![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-119_AGGCAGAA-AAGGAGTA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1524499 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
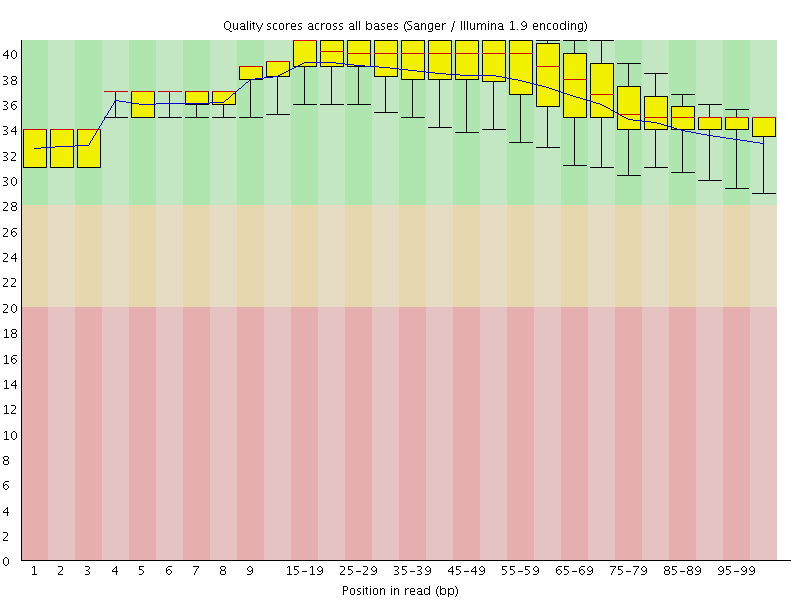
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
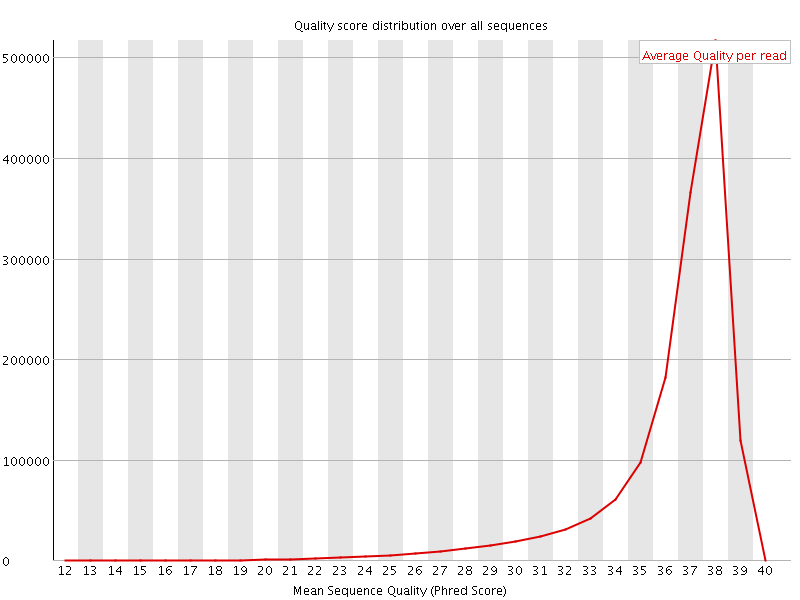
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
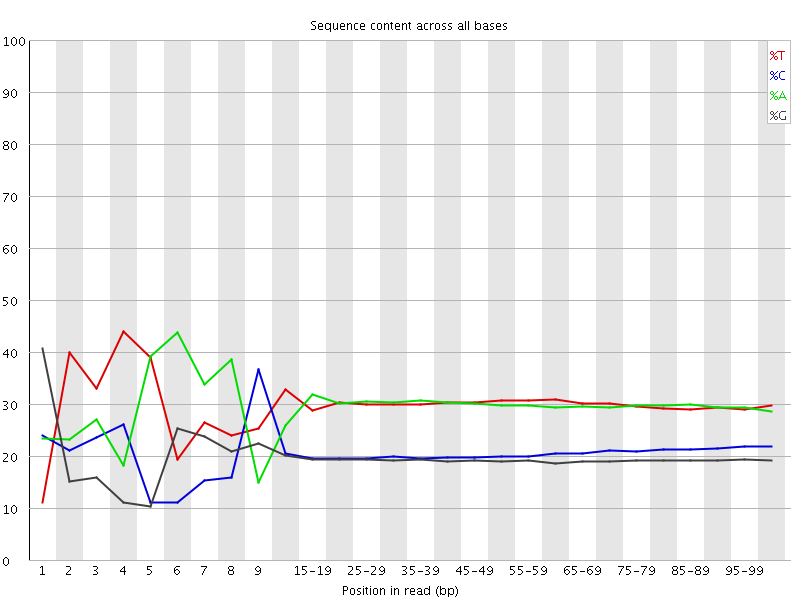
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
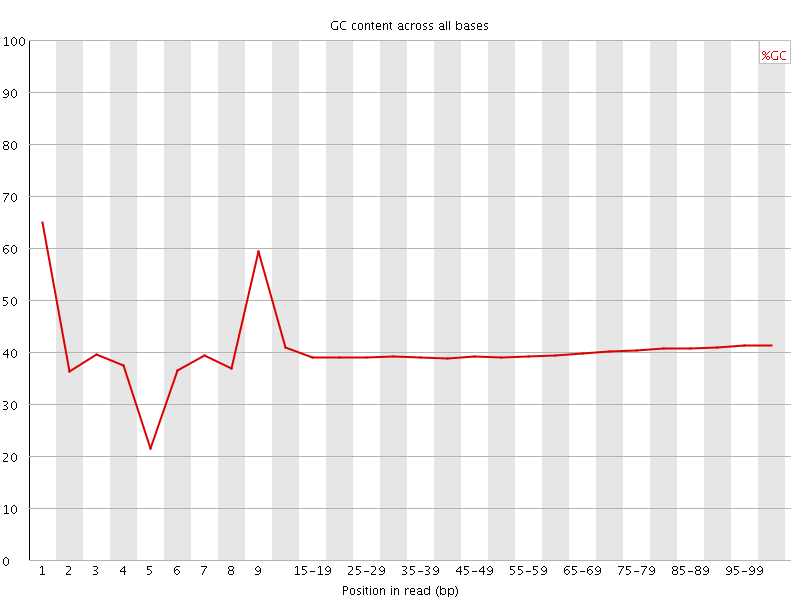
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
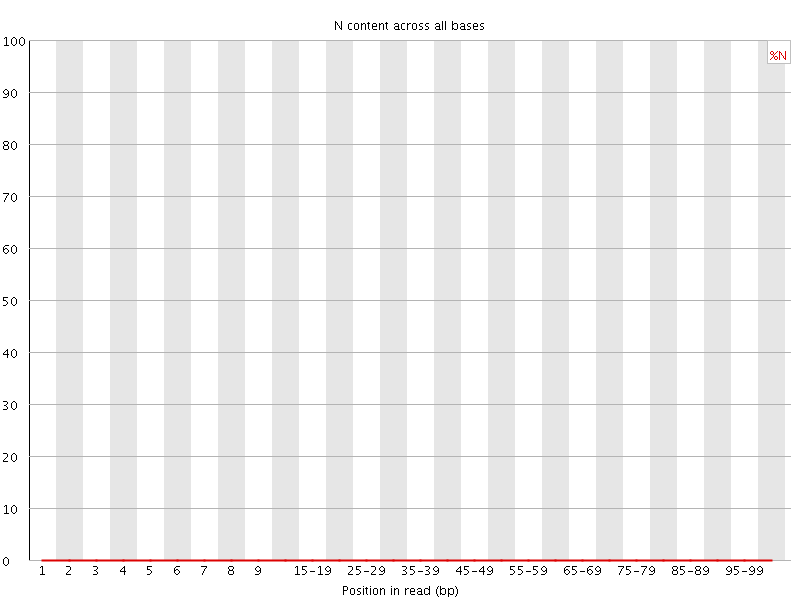
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
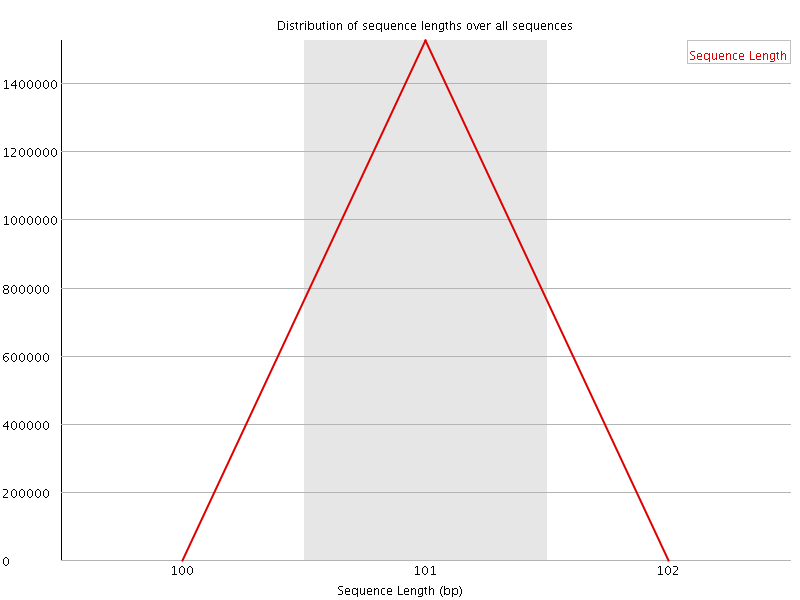
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
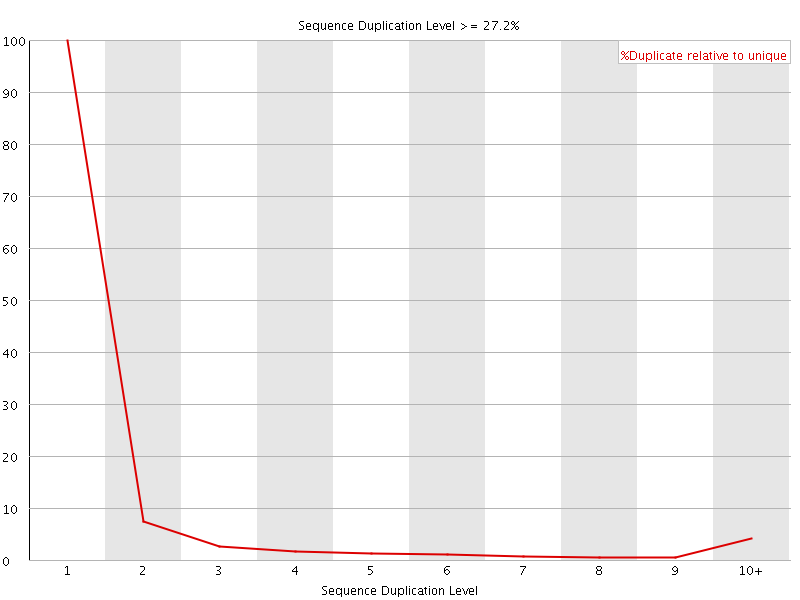
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
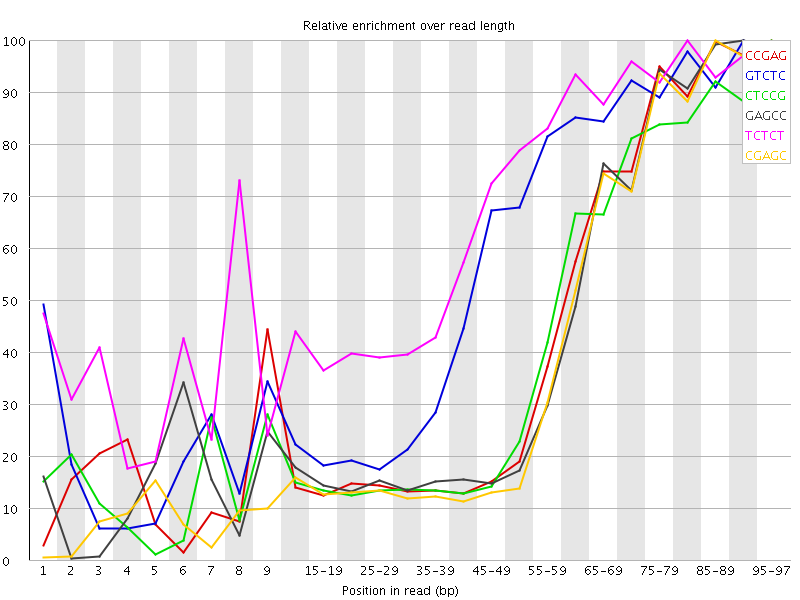
