![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-122_TCCTGAGC-CTCTCTAT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1329196 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
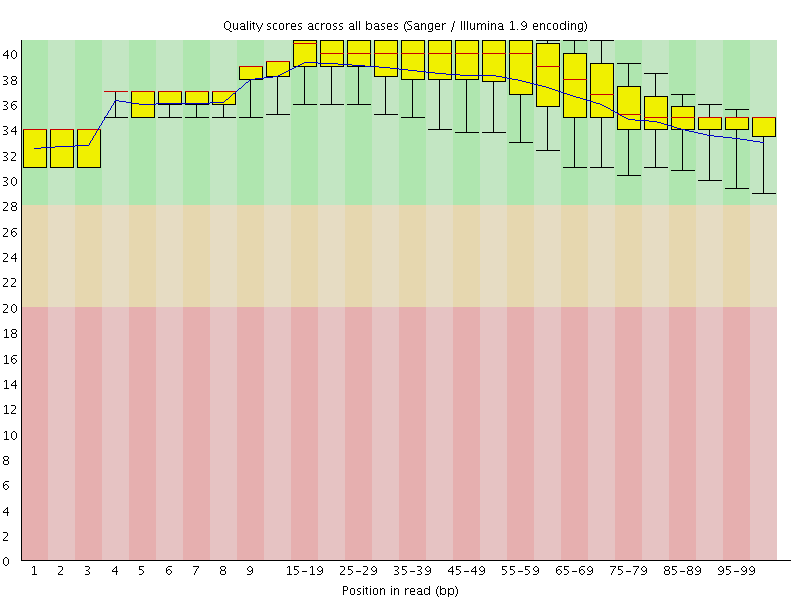
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
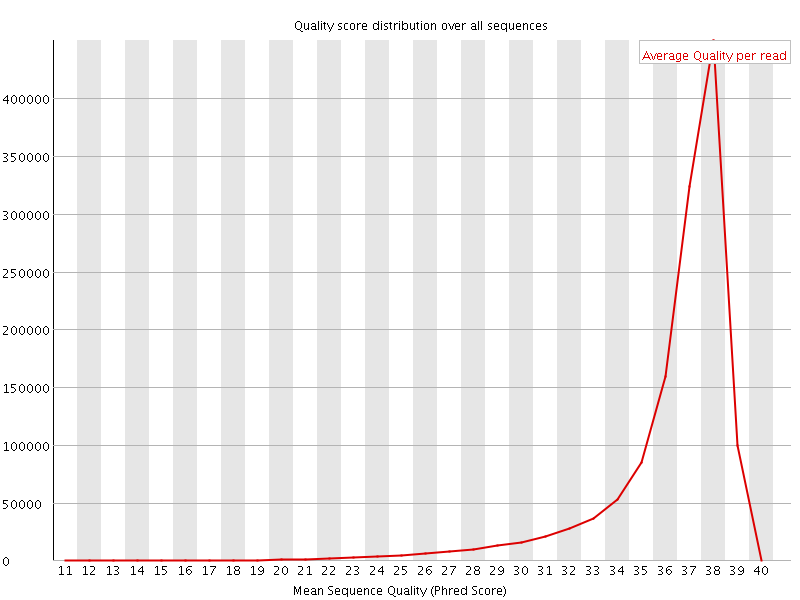
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
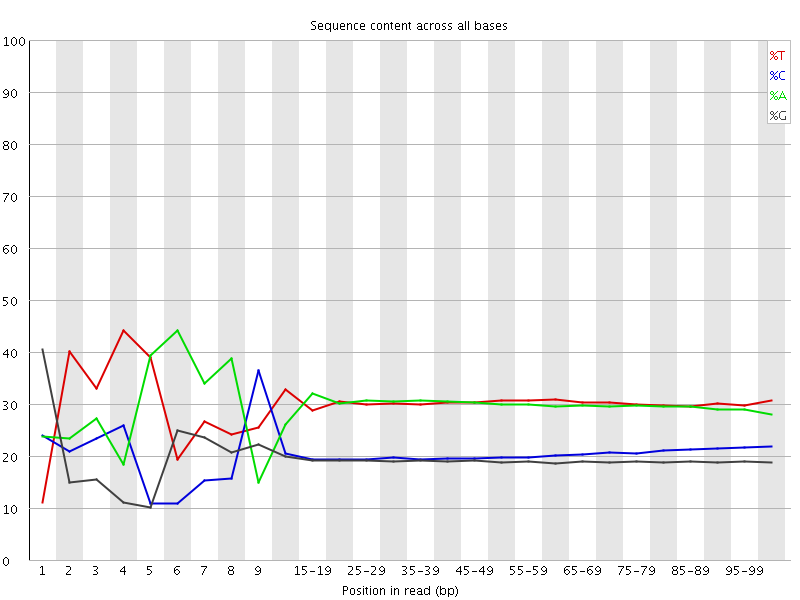
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
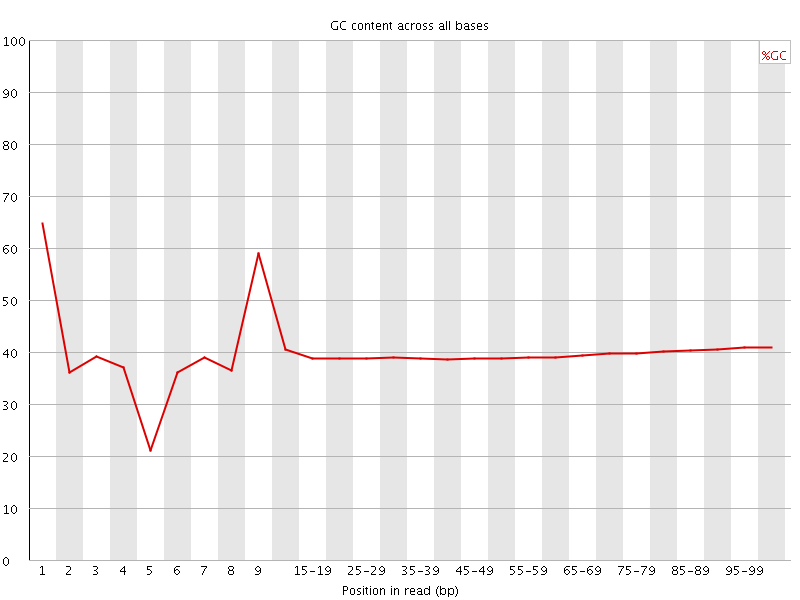
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
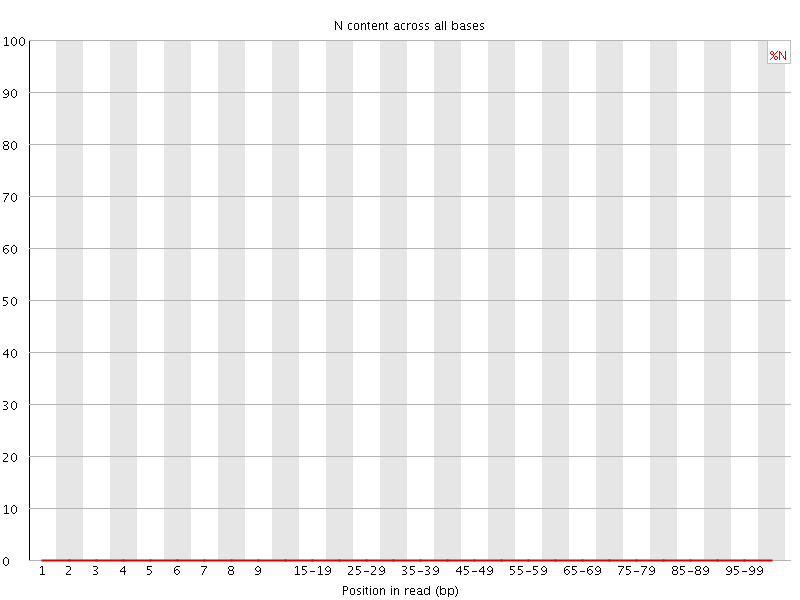
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
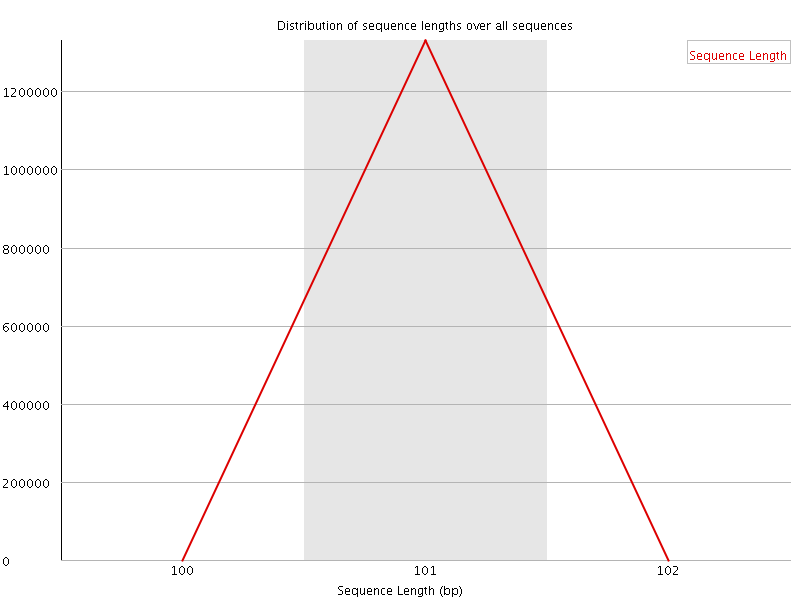
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
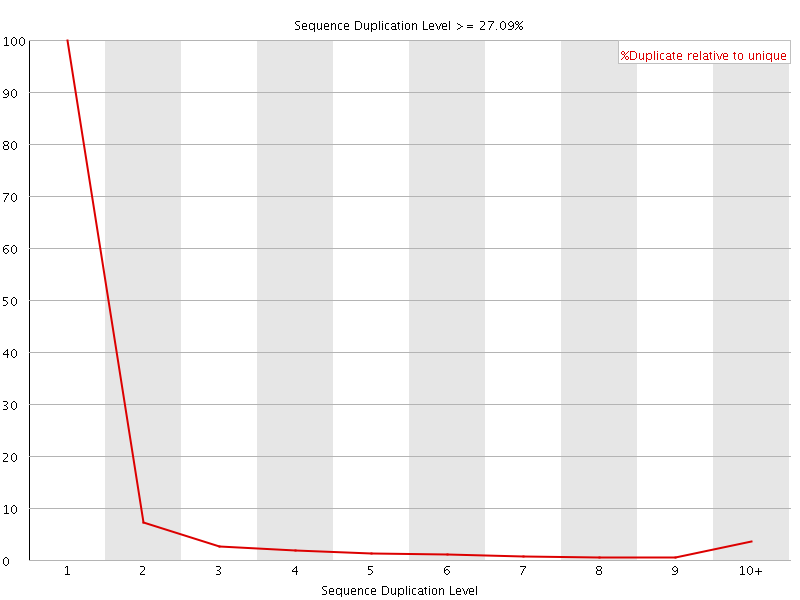
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
