![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-119_AGGCAGAA-AAGGAGTA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1524499 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
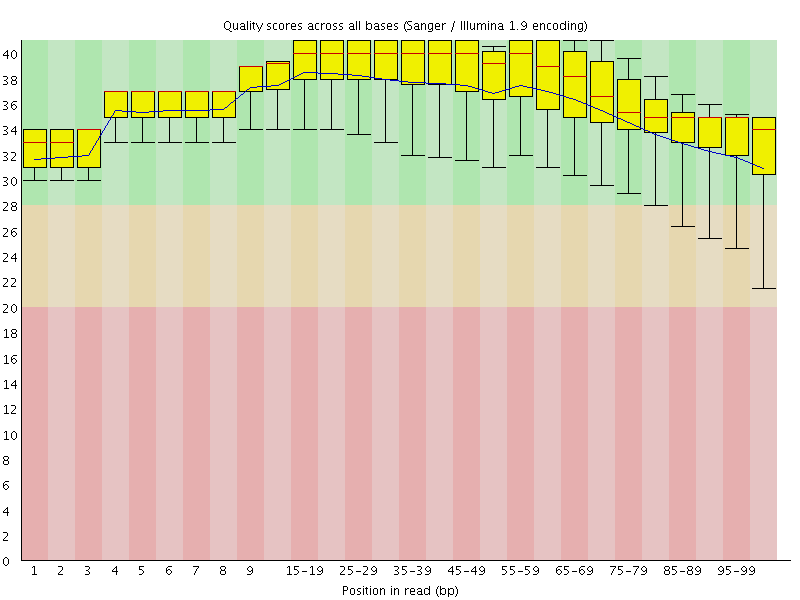
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
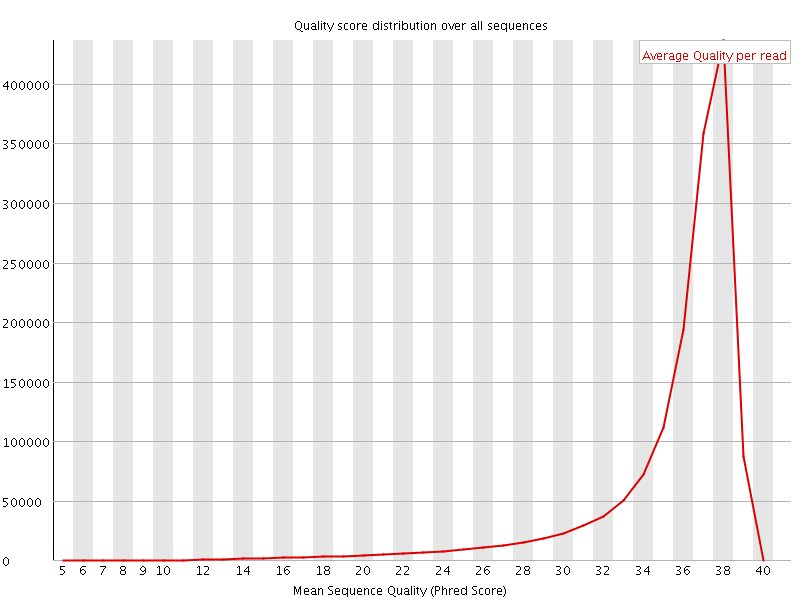
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
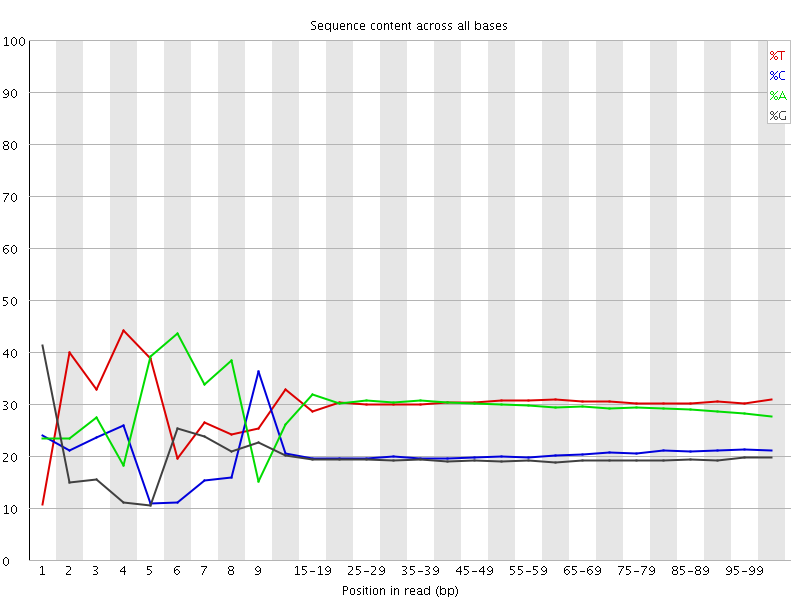
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
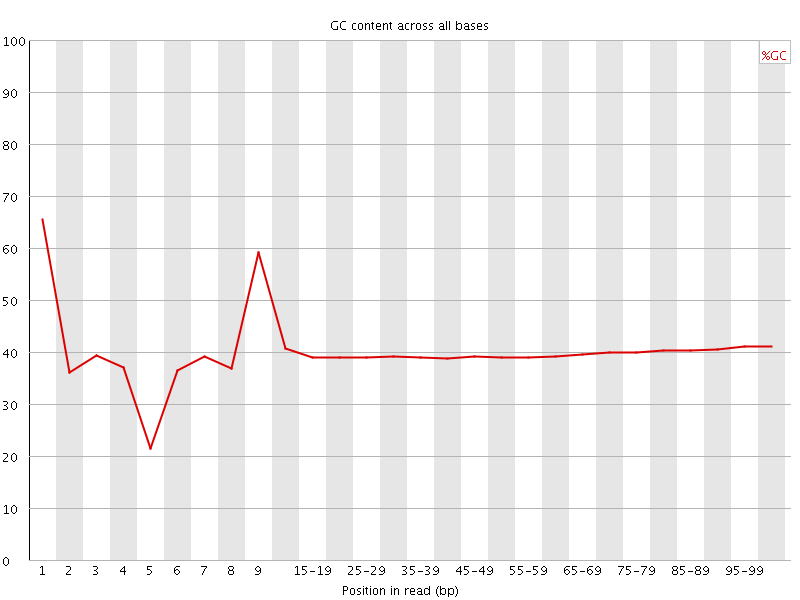
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
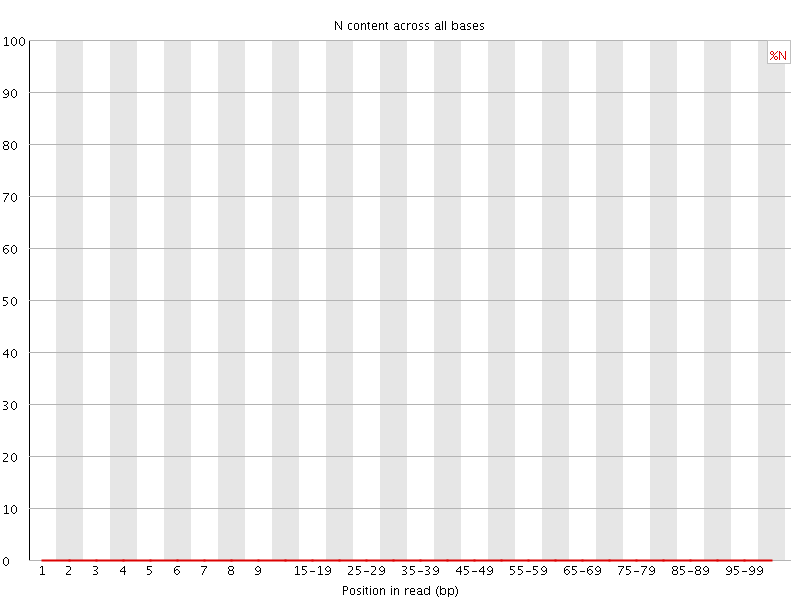
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
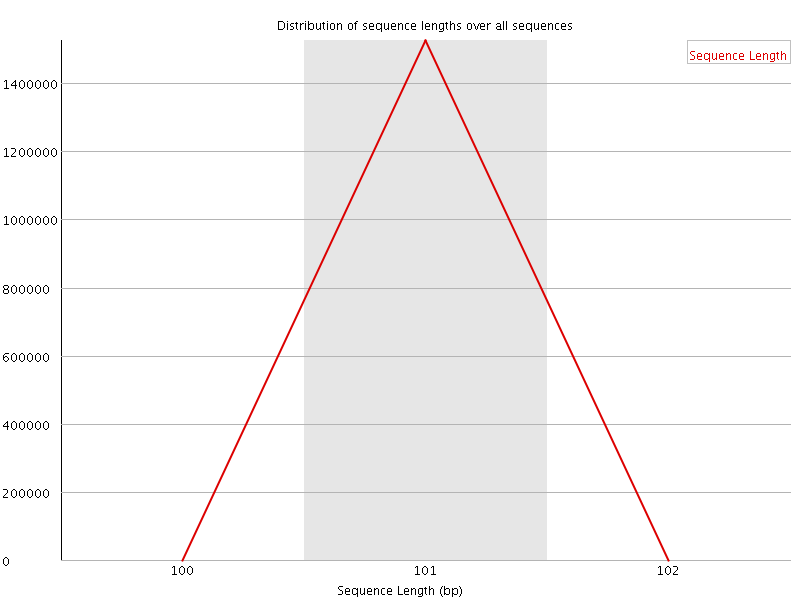
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
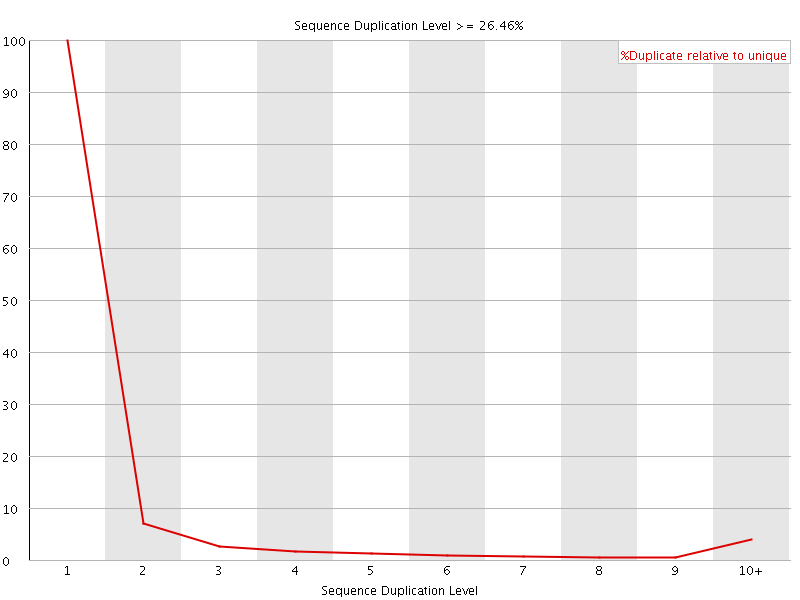
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