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 227165 | 3.201625 | 7.3644147 | 95-97 |
| GTCTC | 354415 | 3.1993752 | 5.538829 | 90-94 |
| CTCCG | 239240 | 3.155829 | 7.4448743 | 95-97 |
| GAGCC | 223325 | 3.147505 | 7.321799 | 90-94 |
| TCTCT | 522320 | 3.0496924 | 4.4961004 | 80-84 |
| CGAGC | 212105 | 2.9893723 | 7.278048 | 85-89 |
| CTGTC | 328140 | 2.9621854 | 5.0949874 | 90-94 |
| TCTCC | 332025 | 2.8328054 | 5.4896116 | 95-97 |
| AGCCC | 208755 | 2.7807302 | 6.900696 | 90-94 |
| GCCCA | 207685 | 2.7664769 | 7.0427914 | 90-94 |
| ACACA | 420955 | 2.5309398 | 7.701463 | 6 |
| CCCAC | 199575 | 2.5125859 | 6.511086 | 90-94 |
| ATCTC | 423280 | 2.4956825 | 5.766234 | 95-97 |
| CCACG | 180745 | 2.4076214 | 6.624254 | 90-94 |
| TCCGA | 254665 | 2.3214784 | 5.1625338 | 95-97 |
| TACAC | 376965 | 2.244424 | 9.710781 | 5 |
| CACAT | 368520 | 2.1941433 | 5.467367 | 7 |
| GAGAC | 224140 | 2.1830523 | 5.49381 | 95-97 |
