![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-118_AGGCAGAA-ACTGCATA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2733160 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
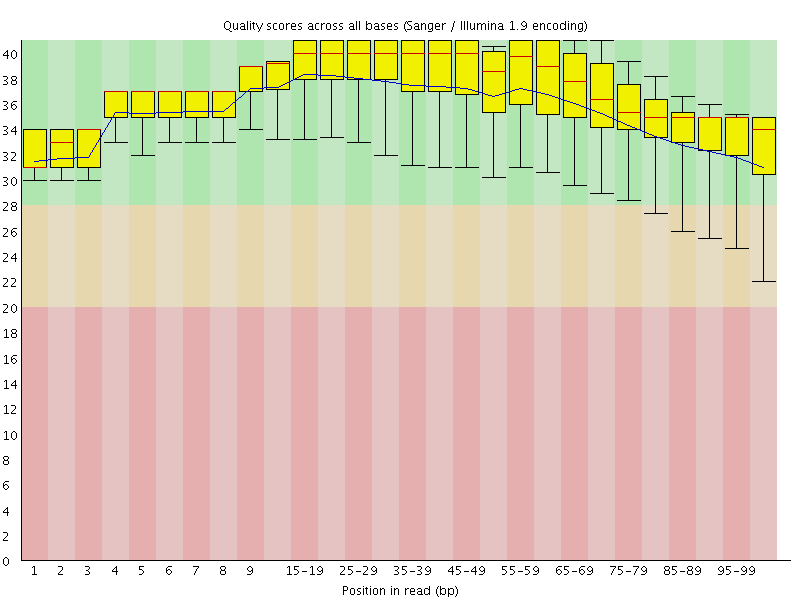
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
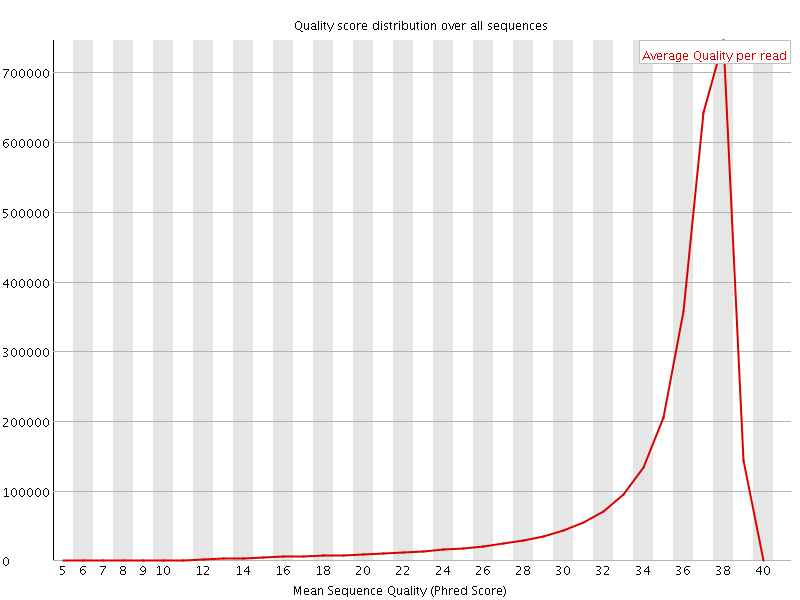
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
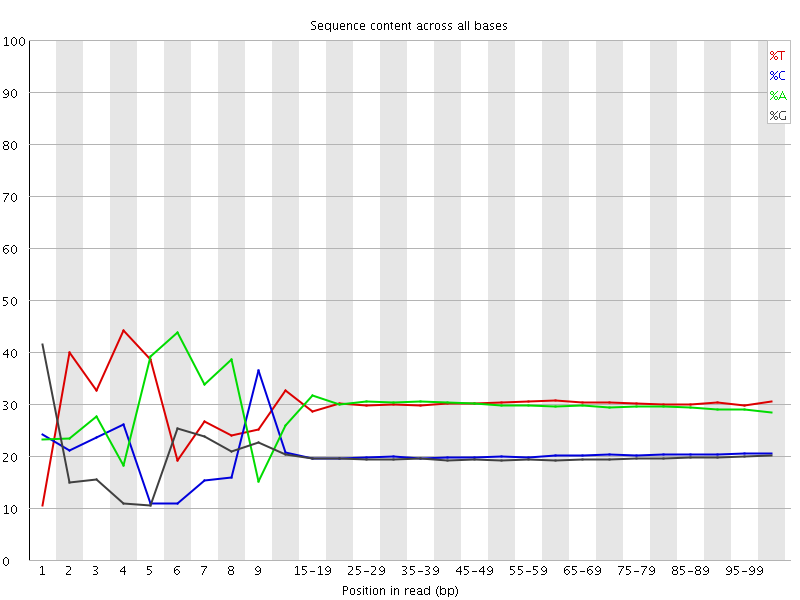
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
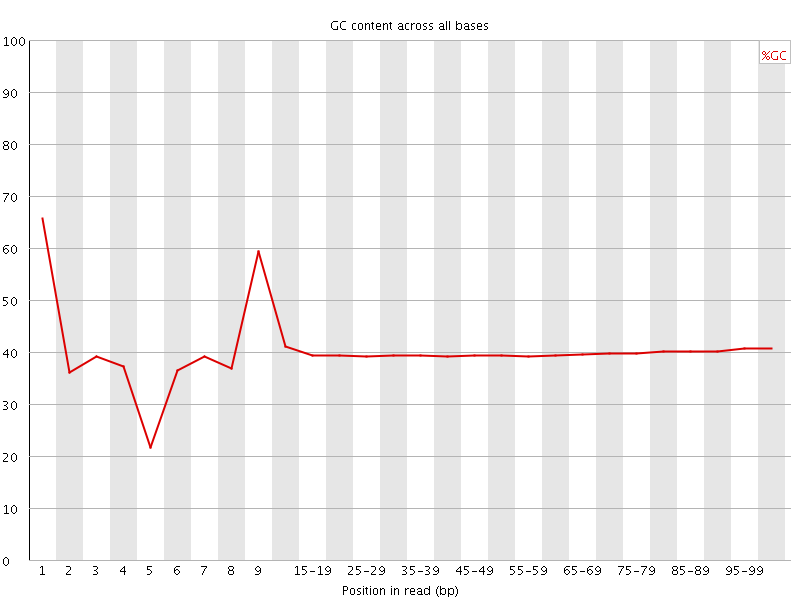
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
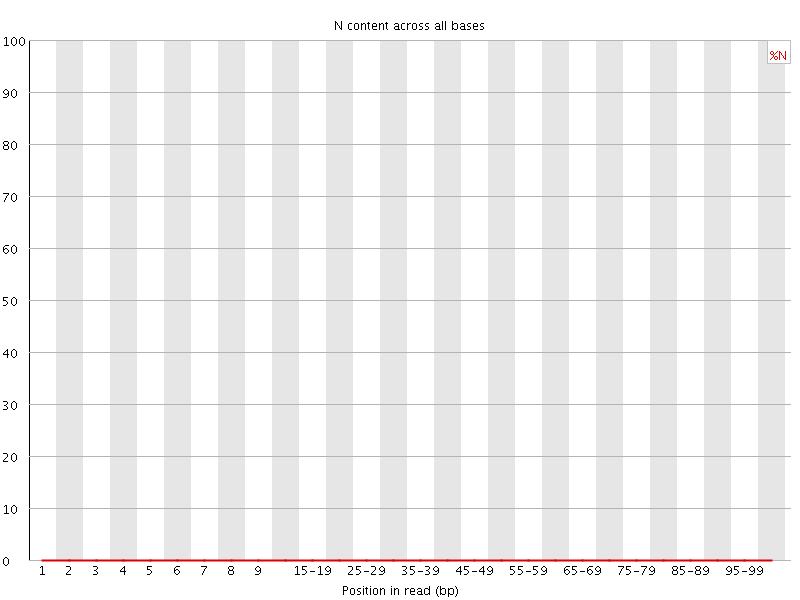
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
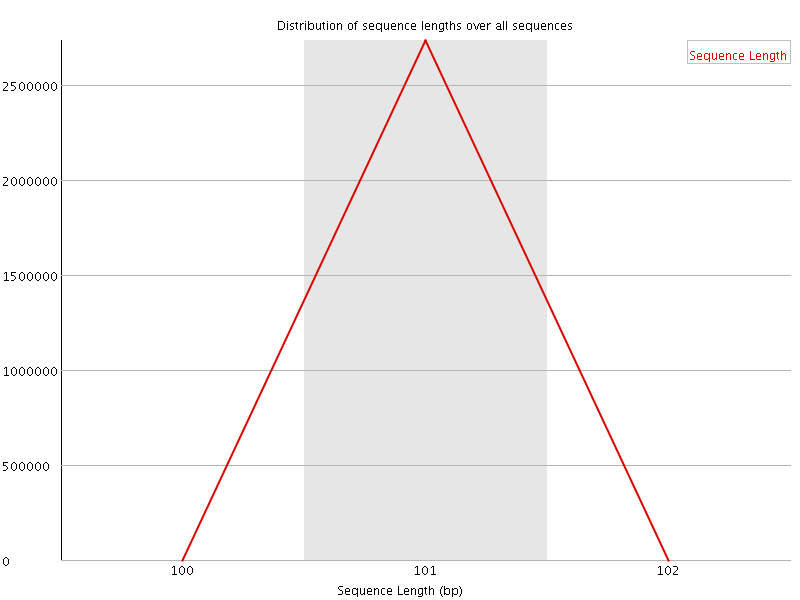
