![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-117_AGGCAGAA-GTAAGGAG_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1555051 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
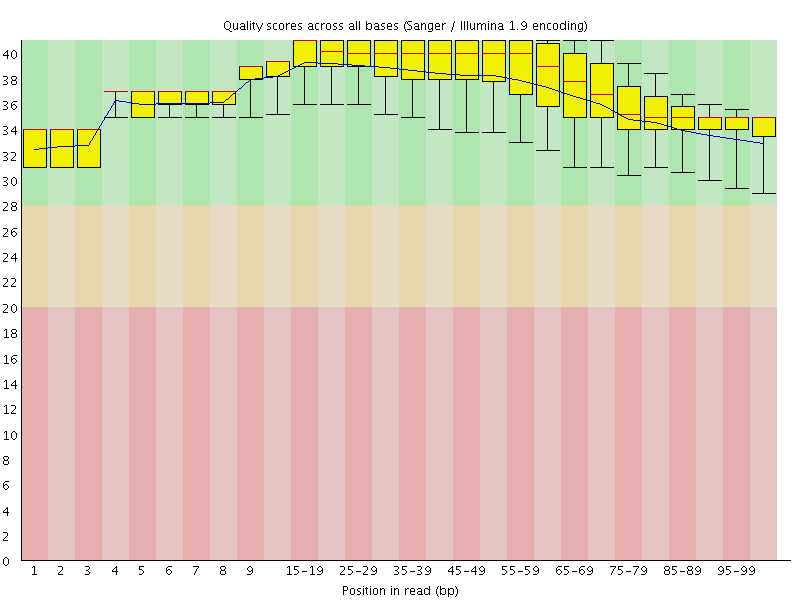
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
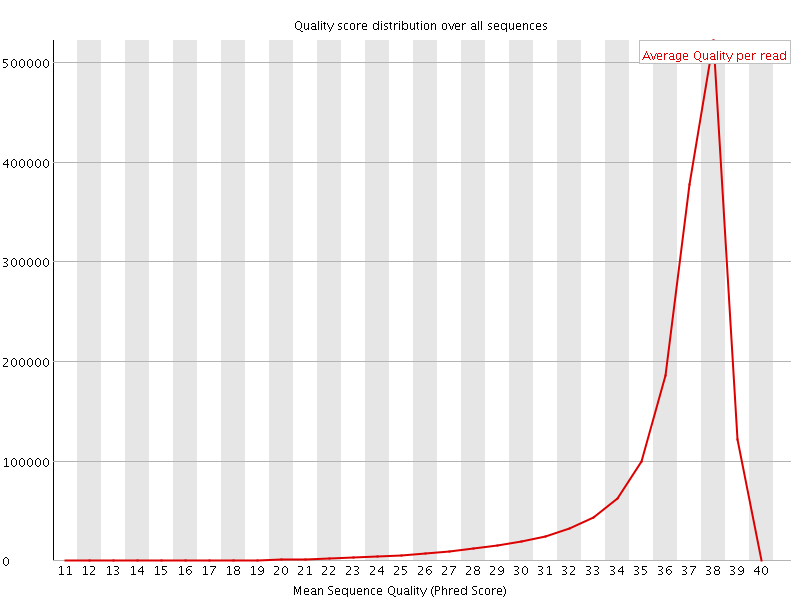
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
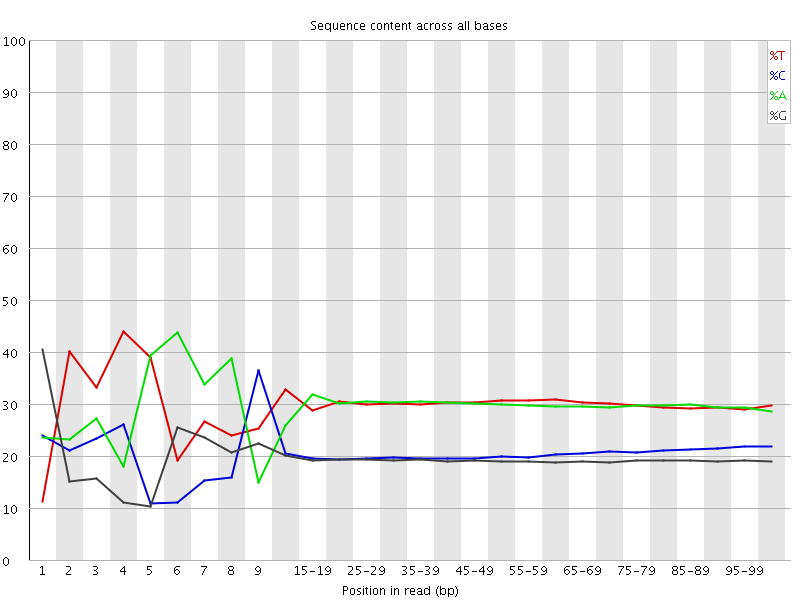
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
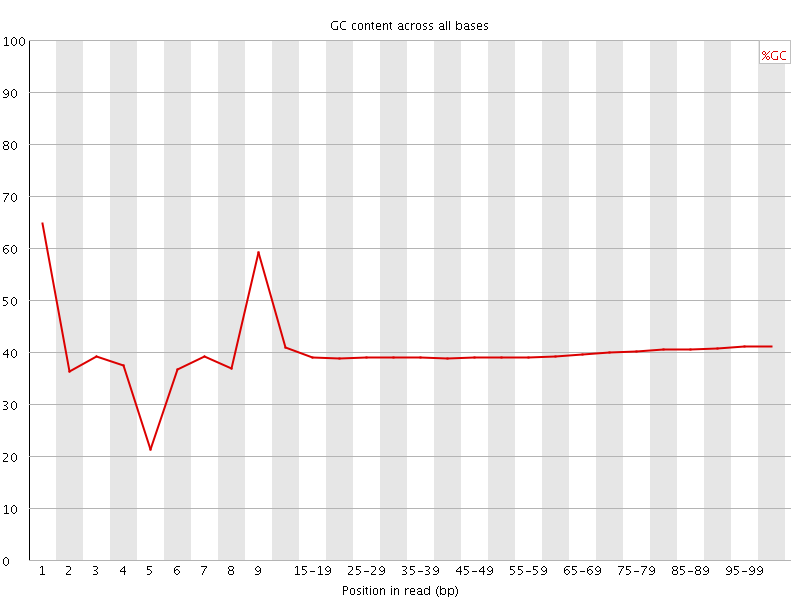
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
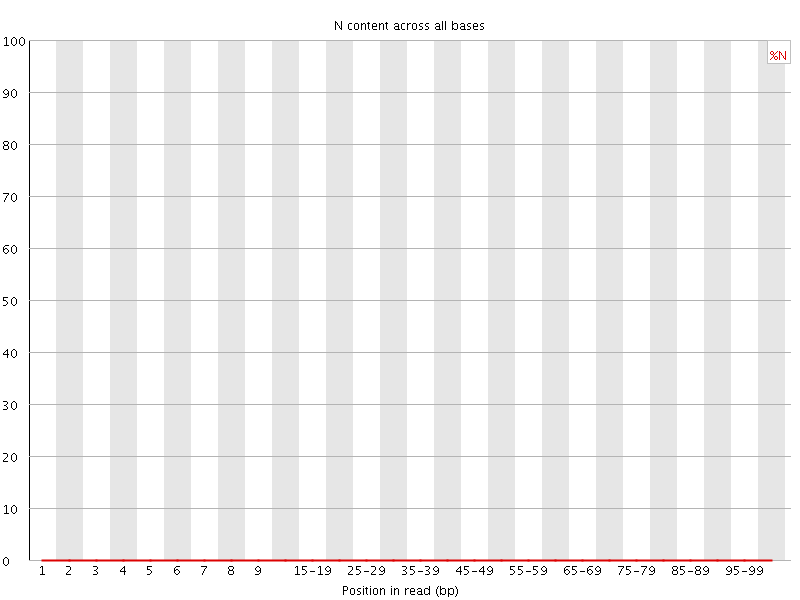
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
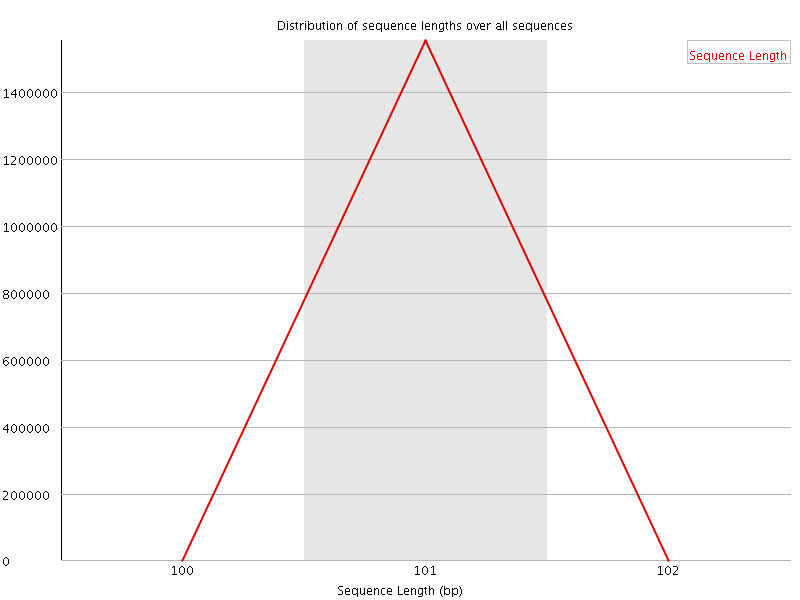
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
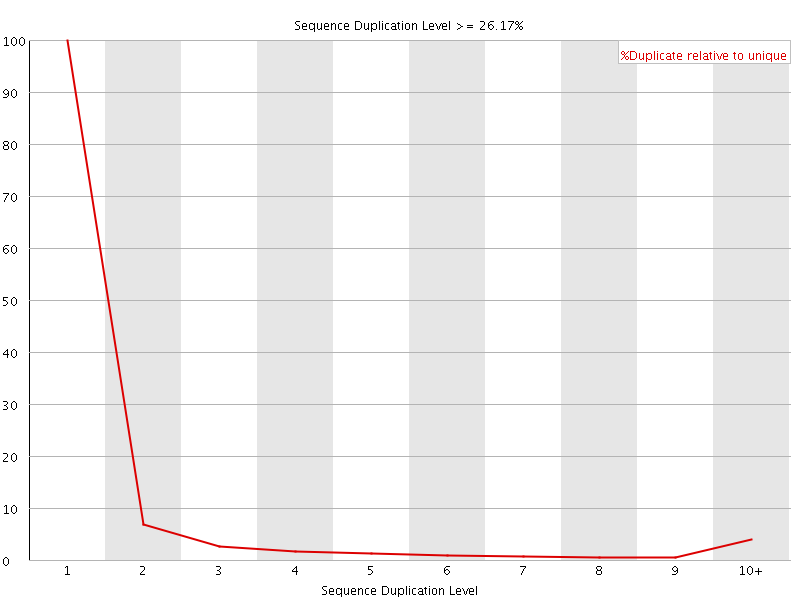
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