| CTTTG | 346345 | 2.139614 | 8.976512 | 9 |
| CGAGA | 217145 | 2.1149232 | 5.4969587 | 95-97 |
| GGCAG | 140370 | 2.093199 | 7.145262 | 90-94 |
| CAGGC | 140240 | 1.976519 | 6.720419 | 90-94 |
| ACGAG | 202505 | 1.9723344 | 5.3049116 | 95-97 |
| AGAGA | 277355 | 1.8667943 | 5.133145 | 8 |
| GAGAG | 176155 | 1.8152939 | 6.160406 | 7 |
| GCTTT | 292945 | 1.8097252 | 5.370966 | 1 |
| TGGCT | 169985 | 1.6235695 | 5.2003717 | 6 |
| TATAC | 390110 | 1.589514 | 5.938279 | 5 |
| TTGAG | 235570 | 1.5548767 | 5.6962667 | 9 |
| CACCA | 169945 | 1.4785597 | 5.057594 | 7 |
| GAGTC | 152030 | 1.4663303 | 7.234144 | 9 |
| GACTC | 159675 | 1.4555672 | 5.1179757 | 7 |
| GTCTT | 227810 | 1.4073409 | 6.3445096 | 1 |
| AAGAC | 216195 | 1.3753048 | 5.6935 | 5 |
| CATGA | 216225 | 1.362125 | 5.173909 | 9 |
| CCATG | 145855 | 1.3295867 | 5.798605 | 9 |
| GTGTT | 201595 | 1.3176906 | 6.1201434 | 1 |
| TGAGT | 199575 | 1.317292 | 5.2642455 | 8 |
