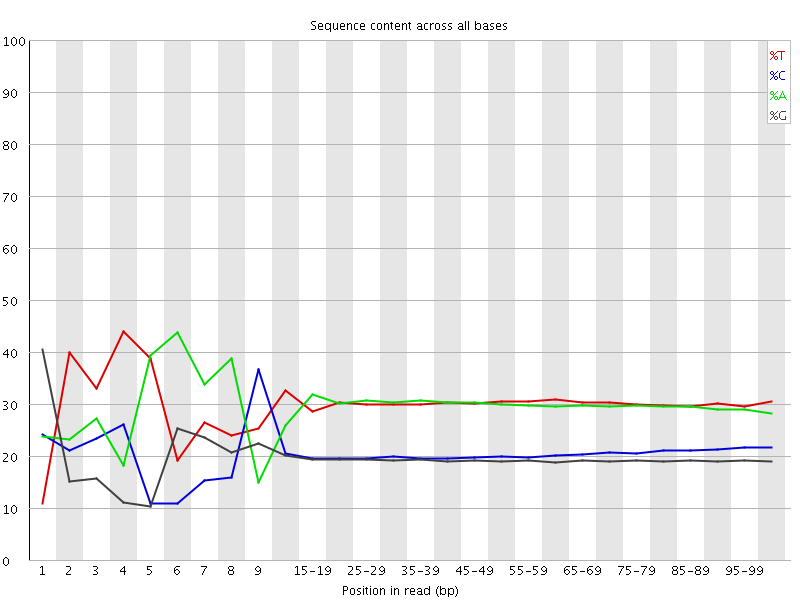
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
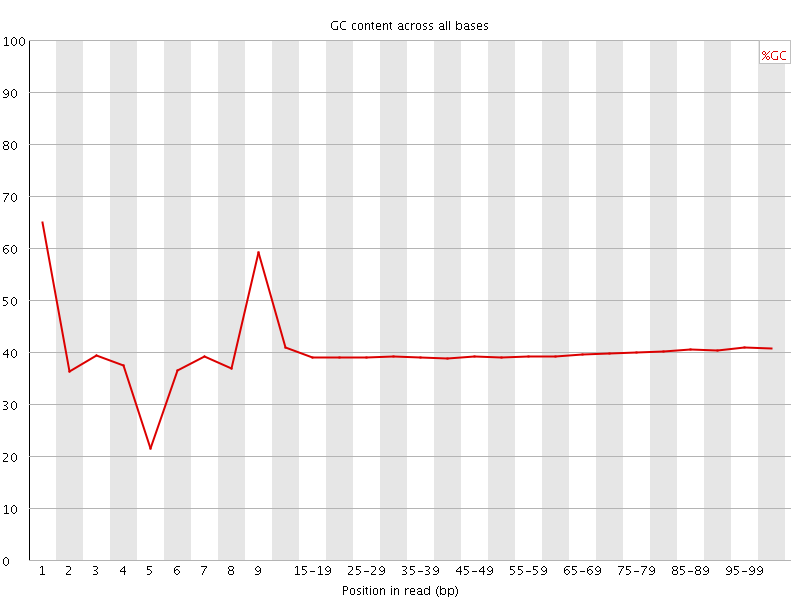
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
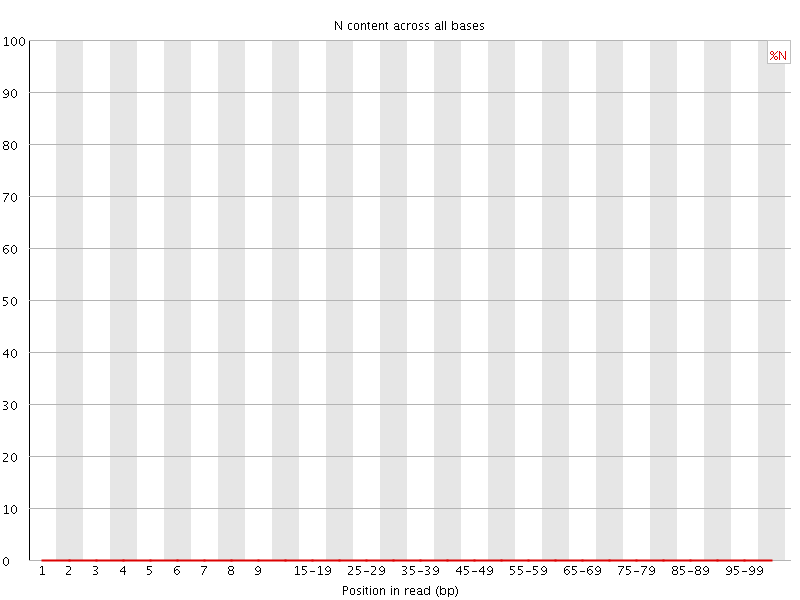
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
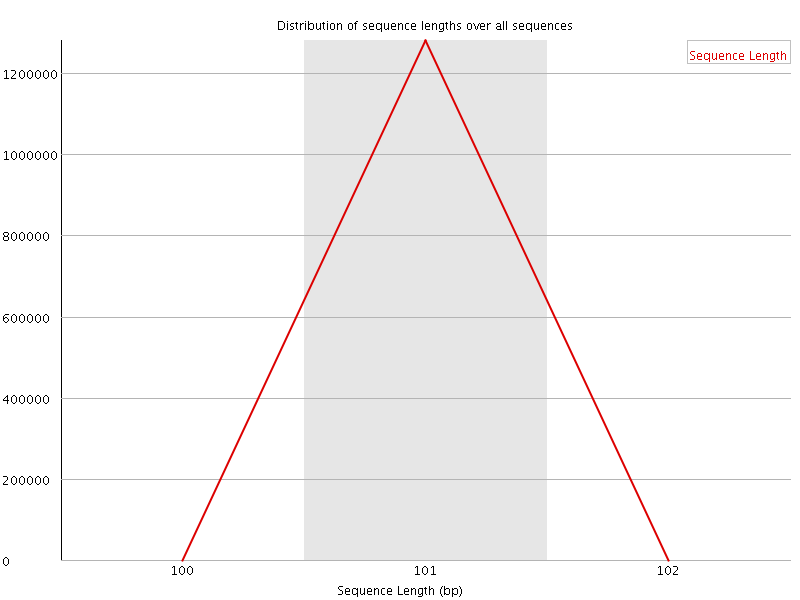
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
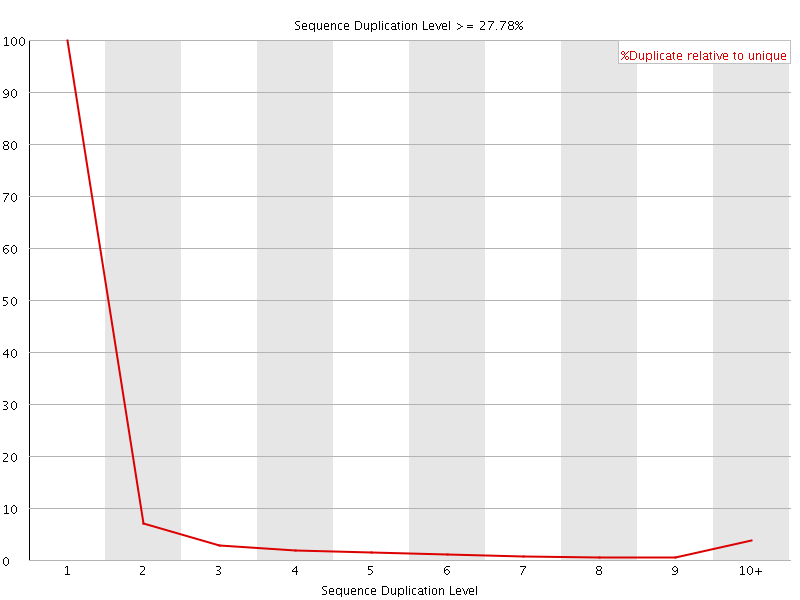
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
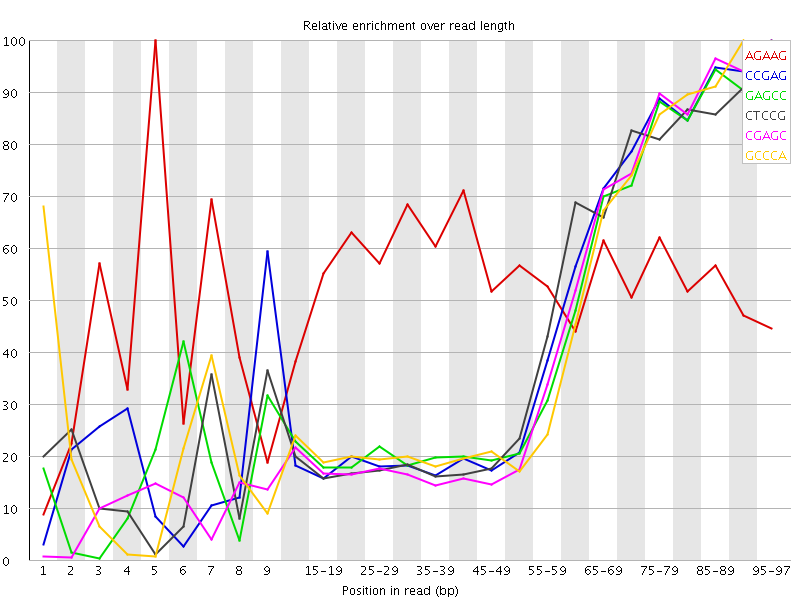
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 343285 | 2.7530246 | 5.074147 | 5 |
| CCGAG | 155890 | 2.6573308 | 5.9343615 | 95-97 |
| GAGCC | 153895 | 2.6233234 | 5.997727 | 95-97 |
| CTCCG | 160920 | 2.5722265 | 5.8075967 | 95-97 |
| CGAGC | 142480 | 2.4287412 | 5.689162 | 95-97 |
| GCCCA | 143020 | 2.31758 | 5.27697 | 90-94 |
