![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-118_AGGCAGAA-ACTGCATA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2733160 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
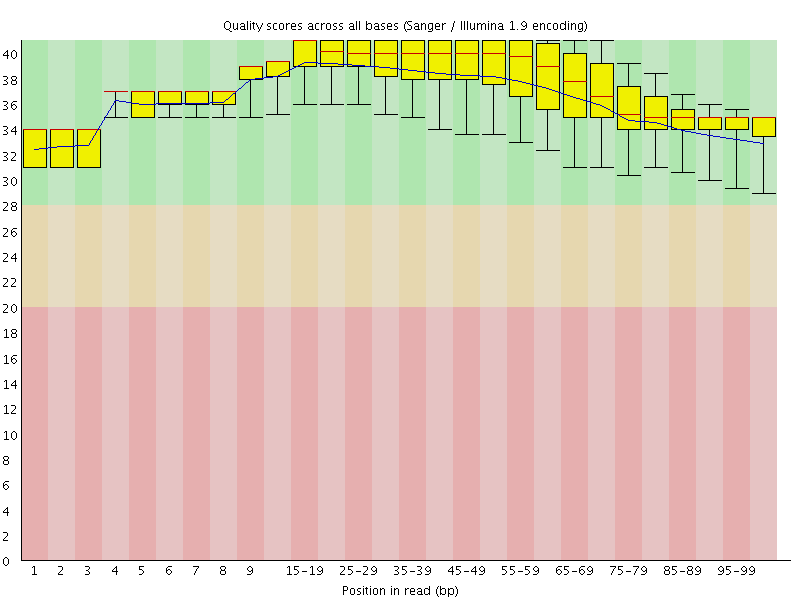
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
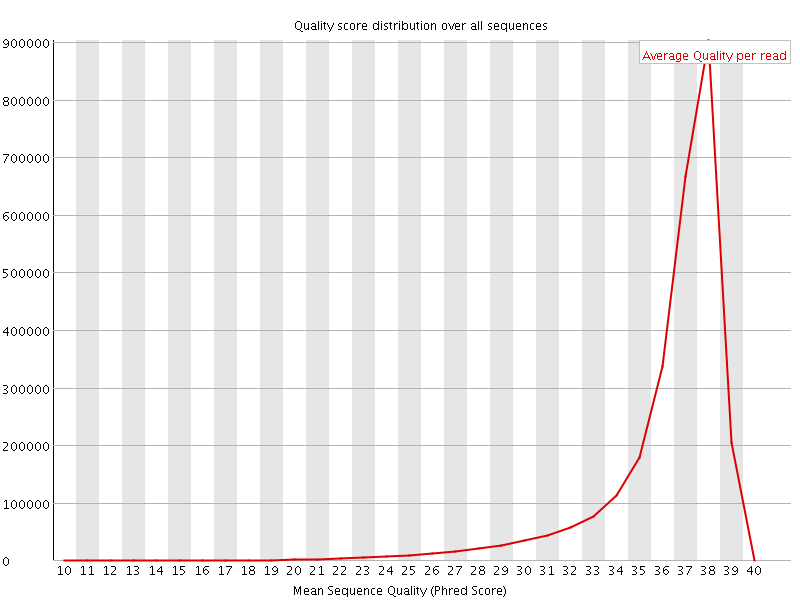
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
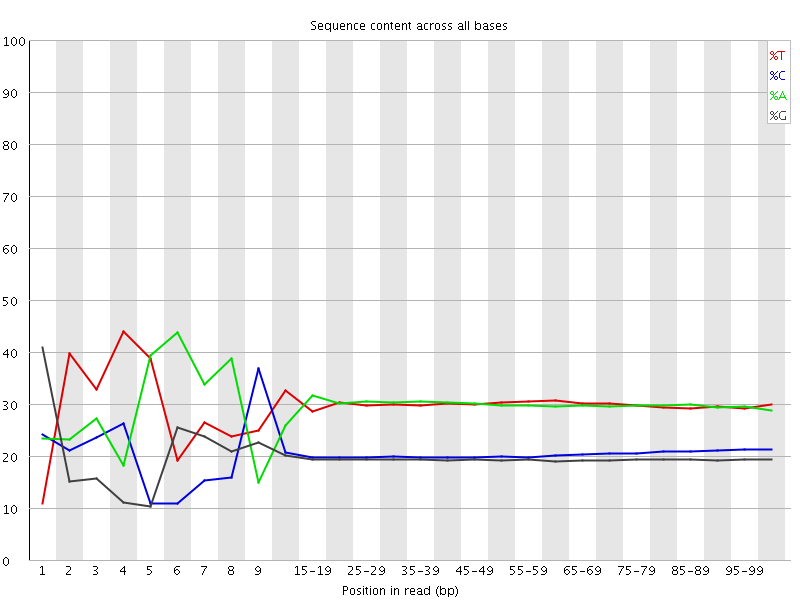
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
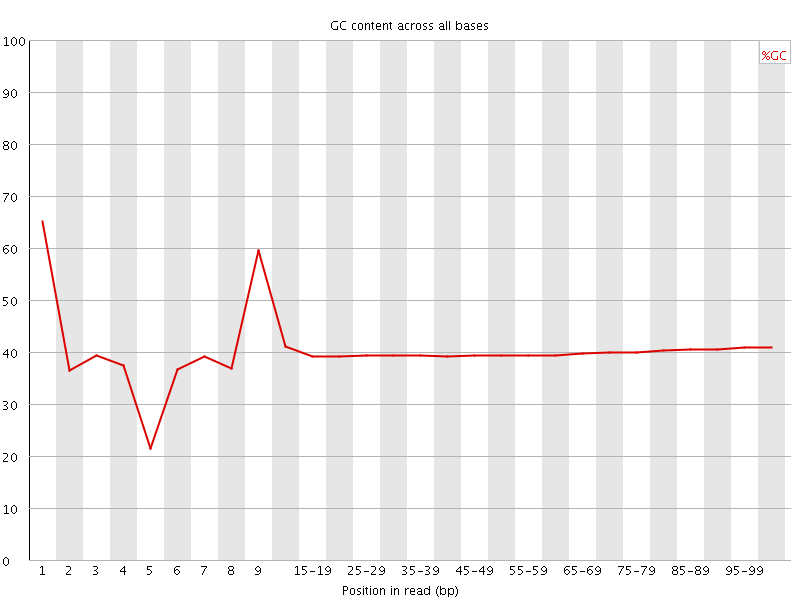
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
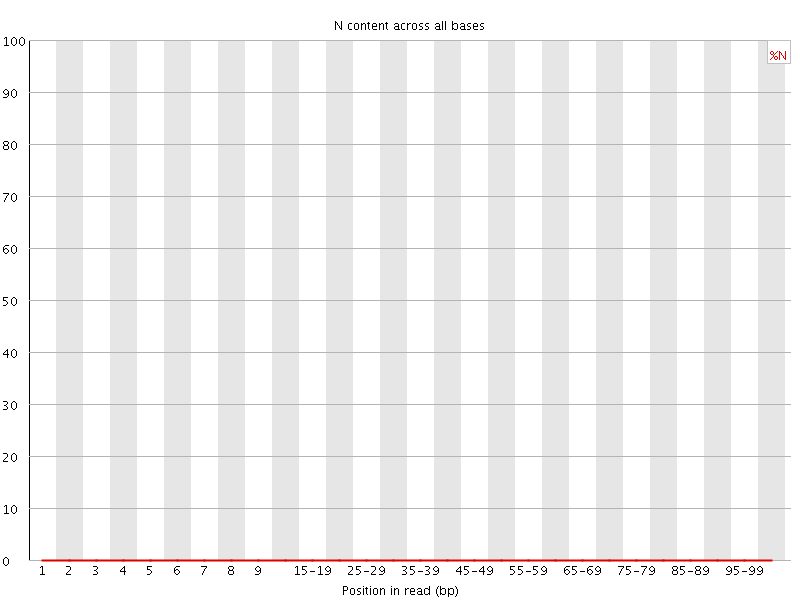
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
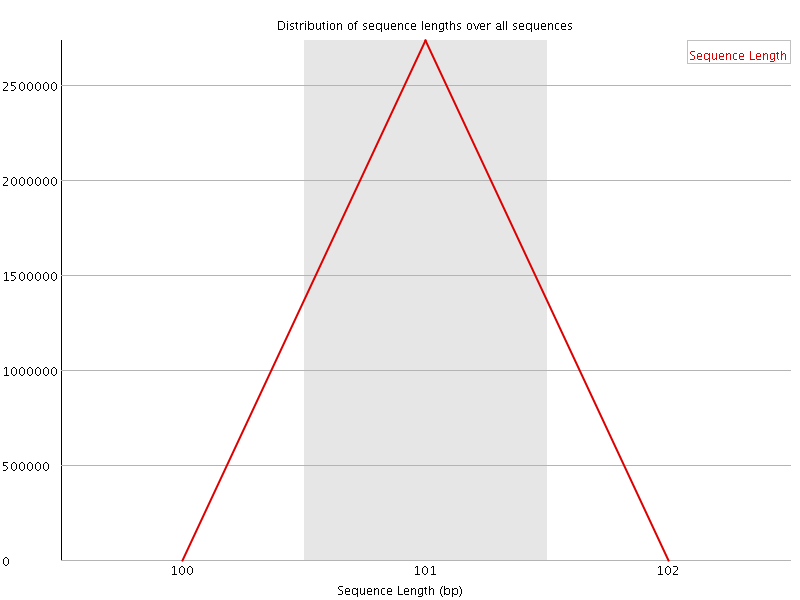
