![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-117_AGGCAGAA-GTAAGGAG_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1555051 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
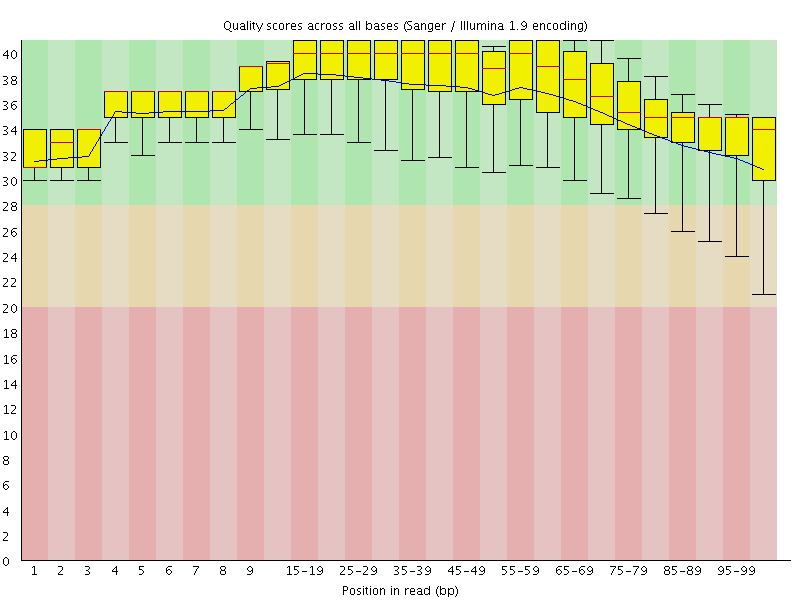
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
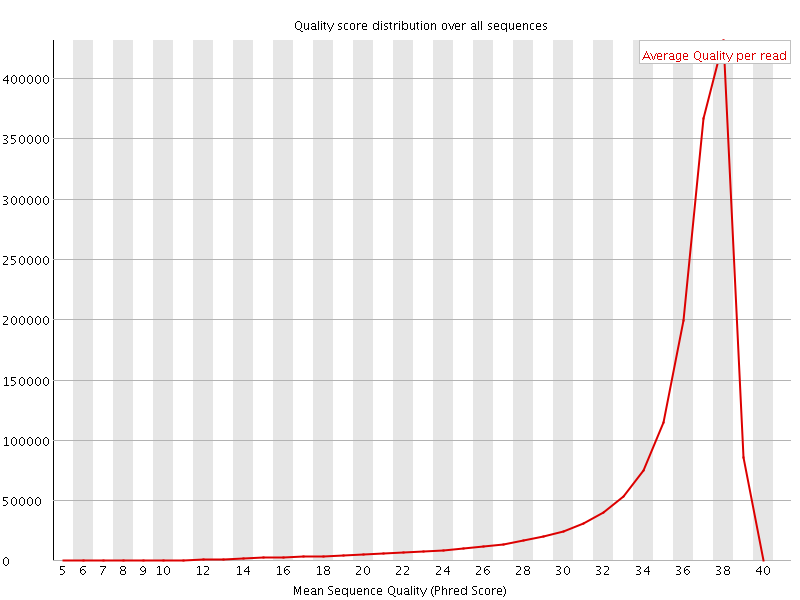
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
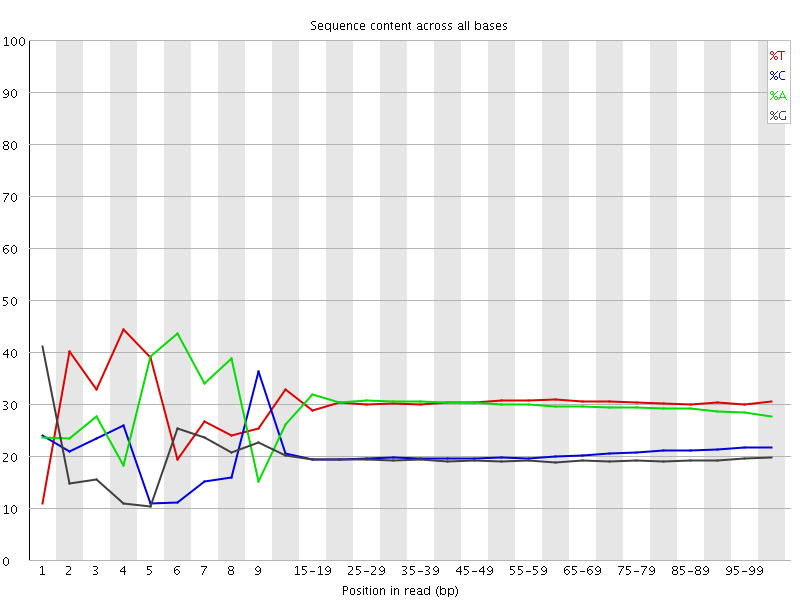
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
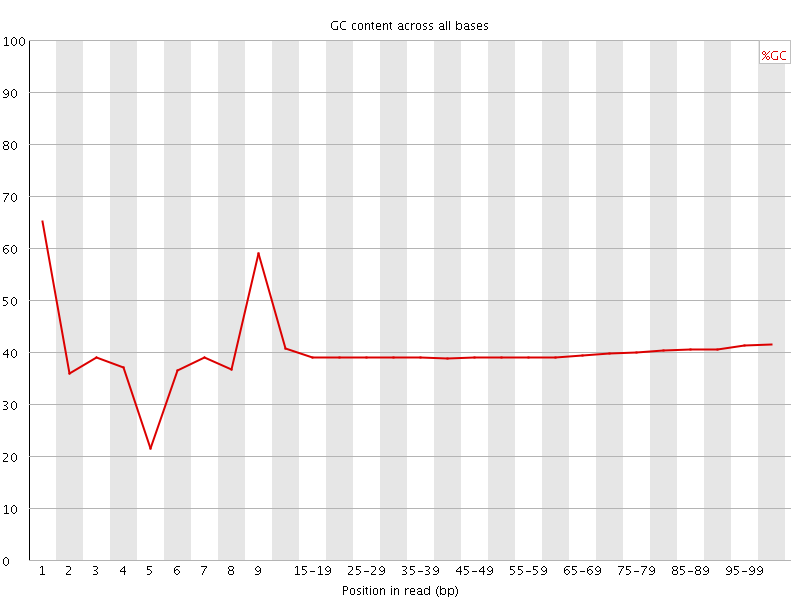
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
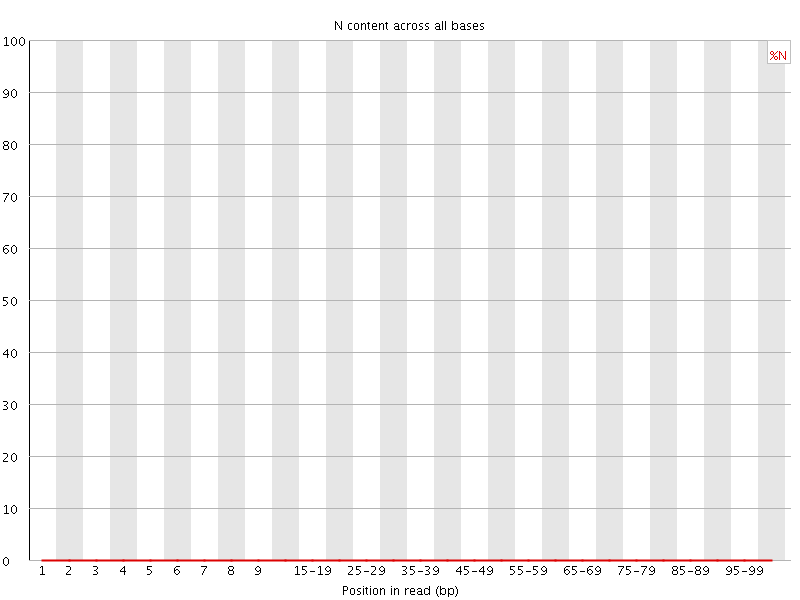
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
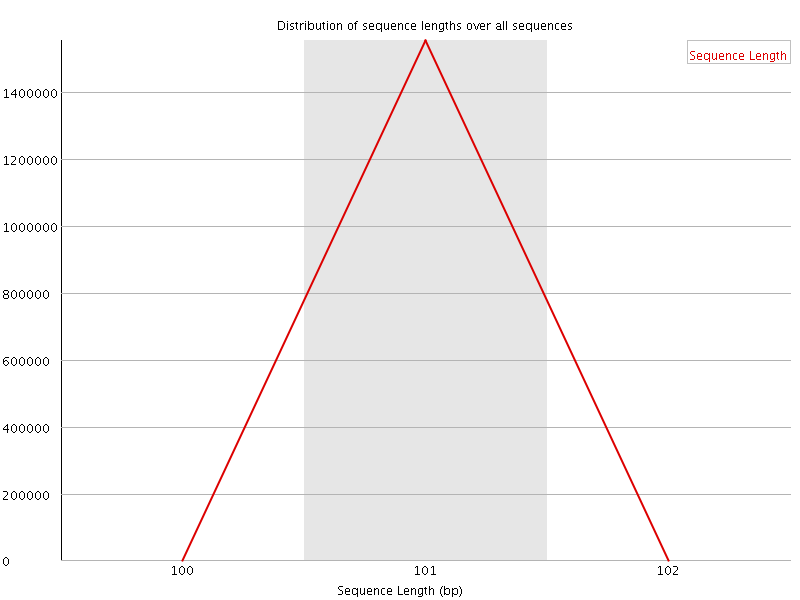
