![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-120_AGGCAGAA-CTAAGCCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1168013 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
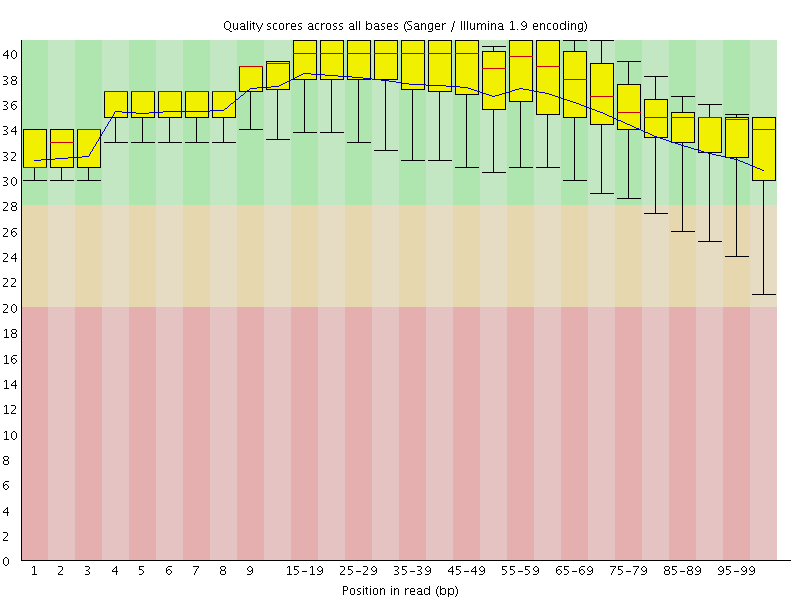
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
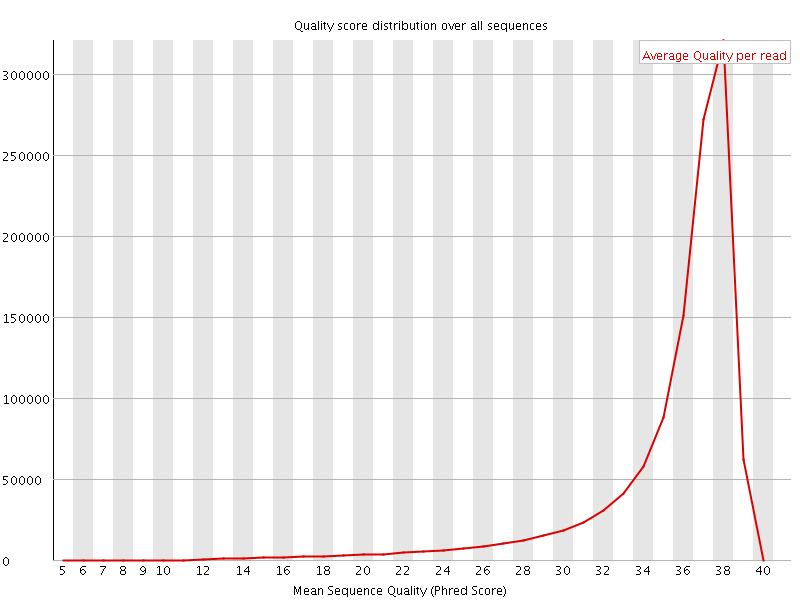
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
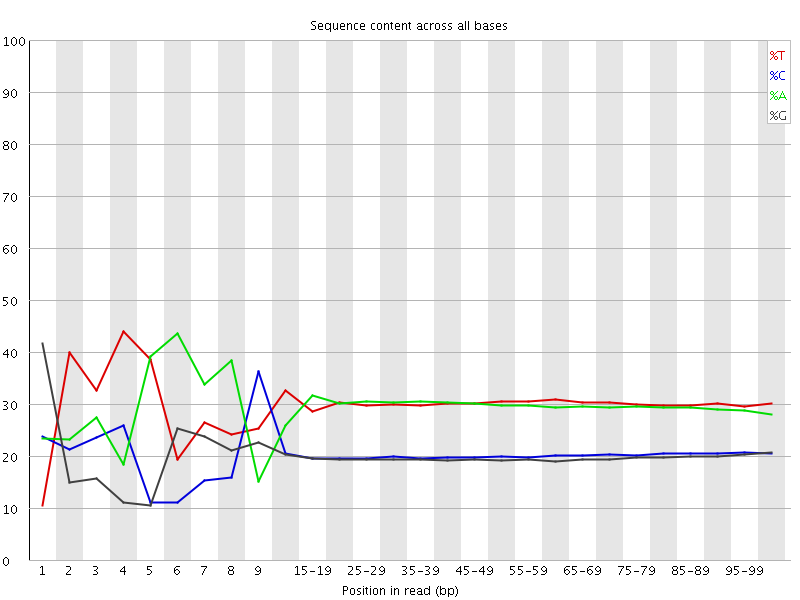
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
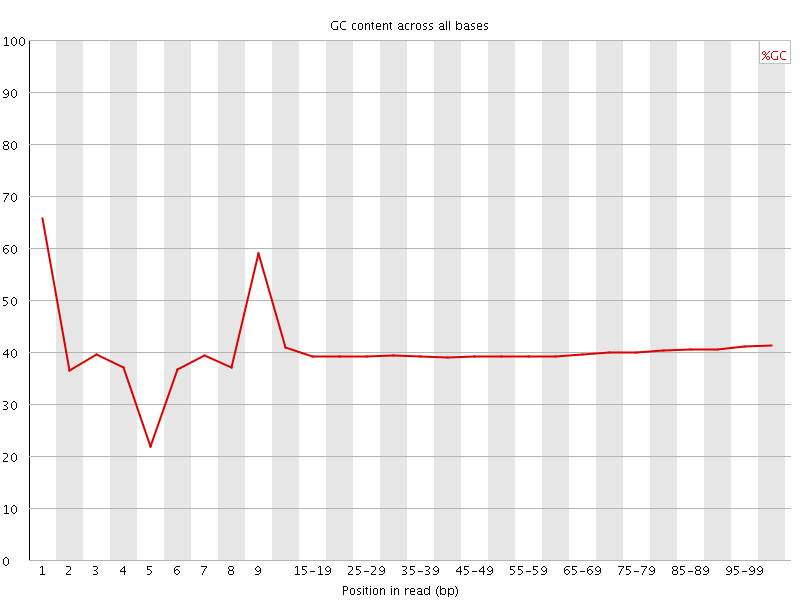
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
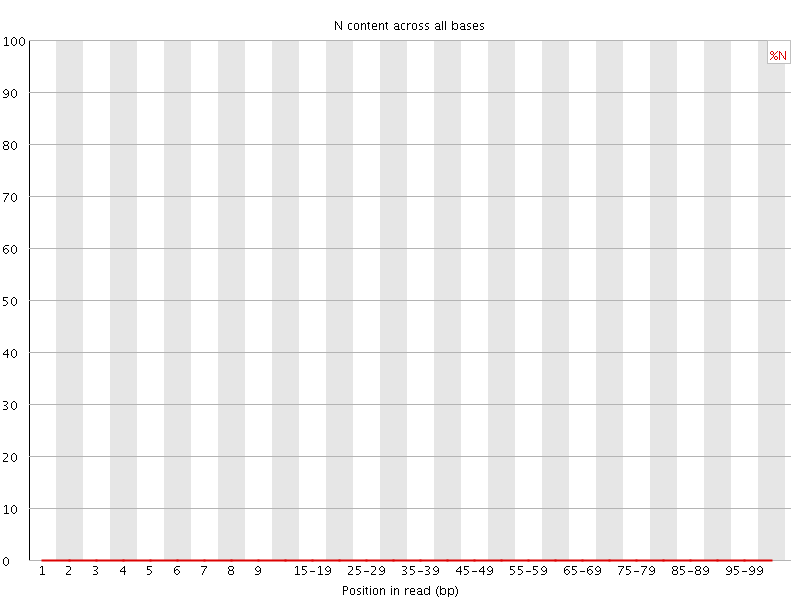
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
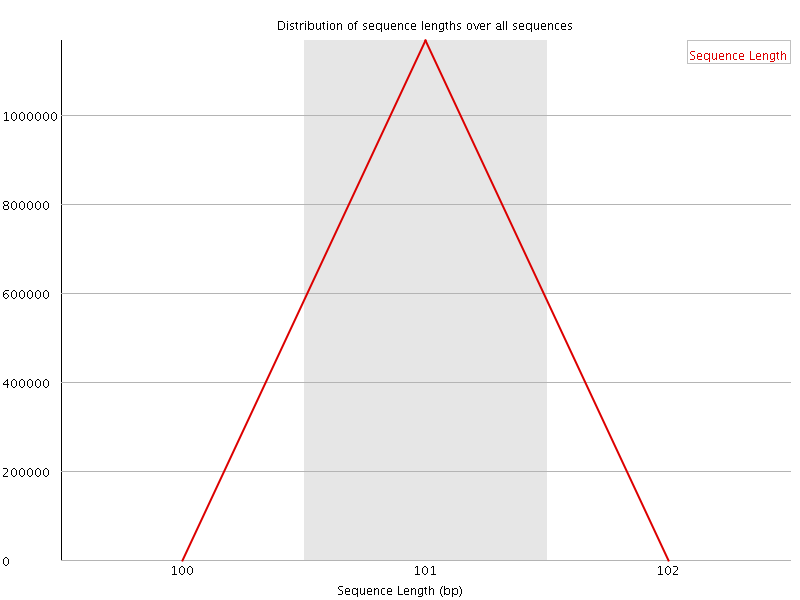
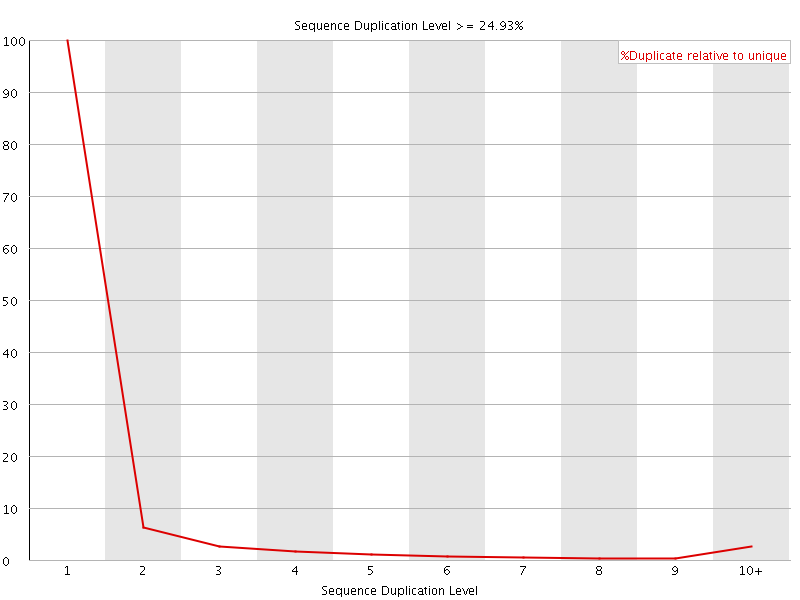
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
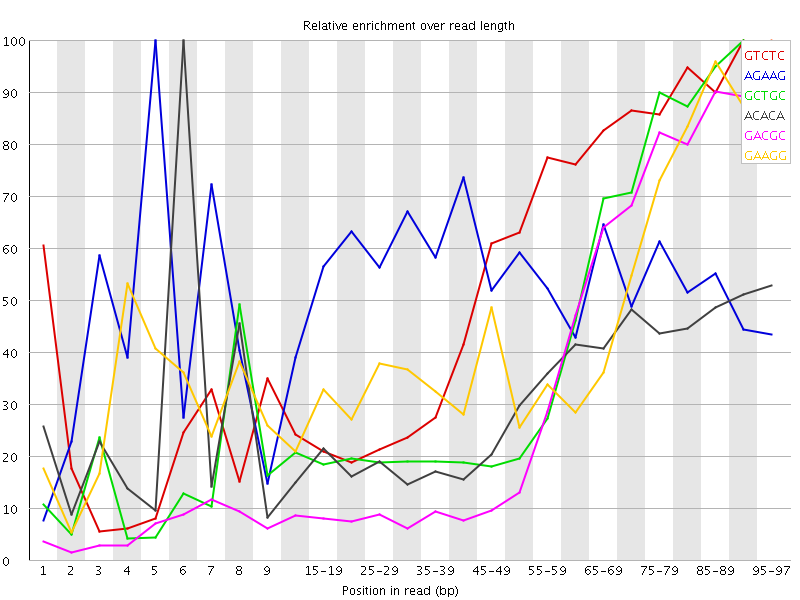