| AGCCC | 141545 | 2.293678 | 5.243278 | 95-97 |
| ACACA | 306920 | 2.2243423 | 7.5440707 | 6 |
| GACTC | 193965 | 2.1266036 | 5.214702 | 7 |
| CTTTG | 288360 | 2.110011 | 8.1206875 | 9 |
| CCACG | 121735 | 1.9726653 | 5.0585365 | 90-94 |
| TACAC | 271065 | 1.9378101 | 9.465787 | 5 |
| CACAT | 265245 | 1.8962036 | 5.1921453 | 7 |
| AGAGA | 234255 | 1.8786426 | 5.074147 | 8 |
| GAGAG | 146835 | 1.8059689 | 5.98107 | 7 |
| TGGAG | 143370 | 1.7394031 | 5.1998587 | 5 |
| TGGCT | 140455 | 1.5979056 | 5.1297255 | 6 |
| TTGAG | 198465 | 1.5486804 | 5.569074 | 9 |
| AGACT | 195725 | 1.4718835 | 5.3670077 | 6 |
| GAGTC | 127150 | 1.4664557 | 7.006497 | 9 |
| CCATG | 127465 | 1.3975074 | 6.0067415 | 9 |
| GTCTT | 189205 | 1.3844663 | 6.102619 | 1 |
| AAGAC | 181145 | 1.3809953 | 5.8031363 | 5 |
| TATAC | 276680 | 1.3382608 | 6.008131 | 5 |
| TATGA | 253100 | 1.2877885 | 6.031545 | 4 |
