![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-115_AGGCAGAA-TATCCTCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2524992 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
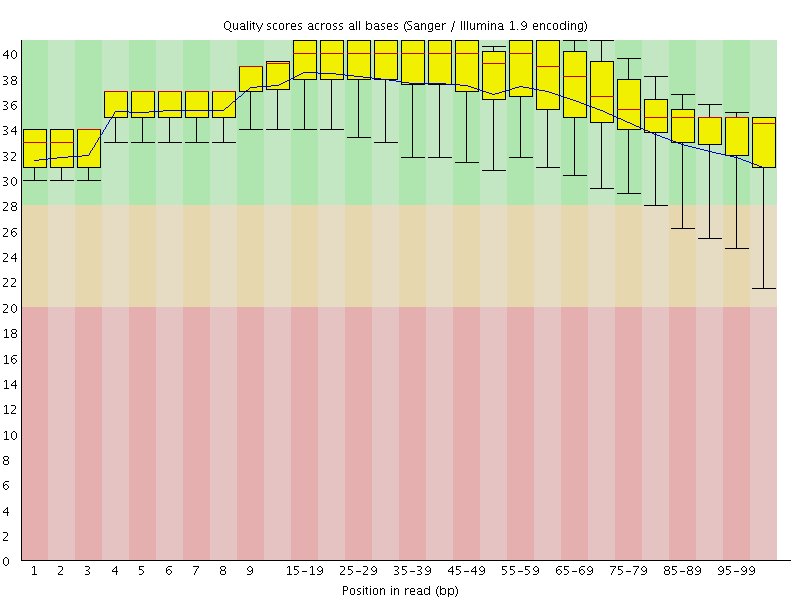
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
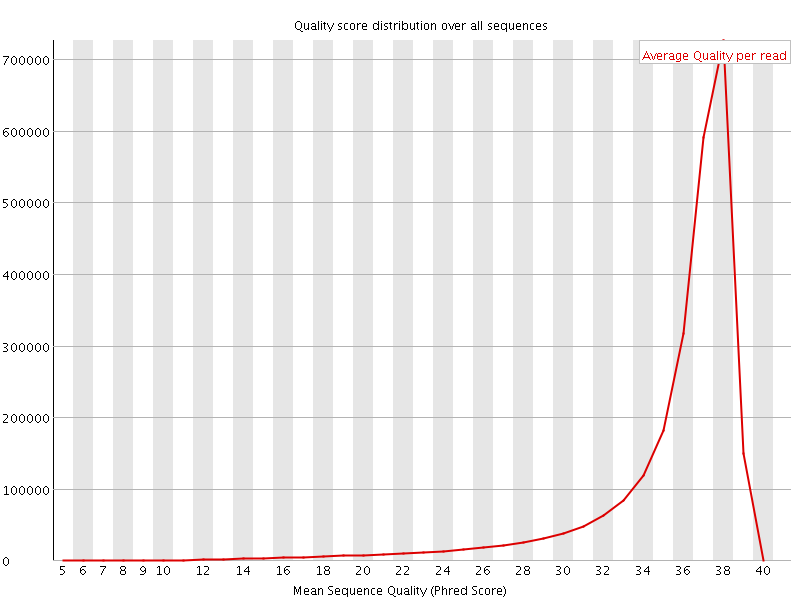
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
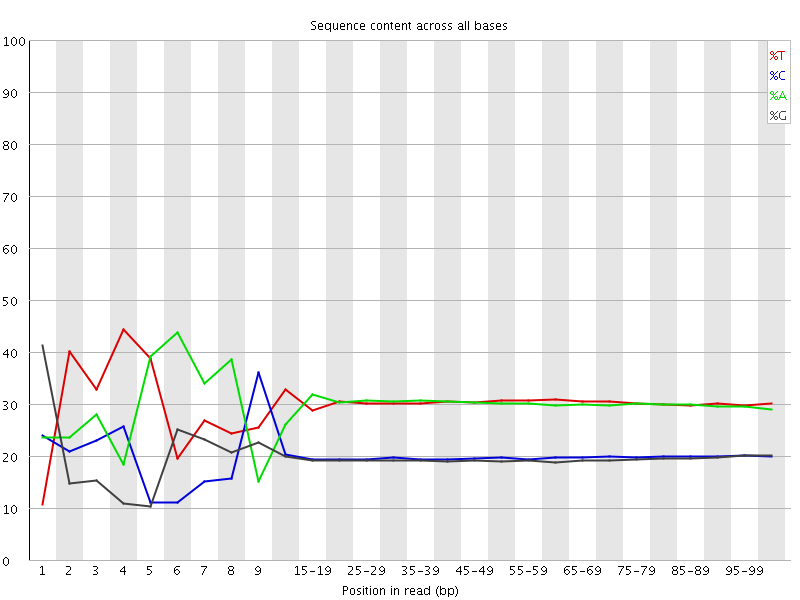
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
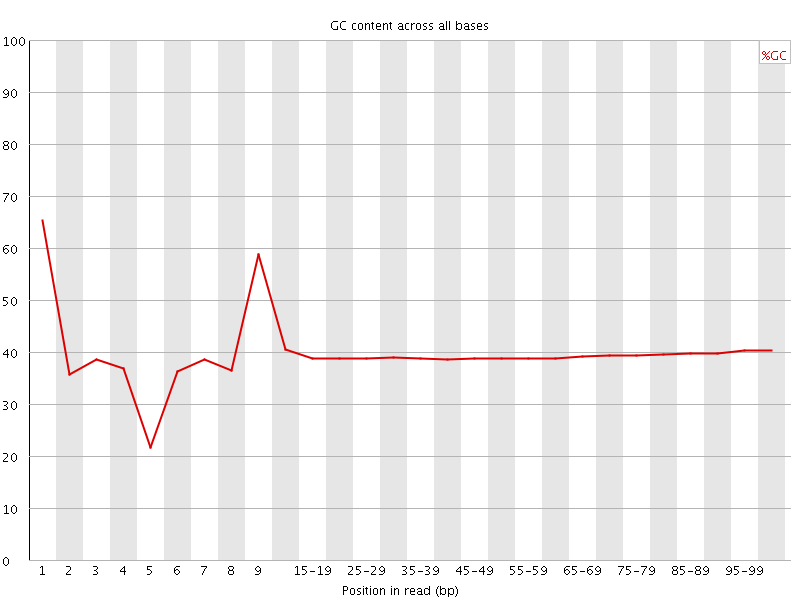
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
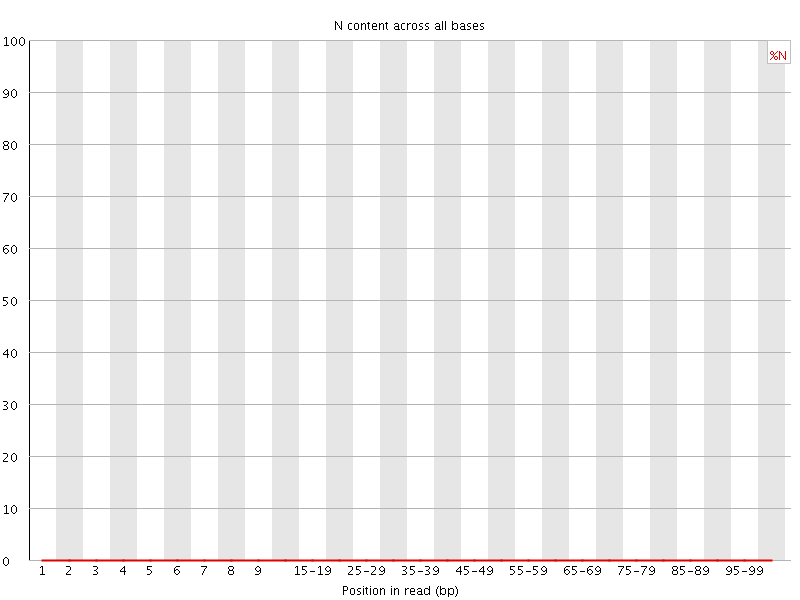
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