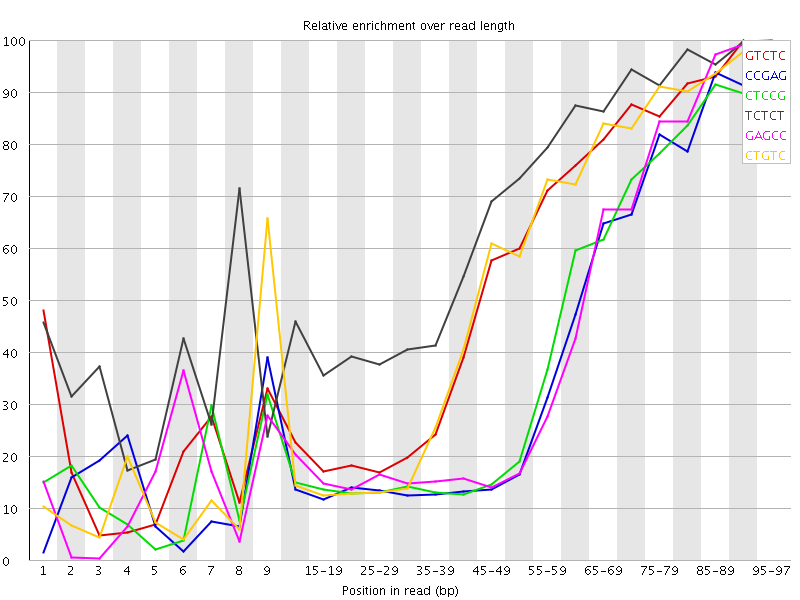
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 361390 | 3.2214935 | 5.924187 | 90-94 |
| CCGAG | 223375 | 3.1269069 | 7.941278 | 95-97 |
| CTCCG | 235205 | 3.0894644 | 7.589543 | 95-97 |
| TCTCT | 537185 | 3.0813515 | 4.615964 | 95-97 |
| GAGCC | 219640 | 3.0746229 | 7.4593725 | 95-97 |
| CTGTC | 334805 | 2.98451 | 5.6375947 | 95-97 |
| CGAGC | 207540 | 2.9052413 | 7.522722 | 95-97 |
| TCTCC | 330305 | 2.7918239 | 5.5777006 | 95-97 |
| GCCCA | 204170 | 2.7099676 | 7.1515684 | 90-94 |
| AGCCC | 202955 | 2.6938407 | 7.0022554 | 90-94 |
| AGAAG | 400475 | 2.6364267 | 5.2761126 | 5 |
| ACACA | 417705 | 2.4722562 | 7.073618 | 6 |
| CCCAC | 195360 | 2.4586658 | 6.638193 | 90-94 |
| ATCTC | 423170 | 2.4528315 | 5.6733766 | 95-97 |
| CCACG | 174930 | 2.3218622 | 6.675974 | 95-97 |