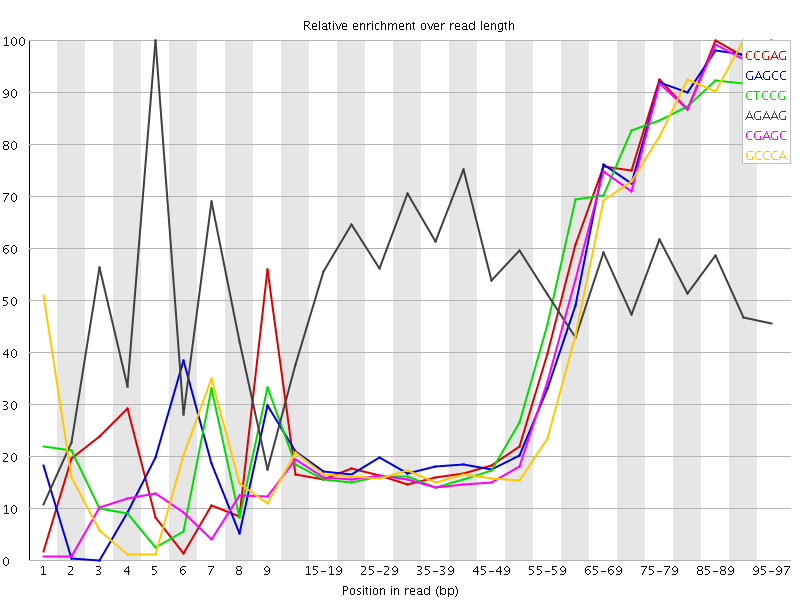
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 172460 | 2.8775933 | 6.386697 | 85-89 |
| GAGCC | 169930 | 2.835379 | 6.3697743 | 95-97 |
| CTCCG | 179755 | 2.7884862 | 6.2228947 | 95-97 |
| AGAAG | 356235 | 2.7783601 | 5.089736 | 5 |
| CGAGC | 160160 | 2.672361 | 6.24842 | 95-97 |
| GCCCA | 156610 | 2.4702861 | 5.9075065 | 90-94 |
| AGCCC | 155705 | 2.456011 | 5.6933556 | 90-94 |
| ACACA | 341405 | 2.3795521 | 7.775713 | 6 |
| CAAAG | 316630 | 2.3344831 | 5.0331464 | 4 |
| CCCAC | 151995 | 2.2664375 | 5.616398 | 90-94 |
| GACTC | 212890 | 2.2578664 | 5.851719 | 7 |
| CATCT | 328840 | 2.2168155 | 5.028283 | 95-97 |
| CTTTG | 309400 | 2.1698925 | 8.820356 | 9 |
| CCACG | 135140 | 2.131629 | 5.7162995 | 90-94 |
| TACAC | 302960 | 2.0766838 | 10.066595 | 5 |