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
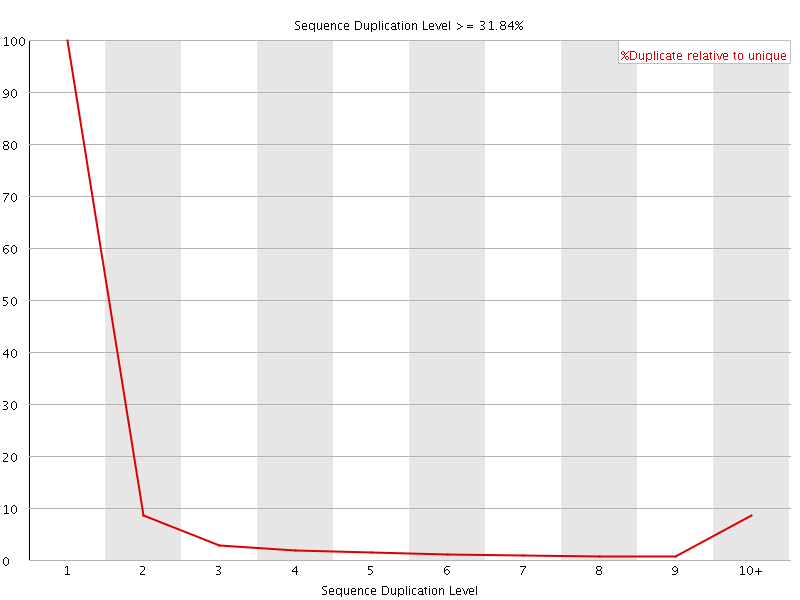
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
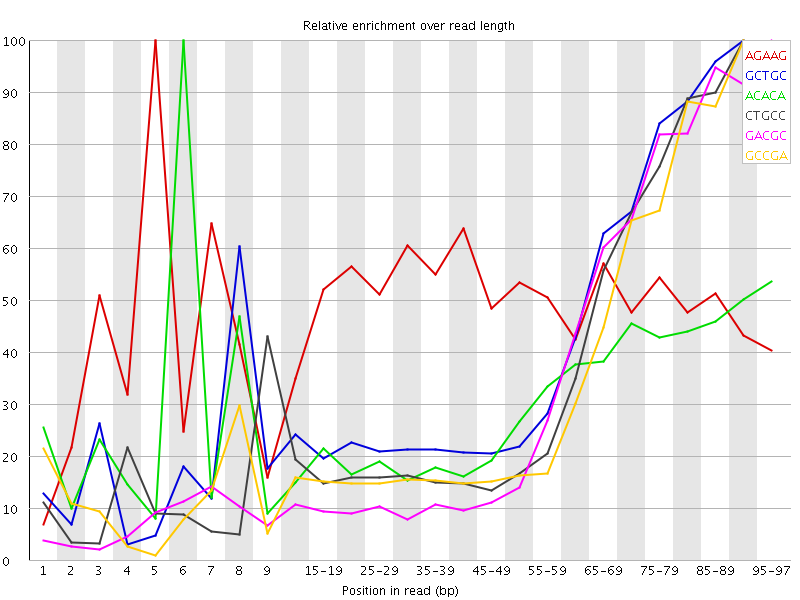
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 705205 | 2.5778475 | 5.1637616 | 5 |
| GCTGC | 317480 | 2.472849 | 5.609103 | 90-94 |
| ACACA | 629015 | 2.2006655 | 7.246264 | 6 |
| CTGCC | 271025 | 2.0652196 | 5.290079 | 90-94 |
| GACGC | 258355 | 2.0437784 | 5.5496707 | 95-97 |
| GCCGA | 257940 | 2.0404956 | 5.511846 | 90-94 |
| CTTTG | 596885 | 2.0375156 | 7.757226 | 9 |
| CGACG | 255140 | 2.0183456 | 5.5944366 | 95-97 |
| CGCTG | 258655 | 2.0146616 | 5.3597364 | 95-97 |