| TCCGA | 253065 | 2.2795477 | 5.4856787 | 95-97 |
| TACAC | 370350 | 2.1692054 | 9.164065 | 5 |
| GAGAC | 220050 | 2.112427 | 5.57779 | 95-97 |
| CGAGA | 211045 | 2.0259812 | 5.473837 | 95-97 |
| CTTTG | 334095 | 2.021136 | 8.057219 | 9 |
| GGCAG | 134205 | 1.9813322 | 6.561534 | 90-94 |
| AGAGA | 290720 | 1.9138821 | 5.1580143 | 8 |
| ACGAG | 196780 | 1.8890407 | 5.2535176 | 95-97 |
| GAGAG | 184805 | 1.8710365 | 6.4255323 | 7 |
| CAGGC | 133540 | 1.869355 | 6.2269597 | 90-94 |
| TGGCT | 167530 | 1.575005 | 5.0550075 | 6 |
| TATAC | 388290 | 1.5434374 | 5.7778373 | 5 |
| TTGAG | 233615 | 1.5061563 | 5.0170135 | 9 |
| GAGTC | 145600 | 1.3832039 | 6.3793364 | 9 |
| GTCTT | 226705 | 1.371471 | 6.1956577 | 1 |
| CCATG | 150050 | 1.3516138 | 5.847875 | 9 |
| GACTC | 149095 | 1.3430114 | 5.140365 | 7 |