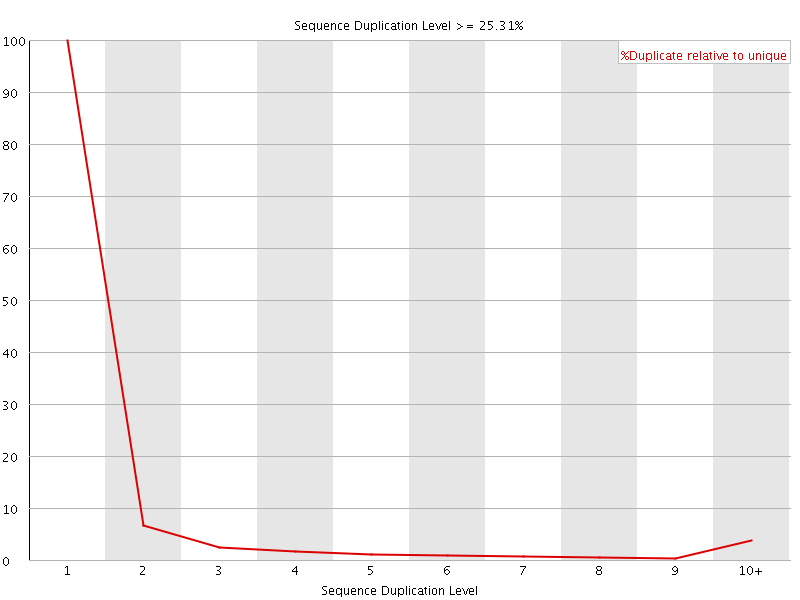
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
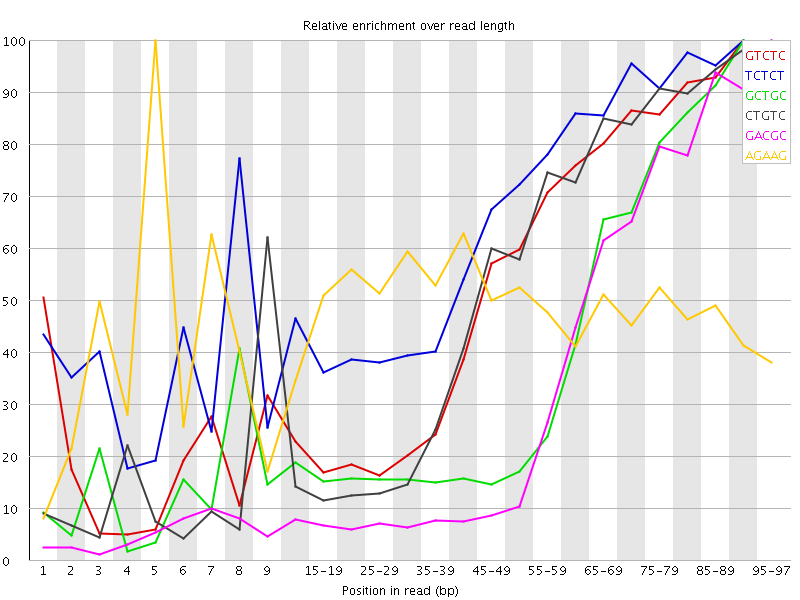
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 358610 | 3.1784334 | 5.862412 | 90-94 |
| TCTCT | 536060 | 3.047688 | 4.5931954 | 90-94 |
| GCTGC | 216705 | 2.994286 | 7.4223633 | 90-94 |
| CTGTC | 332850 | 2.950117 | 5.5713983 | 95-97 |
| GACGC | 189735 | 2.68159 | 7.6639943 | 95-97 |
| AGAAG | 400345 | 2.657992 | 5.48445 | 5 |
| CGCTG | 186355 | 2.5749297 | 7.0592504 | 95-97 |
| ACACA | 417245 | 2.5386925 | 6.9634647 | 6 |
| CTGCC | 188740 | 2.4965377 | 6.9502096 | 90-94 |
| GCCGA | 174575 | 2.4673288 | 7.204704 | 95-97 |
