![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-120_AGGCAGAA-CTAAGCCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1168013 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
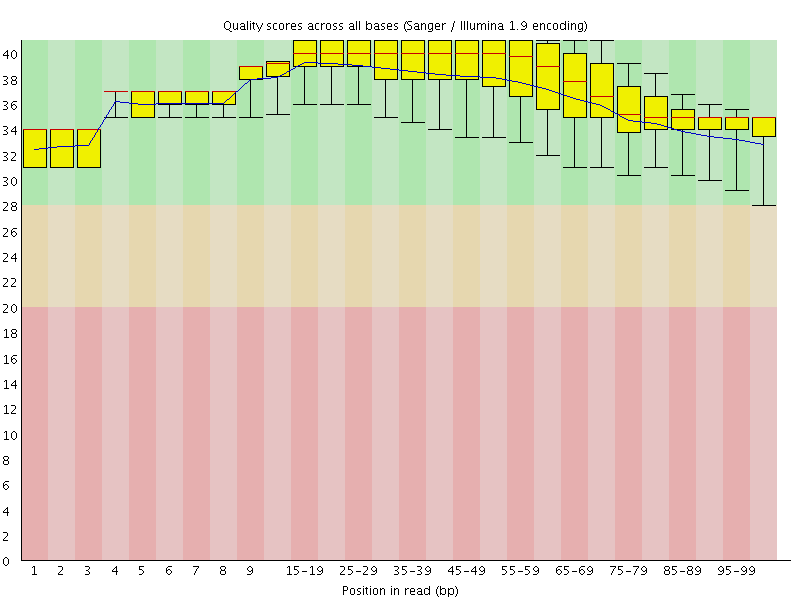
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
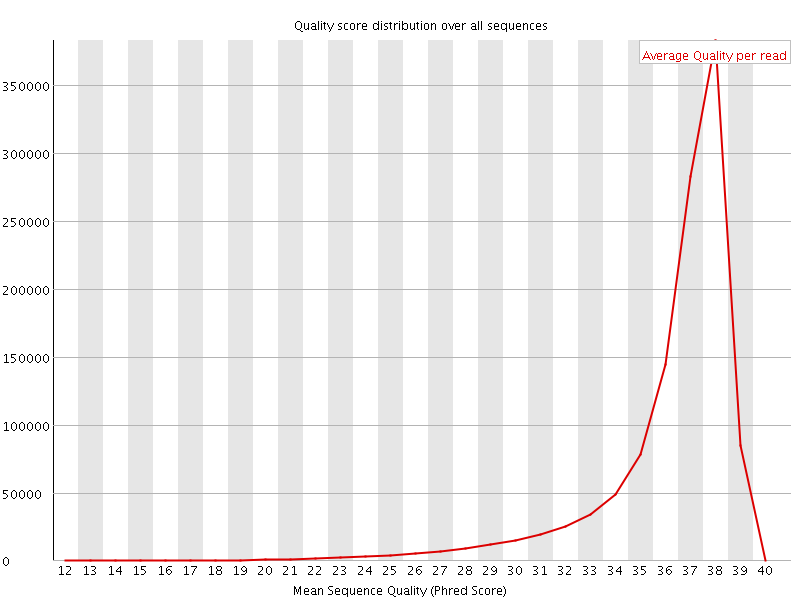
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
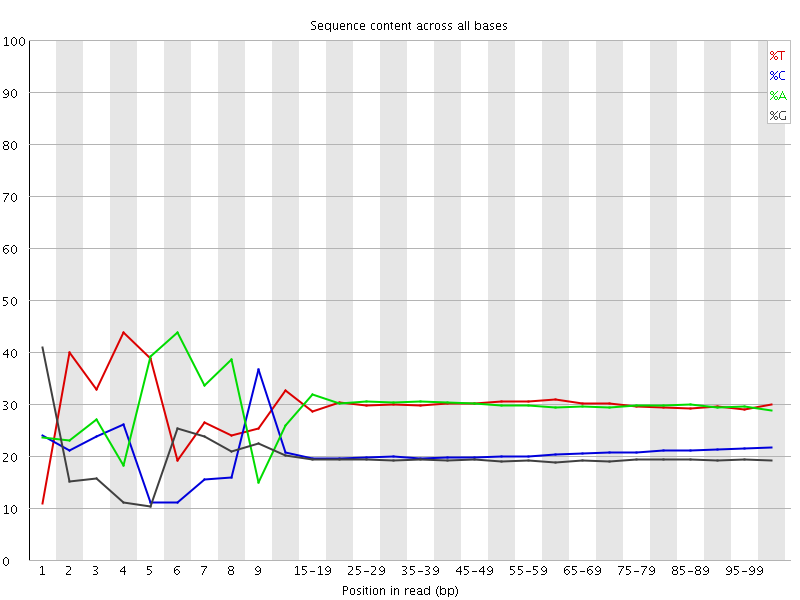
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
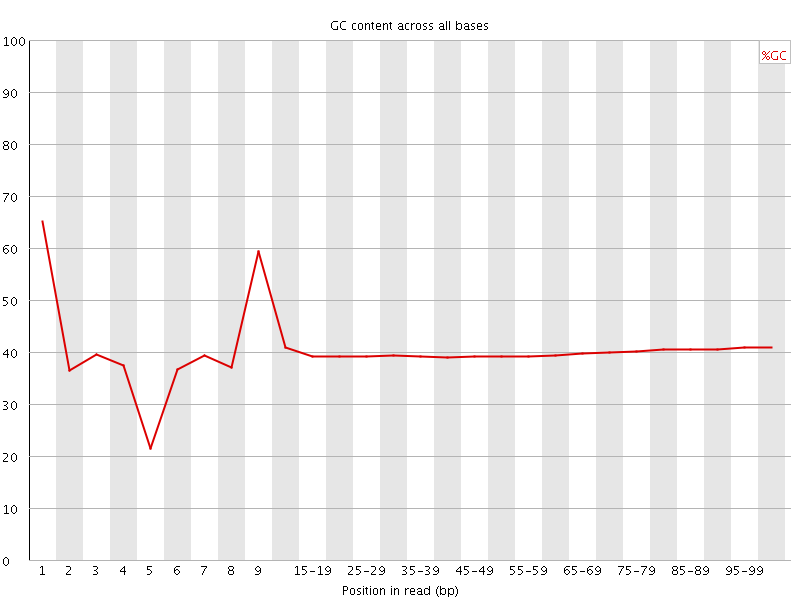
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
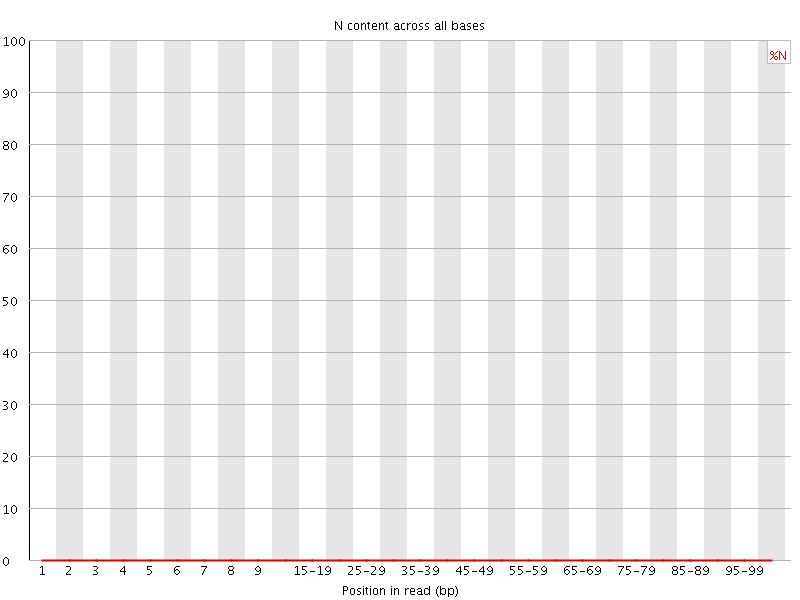
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
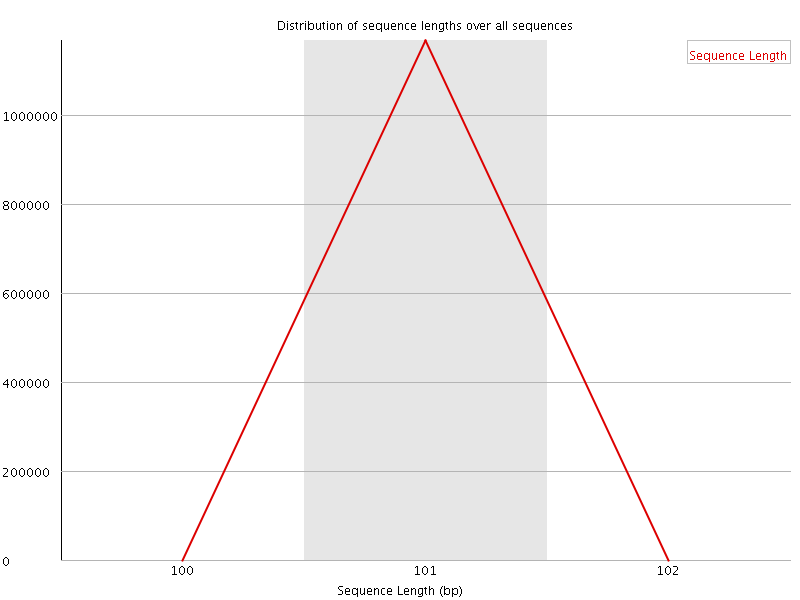
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
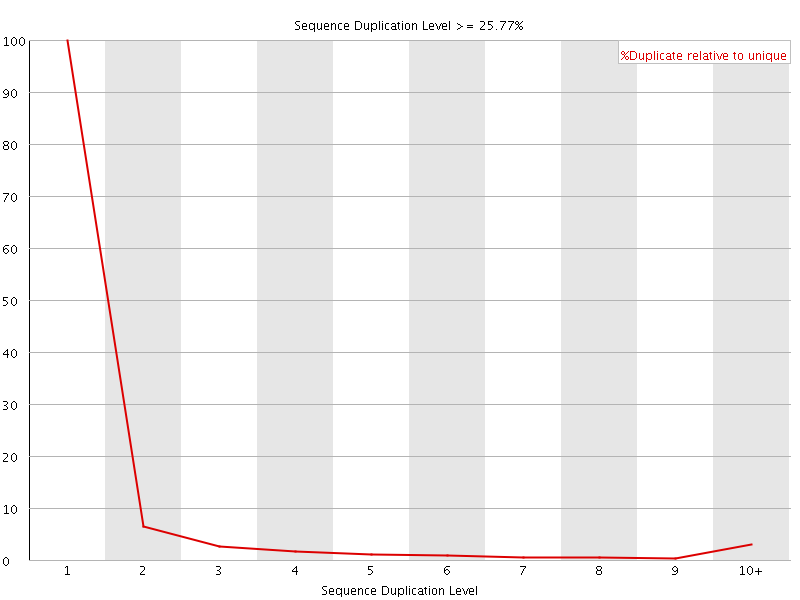
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