| AAGAC | 214115 | 1.3365314 | 5.7502475 | 5 |
| TATGA | 294965 | 1.2365495 | 5.7338324 | 4 |
| GATTG | 191735 | 1.2361486 | 5.635934 | 7 |
| GTGTT | 192190 | 1.2262093 | 5.649402 | 1 |
| GACTT | 197375 | 1.2065719 | 5.912959 | 7 |
| GTTCT | 198760 | 1.2024155 | 5.1219783 | 1 |
| TCATA | 295275 | 1.1737062 | 5.7710185 | 2 |
| GTATG | 180965 | 1.1667125 | 5.1389213 | 3 |
| GTGTA | 171930 | 1.1084623 | 10.570179 | 1 |
| TGGAC | 115890 | 1.1009581 | 7.185389 | 5 |
| ACCAT | 186400 | 1.0917776 | 5.0122633 | 8 |
| ACACC | 122815 | 1.0599741 | 5.5361085 | 6 |
| GTCCA | 117110 | 1.0548983 | 7.264016 | 1 |
| GGACT | 104810 | 0.9956978 | 6.476063 | 6 |
| AGACT | 157155 | 0.97078884 | 5.4119725 | 6 |
| GAGTA | 148810 | 0.9694757 | 5.414814 | 1 |
| ACACT | 160260 | 0.938671 | 5.020783 | 6 |
| GTATA | 181565 | 0.761155 | 7.1597567 | 1 |