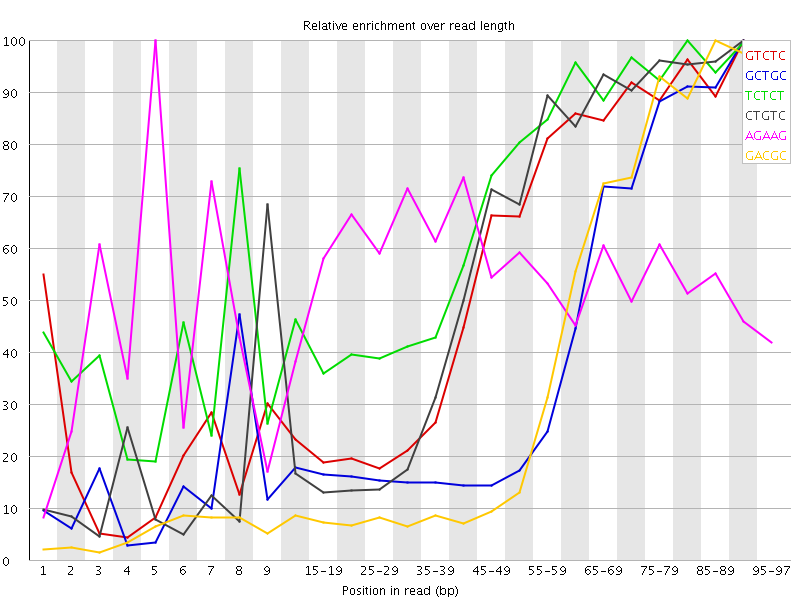
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 353320 | 3.1828544 | 5.5364246 | 90-94 |
| GCTGC | 221880 | 3.1039355 | 7.4373794 | 90-94 |
| TCTCT | 520660 | 3.0203469 | 4.393299 | 80-84 |
| CTGTC | 327065 | 2.9463384 | 5.066318 | 90-94 |
| AGAAG | 410140 | 2.7765124 | 5.0373917 | 5 |
| GACGC | 193475 | 2.7744386 | 7.0953307 | 85-89 |
| CGCTG | 192335 | 2.6906228 | 6.9570227 | 90-94 |
| GCCGA | 183940 | 2.637706 | 7.360981 | 90-94 |
| ACACA | 420170 | 2.6253965 | 7.4995527 | 6 |
| CTGCC | 194575 | 2.6150665 | 6.926297 | 90-94 |
| CGACG | 177915 | 2.5513077 | 7.4836373 | 95-97 |
| CCGAC | 176655 | 2.4337575 | 6.8266997 | 95-97 |
| TGCCG | 172310 | 2.4104884 | 6.9217434 | 90-94 |
| TACAC | 375220 | 2.2871785 | 9.509179 | 5 |