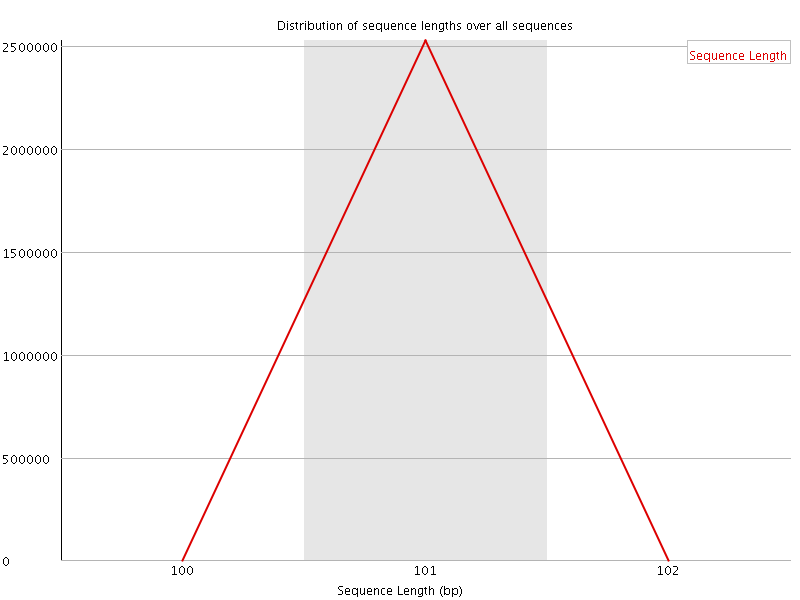
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
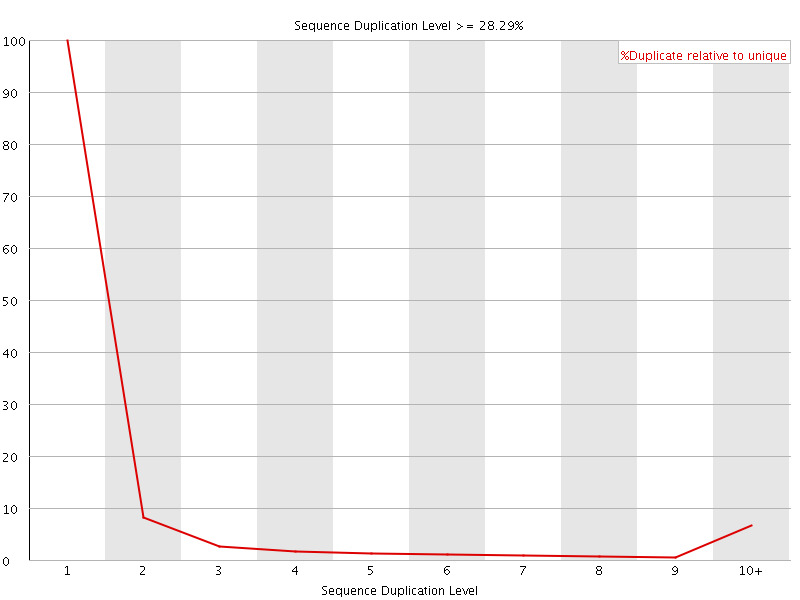
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
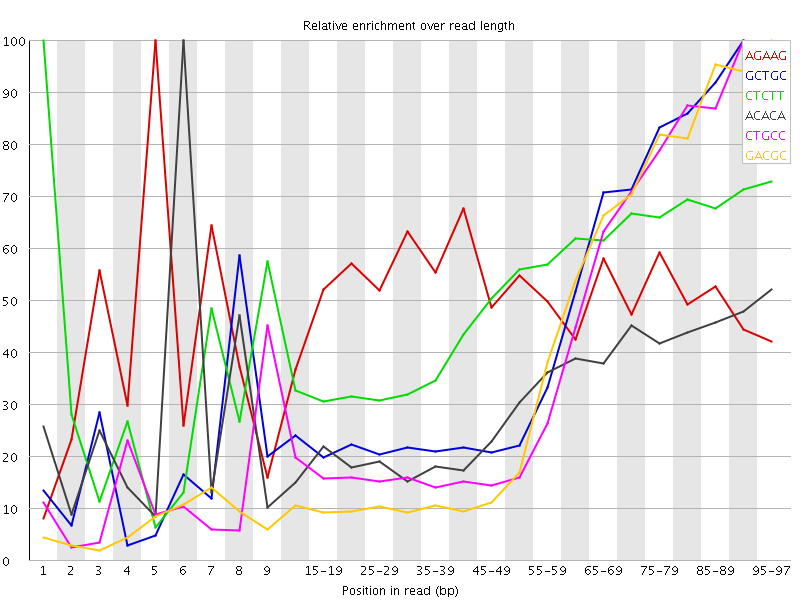
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 665905 | 2.6036909 | 5.102065 | 5 |
| GCTGC | 294445 | 2.5985312 | 5.763645 | 90-94 |
| CTCTT | 691340 | 2.529784 | 5.0465713 | 1 |
| ACACA | 594850 | 2.2493794 | 7.291021 | 6 |
| CTGCC | 251000 | 2.1783943 | 5.388594 | 90-94 |
| GACGC | 239305 | 2.135158 | 5.5032587 | 95-97 |
| CGCTG | 238170 | 2.101894 | 5.324913 | 95-97 |
| GCCGA | 233145 | 2.0801964 | 5.4497523 | 90-94 |
| CTTTG | 553505 | 2.0595589 | 8.277072 | 9 |
