![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-122_TCCTGAGC-CTCTCTAT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1329196 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
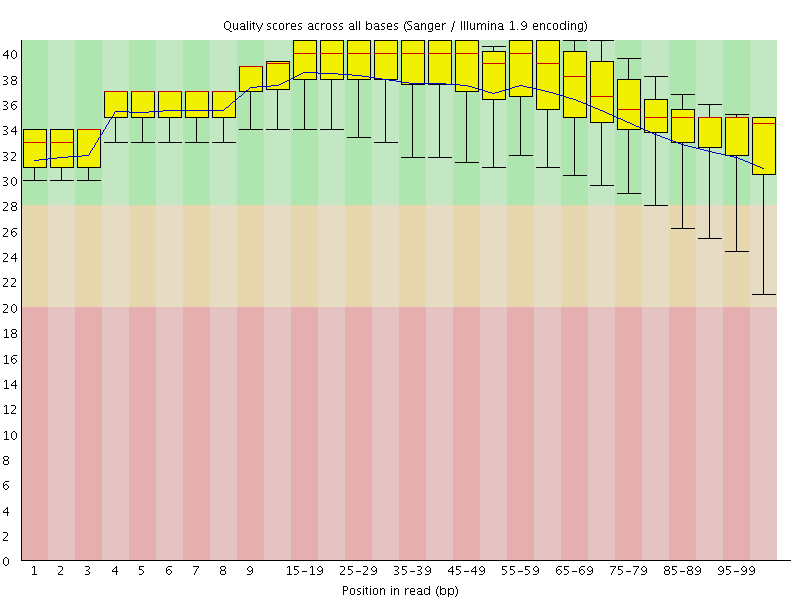
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
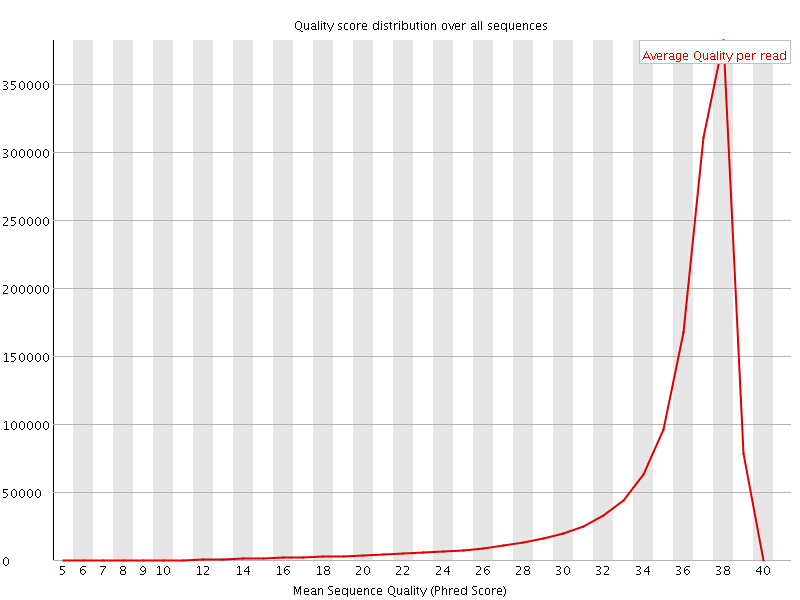
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
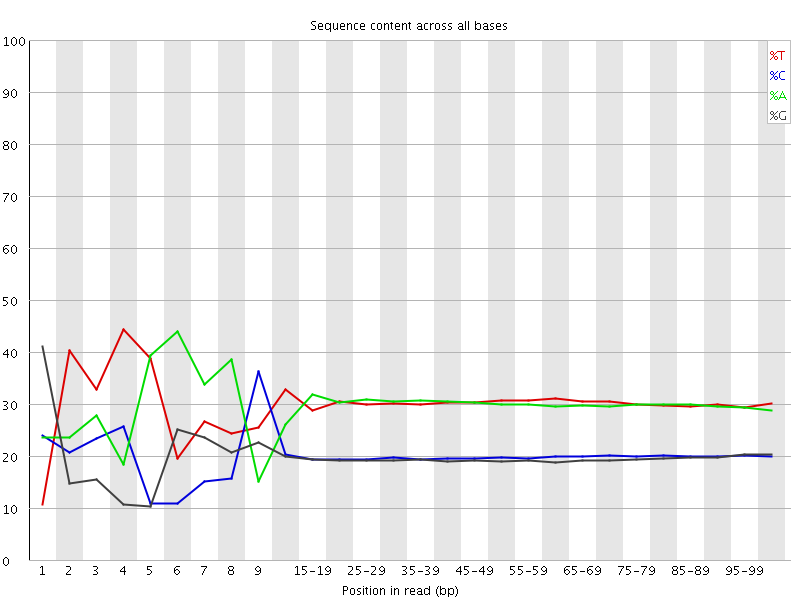
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
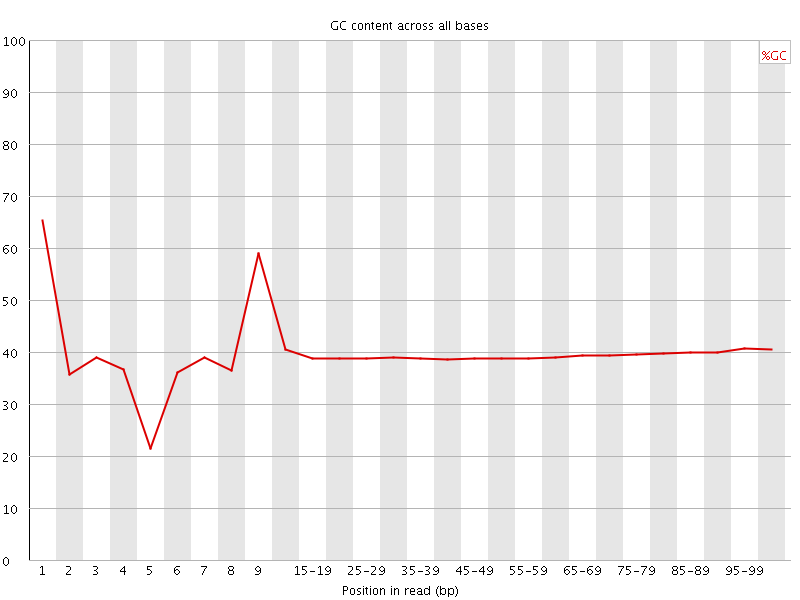
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
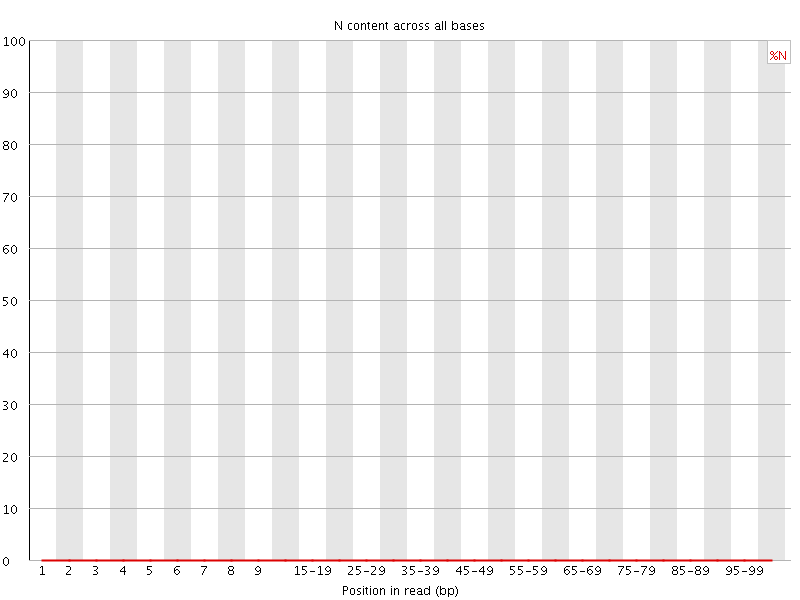
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
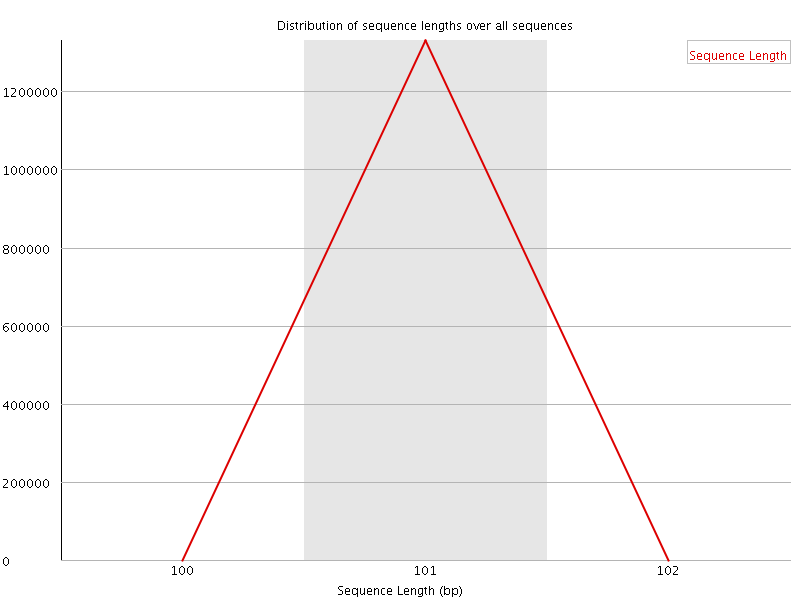
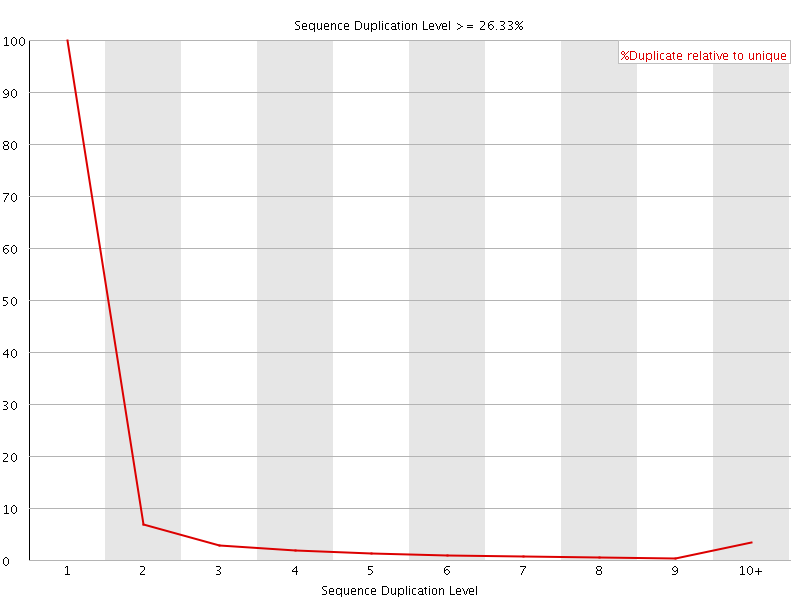