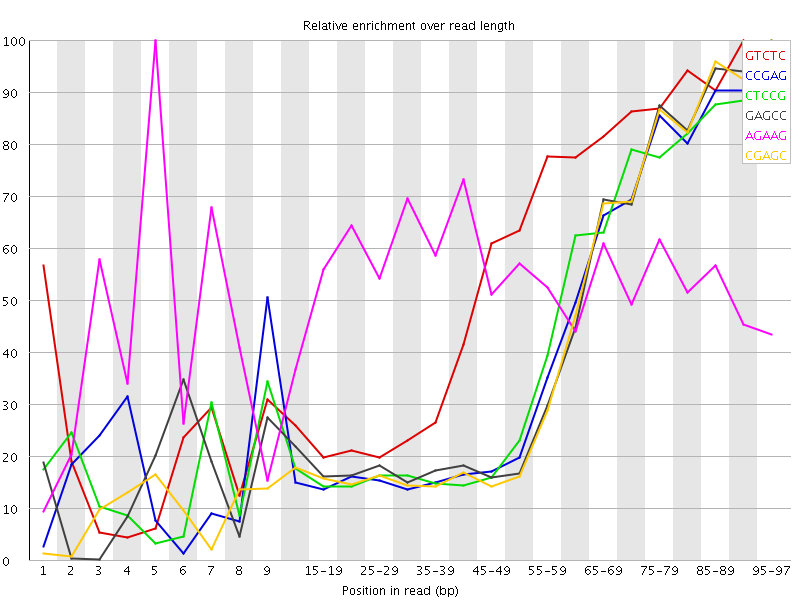
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 241080 | 2.8672879 | 5.0911865 | 90-94 |
| CCGAG | 154920 | 2.853764 | 6.9013877 | 95-97 |
| CTCCG | 161705 | 2.8220053 | 6.6961865 | 95-97 |
| GAGCC | 150605 | 2.774278 | 6.5947256 | 95-97 |
| AGAAG | 316420 | 2.7403495 | 5.0850625 | 5 |
| CGAGC | 143100 | 2.6360292 | 6.475633 | 95-97 |
| GCCCA | 142705 | 2.5056102 | 5.932381 | 90-94 |
| AGCCC | 141410 | 2.482873 | 5.901732 | 90-94 |
| ACACA | 299570 | 2.357045 | 7.805223 | 6 |
| CCCAC | 135980 | 2.2756903 | 5.5937247 | 95-97 |
| ATCTC | 287930 | 2.2380846 | 5.14842 | 95-97 |
| CTTTG | 270305 | 2.1909878 | 8.75611 | 9 |
| CCACG | 121430 | 2.1320643 | 5.748484 | 90-94 |
| TACAC | 271200 | 2.1208954 | 9.998845 | 5 |
| CACAT | 258865 | 2.0244308 | 5.361535 | 7 |
| GCTTT | 226805 | 1.8383937 | 5.4084644 | 1 |