| TATGA | 301340 | 1.299096 | 6.506657 | 4 |
| GACTT | 202495 | 1.263232 | 6.5663333 | 7 |
| GATTG | 189980 | 1.2539604 | 6.5667067 | 7 |
| GTATG | 188615 | 1.2449507 | 5.920276 | 3 |
| TCATA | 303740 | 1.2375972 | 6.31239 | 2 |
| GTTCT | 194215 | 1.1998012 | 5.4069123 | 1 |
| GTGTA | 176090 | 1.1622797 | 11.28506 | 1 |
| TGGAC | 115895 | 1.117808 | 7.776588 | 5 |
| ACATG | 176460 | 1.1116226 | 5.7236753 | 8 |
| GTCCA | 118630 | 1.0814089 | 7.8193173 | 1 |
| ACACC | 123145 | 1.0713892 | 5.538466 | 6 |
| GGACT | 108505 | 1.0465314 | 7.2622013 | 6 |
| ACACT | 173110 | 1.0306852 | 5.2046785 | 6 |
| AGACT | 158865 | 1.0007815 | 5.427412 | 6 |
| GAGTA | 149165 | 0.99422604 | 5.3327074 | 1 |
| TGTAT | 231015 | 0.98623955 | 5.082055 | 2 |
| GCCGT | 70005 | 0.977048 | 5.046105 | 95-97 |
| CATAC | 151560 | 0.9023781 | 5.262413 | 3 |
| TGCCG | 63820 | 0.89072496 | 5.0618954 | 95-97 |
| GTATA | 178520 | 0.7696111 | 7.096919 | 1 |