| CCGAC | 245705 | 1.901545 | 5.222827 | 95-97 |
| TACAC | 550165 | 1.8951793 | 9.194136 | 5 |
| TGCCG | 233880 | 1.8216896 | 5.211692 | 90-94 |
| GAGAG | 323695 | 1.7792614 | 5.883091 | 7 |
| GCTTT | 504630 | 1.7225955 | 5.082101 | 1 |
| ATACA | 677760 | 1.5870686 | 5.0589337 | 6 |
| TTGAG | 413215 | 1.4643553 | 5.06631 | 9 |
| GACTC | 274145 | 1.4200369 | 5.08679 | 7 |
| GAGTC | 262350 | 1.389072 | 6.971198 | 9 |
| CCATG | 263335 | 1.3640423 | 5.659452 | 9 |
| AAGAC | 375145 | 1.3415802 | 5.5947433 | 5 |
| TATAC | 574730 | 1.3250978 | 5.9311466 | 5 |
| GTCTT | 373815 | 1.2760481 | 6.449914 | 1 |
| GTGTA | 356940 | 1.2649275 | 10.149693 | 1 |
| GACTT | 356595 | 1.2362926 | 6.014014 | 7 |
| GTGTT | 350085 | 1.2215413 | 5.795682 | 1 |
| TATGA | 518200 | 1.2212535 | 5.713966 | 4 |
| TCATA | 524265 | 1.2087456 | 5.9064436 | 2 |
| GATTG | 338805 | 1.2006605 | 5.5097685 | 7 |
| GTTCT | 345095 | 1.17801 | 5.3934197 | 1 |
| ACACC | 216730 | 1.1154488 | 5.121527 | 6 |
| TGGAC | 206655 | 1.094182 | 7.0325055 | 5 |
| ACATG | 310710 | 1.0940495 | 5.254283 | 8 |
| GTCCA | 207085 | 1.0726745 | 7.6891246 | 1 |
| ACACT | 297650 | 1.025329 | 5.218671 | 6 |
| GGACT | 191305 | 1.012908 | 6.5320916 | 6 |
| AGACT | 282285 | 0.9939614 | 5.2831573 | 6 |
| GAGTA | 265160 | 0.9543642 | 5.1855965 | 1 |
| GTATG | 266630 | 0.944886 | 5.060936 | 3 |
| GTATA | 315515 | 0.74358124 | 7.403744 | 1 |