| GGCAG | 93725 | 1.8113494 | 5.8119936 | 90-94 |
| GAGAG | 133145 | 1.7643607 | 5.8403187 | 7 |
| ATACA | 321240 | 1.7225621 | 5.0203924 | 6 |
| CAGGC | 93040 | 1.7138795 | 5.4021783 | 90-94 |
| GCCAA | 137485 | 1.6551806 | 5.026467 | 1 |
| TGGCT | 130525 | 1.6286962 | 5.094113 | 6 |
| TTGAG | 182550 | 1.5618664 | 5.666604 | 9 |
| CACCA | 135555 | 1.5554976 | 5.057406 | 7 |
| GAGTC | 119745 | 1.5032932 | 7.456457 | 9 |
| GACTC | 123725 | 1.4804971 | 5.5406866 | 7 |
| TATAC | 274165 | 1.4612257 | 6.0417104 | 5 |
| GTCTT | 175155 | 1.4197388 | 6.477579 | 1 |
| AAGAC | 169570 | 1.399764 | 5.9915295 | 5 |
| CATGA | 169340 | 1.3893939 | 5.1834674 | 9 |
| CCATG | 115020 | 1.3763329 | 6.051242 | 9 |
| GACTT | 164625 | 1.3425226 | 6.955075 | 7 |
| TGAGT | 156060 | 1.3352225 | 5.650011 | 8 |
| GTGTT | 155845 | 1.3253026 | 6.0701523 | 1 |