| CACAT | 366540 | 2.2342691 | 5.2558713 | 7 |
| CTGAC | 233985 | 2.160689 | 5.1983075 | 95-97 |
| TGACG | 218675 | 2.101853 | 5.169933 | 95-97 |
| CTTTG | 346660 | 2.0931733 | 8.462323 | 9 |
| GACGA | 207000 | 2.0395257 | 5.4795666 | 95-97 |
| GAGAG | 177145 | 1.8167142 | 6.346105 | 7 |
| GCTTT | 290580 | 1.7545558 | 5.169591 | 1 |
| TGGCT | 169600 | 1.5902789 | 5.020036 | 6 |
| TATAC | 386620 | 1.5796154 | 5.808924 | 5 |
| CACCA | 169985 | 1.5458659 | 5.1423144 | 7 |
| TTGAG | 238995 | 1.539731 | 5.467445 | 9 |
| GACTC | 158780 | 1.4662231 | 5.100666 | 7 |
| GAGTC | 151715 | 1.4582489 | 7.318012 | 9 |
| GTGTA | 226025 | 1.4561715 | 10.716022 | 1 |
| AAGAC | 218095 | 1.4184511 | 6.0478864 | 5 |
| CCATG | 148320 | 1.3696324 | 6.188746 | 9 |
| GTCTT | 213380 | 1.2884132 | 6.4817586 | 1 |
| TGAGT | 198845 | 1.2810638 | 5.27388 | 8 |