| CGACG | 170445 | 2.408958 | 7.380651 | 95-97 |
| TGCCG | 165210 | 2.282762 | 6.8969812 | 90-94 |
| CCGAC | 168170 | 2.2753243 | 6.9561524 | 95-97 |
| TACAC | 367295 | 2.184808 | 8.601208 | 5 |
| CTGAC | 231350 | 2.097398 | 5.3309164 | 95-97 |
| TGACG | 217145 | 2.0564175 | 5.466091 | 95-97 |
| CTTTG | 333740 | 1.9820542 | 7.698453 | 9 |
| AGAGA | 294290 | 1.9538661 | 5.2321606 | 8 |
| GACGA | 200795 | 1.945069 | 5.475209 | 95-97 |
| GAGAG | 184465 | 1.8665783 | 6.4971104 | 7 |
| TGGAG | 172495 | 1.7064288 | 5.076875 | 5 |
| TATAC | 387100 | 1.542901 | 5.774964 | 5 |
| GAGTC | 144610 | 1.369493 | 6.718975 | 9 |
| CCATG | 150670 | 1.3659605 | 5.6539183 | 9 |
| GTGTA | 215065 | 1.3647333 | 10.093124 | 1 |
| AAGAC | 214635 | 1.3641735 | 5.703481 | 5 |
| GTCTT | 212470 | 1.2618418 | 6.3089566 | 1 |
| TATGA | 293690 | 1.2227966 | 5.5276303 | 4 |
| GATTG | 190215 | 1.207043 | 5.5515537 | 7 |
| GACTT | 196790 | 1.1954485 | 5.729906 | 7 |
| GTGTT | 191140 | 1.1857933 | 5.7447286 | 1 |
| TCATA | 296110 | 1.1802334 | 5.5487056 | 2 |
| GTTCT | 196975 | 1.1698183 | 5.1739206 | 1 |
| ACACC | 124515 | 1.1053593 | 5.1402063 | 6 |
| TGGAC | 114660 | 1.0858591 | 6.803239 | 5 |
| GTCCA | 117725 | 1.067284 | 7.6562524 | 1 |
| AGACT | 159645 | 0.99198204 | 5.2853346 | 6 |
| GGACT | 104360 | 0.9883154 | 6.2458973 | 6 |
| GAGTA | 146330 | 0.9497996 | 5.0565033 | 1 |
| GTATA | 180545 | 0.7517102 | 7.5715814 | 1 |