| TATGA | 235705 | 1.3179845 | 6.2545958 | 4 |
| TCATA | 238220 | 1.2696486 | 6.487071 | 2 |
| ACTCA | 160460 | 1.2548633 | 5.1909065 | 8 |
| GATTG | 146375 | 1.2523593 | 6.093881 | 7 |
| GTGTA | 143155 | 1.2248098 | 11.189551 | 1 |
| GTTCT | 148460 | 1.2033594 | 5.365228 | 1 |
| GTATG | 138530 | 1.1852388 | 5.666604 | 3 |
| ACATG | 139330 | 1.1431689 | 5.7921166 | 8 |
| ACCAT | 145445 | 1.1374397 | 5.164364 | 8 |
| TGGAC | 90295 | 1.1335745 | 7.626891 | 5 |
| ACACC | 97885 | 1.1232333 | 5.196499 | 6 |
| GTCCA | 93530 | 1.1191828 | 7.4388833 | 1 |
| GGACT | 84850 | 1.0652171 | 7.2434154 | 6 |
| ACACT | 134045 | 1.048287 | 5.255366 | 6 |
| AGACT | 123920 | 1.0167336 | 5.5574083 | 6 |
| TGTAT | 182355 | 1.0134896 | 5.0397944 | 2 |
| GAGTA | 117345 | 1.0101053 | 5.3471303 | 1 |
| CATAC | 115860 | 0.9060729 | 5.3122425 | 3 |
| GTATA | 138980 | 0.7771303 | 6.865543 | 1 |