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
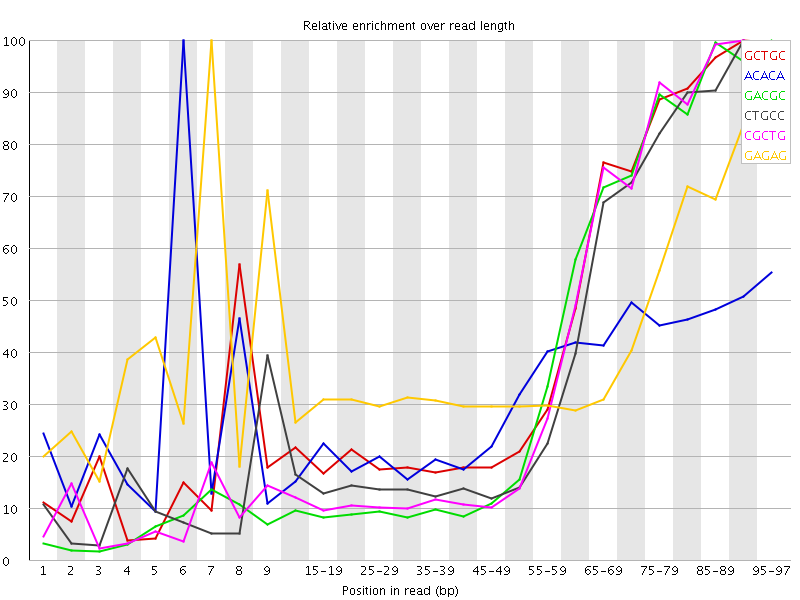
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 168065 | 2.7946467 | 6.2727466 | 90-94 |
| ACACA | 339500 | 2.4277492 | 7.4731183 | 6 |
| GACGC | 142190 | 2.3854074 | 6.0051184 | 95-97 |
| CTGCC | 145655 | 2.378801 | 6.008773 | 90-94 |
| CGCTG | 142585 | 2.370956 | 5.9178987 | 90-94 |
| GAGAG | 205460 | 2.3328662 | 5.63849 | 7 |
| GCCGA | 138955 | 2.3311365 | 6.2780666 | 90-94 |
| CGACG | 133825 | 2.2450747 | 6.205832 | 95-97 |
| CTTTG | 308230 | 2.1853657 | 8.953553 | 9 |
| CCGAC | 131010 | 2.1586442 | 5.8926706 | 95-97 |
| TACAC | 302100 | 2.141267 | 9.643507 | 5 |
| TGCCG | 127840 | 2.1257706 | 5.9566092 | 90-94 |
| CACAT | 292265 | 2.0715568 | 5.283166 | 7 |
| GCTTT | 258390 | 1.8319979 | 5.4027796 | 1 |