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 242260 | 2.8855095 | 5.1113234 | 90-94 |
| AGAAG | 317490 | 2.725161 | 5.0295267 | 5 |
| GCTGC | 148630 | 2.7074761 | 6.2109914 | 90-94 |
| ACACA | 298870 | 2.4572687 | 7.8506603 | 6 |
| GACGC | 127295 | 2.3611226 | 6.5317664 | 95-97 |
| GAAGG | 182990 | 2.3591628 | 5.1027884 | 95-97 |
| CAAAG | 276710 | 2.324562 | 5.171016 | 4 |
| CGCTG | 125945 | 2.2942417 | 5.967099 | 95-97 |
| GCCGA | 123535 | 2.2913802 | 6.19112 | 90-94 |
| CTGCC | 128215 | 2.2858677 | 5.8695054 | 90-94 |
| CGACG | 122420 | 2.2706988 | 6.2678566 | 95-97 |
| CTTTG | 271940 | 2.1639264 | 8.578292 | 9 |
| TACAC | 267470 | 2.1597147 | 9.917062 | 5 |
| CCGAC | 117205 | 2.1276848 | 5.9096336 | 90-94 |
| CACAT | 256605 | 2.071984 | 5.5408607 | 7 |
| TGCCG | 113605 | 2.0694535 | 5.728533 | 95-97 |