| GATTG | 161905 | 1.2633921 | 6.1781917 | 7 |
| GACTT | 170240 | 1.262845 | 5.963075 | 7 |
| GTGTT | 163595 | 1.2592419 | 5.6021905 | 1 |
| TCATA | 260230 | 1.2586945 | 6.1494136 | 2 |
| GTTCT | 165885 | 1.2138273 | 5.456877 | 1 |
| AACAC | 163345 | 1.1838107 | 5.003613 | 5 |
| GTATG | 149715 | 1.1682699 | 5.9209247 | 3 |
| GTGTA | 149225 | 1.1644462 | 10.847847 | 1 |
| TGGAC | 99530 | 1.1479067 | 7.2413516 | 5 |
| ACATG | 152100 | 1.1438165 | 5.600356 | 8 |
| ACCAT | 158275 | 1.1314883 | 5.1366887 | 8 |
| GTCCA | 101540 | 1.1132696 | 7.474575 | 1 |
| ACACC | 102455 | 1.0825441 | 5.5070395 | 6 |
| GGACT | 91205 | 1.0518922 | 7.0232725 | 6 |
| ACACT | 142290 | 1.0172136 | 5.039639 | 6 |
| GAGTA | 125470 | 0.9925592 | 5.193657 | 1 |
| CATAC | 128710 | 0.92013186 | 5.1990776 | 3 |
| GTATA | 150345 | 0.7649647 | 7.033762 | 1 |
| ATACT | 156670 | 0.75778997 | 5.0044312 | 6 |