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
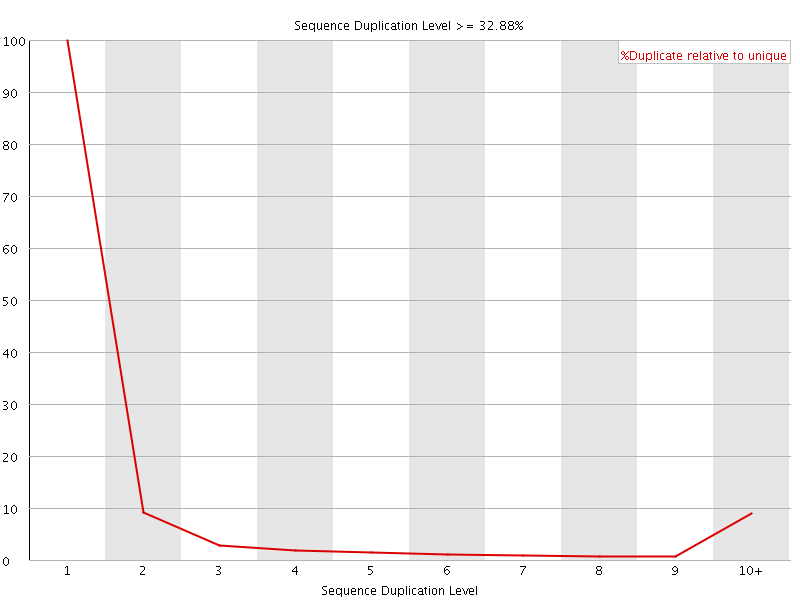
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
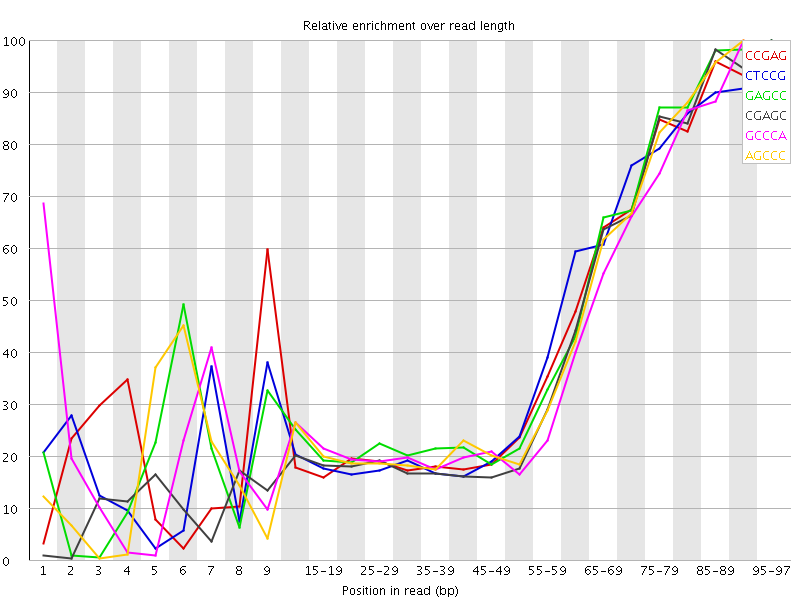
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 330845 | 2.5903914 | 6.00332 | 95-97 |
| CTCCG | 340490 | 2.542904 | 5.8542776 | 95-97 |
| GAGCC | 323785 | 2.535114 | 5.694549 | 95-97 |
| CGAGC | 298775 | 2.3392954 | 5.6477275 | 95-97 |
| GCCCA | 302810 | 2.2752752 | 5.4424734 | 90-94 |
| AGCCC | 300920 | 2.261074 | 5.2464595 | 90-94 |
| ACACA | 637140 | 2.1567087 | 7.2950296 | 6 |
| CTTTG | 597935 | 2.0709672 | 8.428205 | 9 |
| CCCAC | 284560 | 2.0519202 | 5.1313834 | 90-94 |