| CGACG | 230350 | 2.0552583 | 5.6359715 | 95-97 |
| CCGAC | 224695 | 1.9715631 | 5.2219205 | 95-97 |
| TACAC | 514745 | 1.9252758 | 8.995472 | 5 |
| TGCCG | 214385 | 1.891987 | 5.177265 | 90-94 |
| GAGAG | 299650 | 1.7997115 | 5.9009347 | 7 |
| GCTTT | 466445 | 1.7356138 | 5.26703 | 1 |
| TTGAG | 385550 | 1.4748533 | 5.029131 | 9 |
| GACTC | 251100 | 1.4426433 | 5.3772755 | 7 |
| GAGTC | 242430 | 1.4163139 | 6.8902035 | 9 |
| CCATG | 239400 | 1.3754233 | 5.6113453 | 9 |
| AAGAC | 353760 | 1.3602694 | 5.674035 | 5 |
| TATAC | 538535 | 1.3188932 | 5.873416 | 5 |
| GTCTT | 353070 | 1.3137523 | 6.5612245 | 1 |
| GTGTA | 335705 | 1.2841802 | 10.1003 | 1 |
| GACTT | 330485 | 1.2432513 | 5.9837546 | 7 |
| GTGTT | 324160 | 1.2265157 | 5.8550706 | 1 |
| GATTG | 319870 | 1.223606 | 5.65426 | 7 |
| TATGA | 489130 | 1.2180945 | 5.492856 | 4 |
| GTTCT | 325500 | 1.2111661 | 5.470997 | 1 |
| TCATA | 492900 | 1.2071313 | 5.6084948 | 2 |
| ACACC | 198985 | 1.1366483 | 5.343487 | 6 |
| TGGAC | 193815 | 1.1322975 | 7.3087163 | 5 |
| ACATG | 294835 | 1.1213489 | 5.1127496 | 8 |
| GTCCA | 194675 | 1.1184652 | 7.851035 | 1 |
| ACACT | 275820 | 1.031636 | 5.047783 | 6 |
| GGACT | 174275 | 1.0181419 | 6.7512465 | 6 |
| AGACT | 259110 | 0.9854756 | 5.2841244 | 6 |
| GTATG | 247715 | 0.9475899 | 5.043455 | 3 |
| GTATA | 298255 | 0.74275297 | 7.099702 | 1 |