| TATGA | 301045 | 1.2802573 | 6.1330094 | 4 |
| GTGTT | 203505 | 1.2790143 | 5.987587 | 1 |
| GACTT | 205360 | 1.2710805 | 6.6125746 | 7 |
| GATTG | 191530 | 1.2339367 | 6.1392813 | 7 |
| TCATA | 300465 | 1.2276112 | 6.2250175 | 2 |
| GTTCT | 193630 | 1.1691604 | 5.2867484 | 1 |
| ACACC | 124215 | 1.1296275 | 5.6758394 | 6 |
| ACCAT | 185100 | 1.1282895 | 5.211565 | 8 |
| TGGAC | 115475 | 1.1099186 | 7.4599514 | 5 |
| ACATG | 174805 | 1.1090901 | 5.6707435 | 8 |
| GTCCA | 115425 | 1.0658698 | 7.6238413 | 1 |
| GGACT | 108805 | 1.0458082 | 7.0989394 | 6 |
| ACACT | 170190 | 1.0374045 | 5.001552 | 6 |
| AGACT | 160470 | 1.0181383 | 5.572136 | 6 |
| GTATG | 153490 | 0.98886305 | 5.6625795 | 3 |
| GAGTA | 149630 | 0.9881673 | 5.29535 | 1 |
| CATAC | 149615 | 0.91198826 | 5.1181283 | 3 |
| GTATA | 180070 | 0.7657857 | 7.2386775 | 1 |