| GCTTT | 229635 | 1.8272897 | 5.8129425 | 1 |
| ATACA | 317560 | 1.7443138 | 5.058409 | 6 |
| GAGAG | 134665 | 1.7361424 | 5.5581703 | 7 |
| GCCAA | 138455 | 1.7098082 | 5.3432484 | 1 |
| CACCA | 132800 | 1.6050587 | 5.1051726 | 7 |
| TGGCT | 131625 | 1.601862 | 5.234805 | 6 |
| GACTC | 125405 | 1.5209137 | 5.540387 | 7 |
| TTGAG | 182560 | 1.5113666 | 5.7651515 | 9 |
| TATAC | 271725 | 1.4658161 | 5.7906766 | 5 |
| GAGTC | 118170 | 1.4643433 | 7.0369525 | 9 |
| AAGAC | 168915 | 1.4190066 | 5.7007494 | 5 |
| GTGTA | 170290 | 1.4097865 | 10.862586 | 1 |
| CATGA | 168895 | 1.3934264 | 5.097165 | 9 |
| CCATG | 114740 | 1.3915685 | 6.0696197 | 9 |
| GTCTT | 167920 | 1.3362008 | 6.511576 | 1 |
| GACTT | 163595 | 1.3255262 | 6.573758 | 7 |
| TATGA | 235510 | 1.298091 | 6.2267766 | 4 |
| TGAGT | 156040 | 1.2918144 | 5.211252 | 8 |
| ACTCA | 158310 | 1.2782907 | 5.0240173 | 8 |