| ATACA | 364220 | 1.7160739 | 5.0038795 | 6 |
| GCCAA | 154590 | 1.6932074 | 5.153382 | 1 |
| CACCA | 149570 | 1.6090012 | 5.164727 | 7 |
| TTGAG | 209365 | 1.524804 | 5.848918 | 9 |
| GACTC | 140365 | 1.5238551 | 5.4649043 | 7 |
| GAGTC | 133585 | 1.4765884 | 7.8692446 | 9 |
| TATAC | 308040 | 1.4385848 | 5.7242107 | 5 |
| CCATG | 129975 | 1.4110574 | 6.0493693 | 9 |
| AAGAC | 193115 | 1.4060376 | 5.809367 | 5 |
| GTGTA | 188300 | 1.3713877 | 11.071355 | 1 |
| GTCTT | 188700 | 1.3378923 | 6.4138665 | 1 |
| GACTT | 183660 | 1.3137348 | 6.95864 | 7 |
| TATGA | 272340 | 1.294961 | 6.425522 | 4 |
| ACTCA | 182385 | 1.292734 | 5.0701475 | 8 |
| TGAGT | 177110 | 1.2898912 | 5.4957643 | 8 |
| TCATA | 275465 | 1.2864555 | 6.3222284 | 2 |
| GTGTT | 176740 | 1.2758539 | 6.0716534 | 1 |
| ATGAG | 171250 | 1.2583008 | 5.163244 | 7 |
| GATTG | 171975 | 1.252493 | 6.5552354 | 7 |
| GTTCT | 172780 | 1.2250185 | 5.5506587 | 1 |
| TGGAC | 105465 | 1.1657625 | 8.138709 | 5 |
| ACCAT | 164215 | 1.1639462 | 5.321077 | 8 |
| ACACC | 107960 | 1.1613811 | 5.467184 | 6 |
| GTCCA | 104805 | 1.1378024 | 8.446583 | 1 |
| ACATG | 156990 | 1.1329454 | 5.5717106 | 8 |
| GGACT | 98470 | 1.0884429 | 7.9333024 | 6 |
| ACACT | 153000 | 1.084455 | 5.2727356 | 6 |
| AGACT | 140260 | 1.0122105 | 5.5924783 | 6 |
| GTATG | 136780 | 0.996168 | 6.008919 | 3 |
| TGTAT | 208300 | 0.98172665 | 5.118438 | 2 |
| GAGTA | 132615 | 0.97442085 | 5.146511 | 1 |
| CATAC | 134995 | 0.95683664 | 5.3563576 | 3 |
| GTATA | 158785 | 0.75501347 | 7.274427 | 1 |