| CACAT | 293850 | 2.0142379 | 5.440415 | 7 |
| GCTTT | 259015 | 1.8165312 | 5.1862254 | 1 |
| GAGAG | 148735 | 1.7948287 | 5.944315 | 7 |
| TGGAG | 148005 | 1.7564912 | 5.253377 | 5 |
| TGGCT | 150920 | 1.6651862 | 5.2104254 | 6 |
| TTGAG | 209825 | 1.5828097 | 5.7530537 | 9 |
| AGACT | 214280 | 1.5537461 | 5.5264874 | 6 |
| CACCA | 149690 | 1.5260286 | 5.081141 | 7 |
| GAGTC | 134665 | 1.5108141 | 7.72945 | 9 |
| ATGGA | 196955 | 1.5107018 | 5.0650887 | 4 |
| TATAC | 309175 | 1.4249666 | 5.9060307 | 5 |
| AAGAC | 191975 | 1.4154135 | 6.0983996 | 5 |
| GTCTT | 201710 | 1.4146382 | 6.441122 | 1 |
| CATGA | 194710 | 1.4118439 | 5.1081343 | 9 |
| CCATG | 130425 | 1.3832599 | 6.09854 | 9 |
| TGAGT | 178490 | 1.3464351 | 5.460464 | 8 |
| TATGA | 271285 | 1.322633 | 6.387004 | 4 |
| ATGAG | 172030 | 1.3195199 | 5.087402 | 7 |
| GTGTT | 175800 | 1.3042177 | 6.029327 | 1 |