| AACAC | 155420 | 1.2778422 | 5.407892 | 5 |
| TCATA | 236780 | 1.2773058 | 6.3897834 | 2 |
| GTGTT | 155245 | 1.2622137 | 6.069538 | 1 |
| GATTG | 147620 | 1.2221076 | 6.0864344 | 7 |
| GTTCT | 149215 | 1.1873584 | 5.2802825 | 1 |
| ACACC | 97365 | 1.176781 | 5.550511 | 6 |
| ACCAT | 143350 | 1.1574947 | 5.067092 | 8 |
| ACATG | 138365 | 1.1415464 | 5.7134533 | 8 |
| TGGAC | 91445 | 1.1331716 | 7.4413953 | 5 |
| GTCCA | 91430 | 1.1088644 | 7.6418877 | 1 |
| ACACT | 134205 | 1.0836525 | 5.638635 | 6 |
| GGACT | 84530 | 1.0474819 | 6.916735 | 6 |
| AGACT | 126180 | 1.0410169 | 5.725284 | 6 |
| GAGTA | 118165 | 0.99609804 | 5.237983 | 1 |
| TGTAT | 182010 | 0.9852401 | 5.007121 | 2 |
| GTATG | 117195 | 0.970227 | 5.5415134 | 3 |
| CATAC | 114655 | 0.9257938 | 5.3382897 | 3 |
| GTATA | 139490 | 0.76884514 | 7.149225 | 1 |