| CCACG | 249830 | 1.8771905 | 5.129873 | 90-94 |
| TACAC | 556565 | 1.8725542 | 9.237698 | 5 |
| AGAGA | 504320 | 1.8536044 | 5.019915 | 8 |
| GAGAG | 320395 | 1.7909713 | 5.905529 | 7 |
| GCTTT | 507995 | 1.7594568 | 5.1528354 | 1 |
| TTGAG | 415165 | 1.5074925 | 5.0138702 | 9 |
| GAGTC | 263285 | 1.4038275 | 6.889434 | 9 |
| CCATG | 266925 | 1.36584 | 5.6564517 | 9 |
| GTCTT | 390445 | 1.3523186 | 6.4259286 | 1 |
| TATAC | 580150 | 1.3292466 | 6.169782 | 5 |
| AAGAC | 376450 | 1.3278257 | 5.7135873 | 5 |
| GTGTT | 351820 | 1.2697457 | 5.9136577 | 1 |
| TGAGT | 348005 | 1.2636302 | 5.0279546 | 8 |
| TATGA | 520115 | 1.2417717 | 5.853748 | 4 |
| GACTT | 354535 | 1.2354251 | 6.158197 | 7 |
| GATTG | 338285 | 1.228336 | 5.7145443 | 7 |
| TCATA | 524845 | 1.2025311 | 5.873057 | 2 |
| GTTCT | 345605 | 1.1970139 | 5.520655 | 1 |
| GTATG | 306090 | 1.1114339 | 5.2832255 | 3 |
| GTGTA | 304730 | 1.1064955 | 10.309394 | 1 |
| ACATG | 312135 | 1.0943041 | 5.135053 | 8 |
| TGGAC | 203860 | 1.0869752 | 7.160875 | 5 |
| ACACC | 219720 | 1.0855285 | 5.1691813 | 6 |
| GTCCA | 209815 | 1.0736113 | 7.5754614 | 1 |
| GGACT | 190390 | 1.0151536 | 6.757591 | 6 |
| ACACT | 297800 | 1.0019432 | 5.146553 | 6 |
| AGACT | 279340 | 0.97932917 | 5.2540383 | 6 |
| GAGTA | 266120 | 0.97218776 | 5.19762 | 1 |
| GTATA | 317035 | 0.7569194 | 7.1201525 | 1 |
| ATACT | 326860 | 0.74890554 | 5.052245 | 6 |