| GACTT | 182830 | 1.3037839 | 6.831929 | 7 |
| GATTG | 170990 | 1.2898588 | 6.312632 | 7 |
| TCATA | 278340 | 1.2828503 | 6.53717 | 2 |
| ACTCA | 182990 | 1.254332 | 5.0083723 | 8 |
| GTATG | 162910 | 1.2289076 | 5.9542093 | 3 |
| GTGTA | 161860 | 1.220987 | 11.474918 | 1 |
| GTTCT | 172285 | 1.2082739 | 5.0535936 | 1 |
| TGGAC | 104980 | 1.1777765 | 7.9796643 | 5 |
| ACATG | 160100 | 1.1608865 | 5.776094 | 8 |
| GTCCA | 106625 | 1.1308422 | 8.053753 | 1 |
| ACCAT | 164625 | 1.1284462 | 5.24101 | 8 |
| ACACC | 109705 | 1.1183978 | 5.5853014 | 6 |
| GGACT | 97755 | 1.0967188 | 7.7457685 | 6 |
| ACACT | 154815 | 1.0612023 | 5.2144237 | 6 |
| TGTAT | 212790 | 1.0202924 | 5.1942177 | 2 |
| GAGTA | 132370 | 1.0153161 | 5.2890162 | 1 |
| CATAC | 135750 | 0.93051845 | 5.5799985 | 3 |
| GTATA | 160090 | 0.78050894 | 7.0948505 | 1 |