![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-114_AGGCAGAA-CTCTCTAT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1570776 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
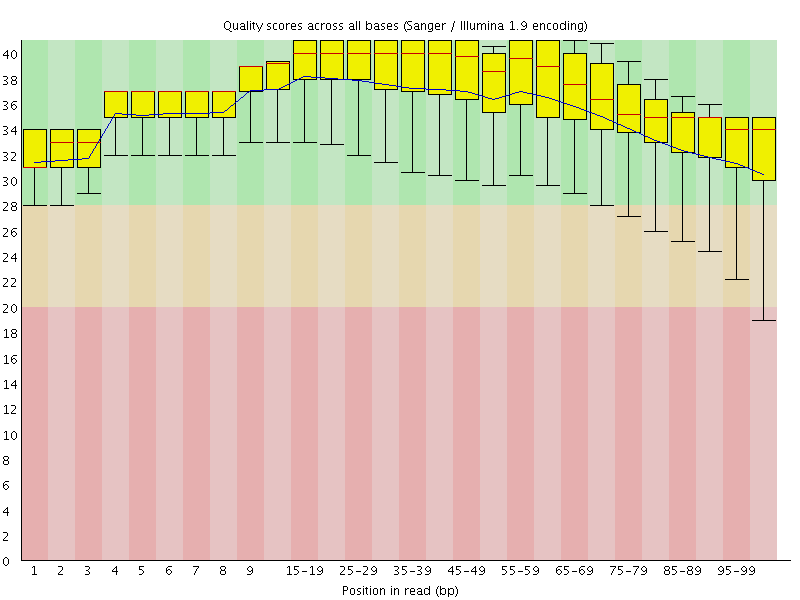
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
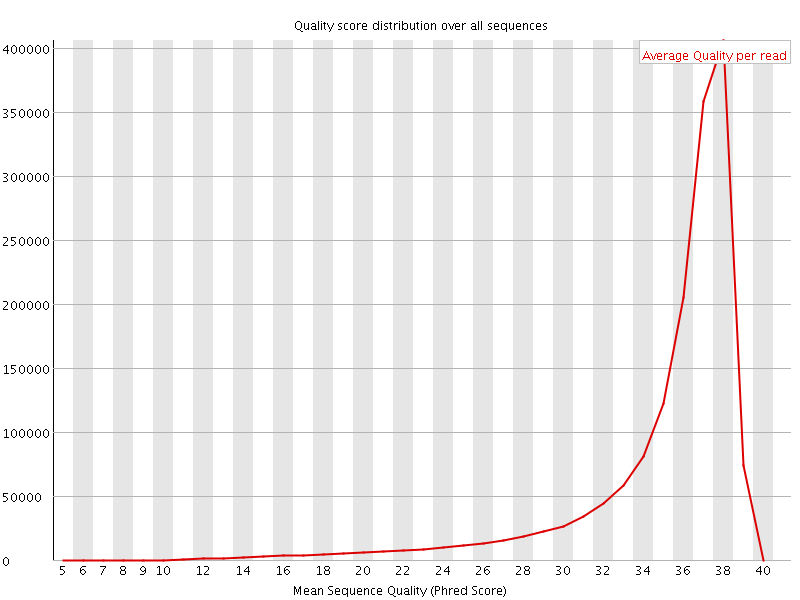
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
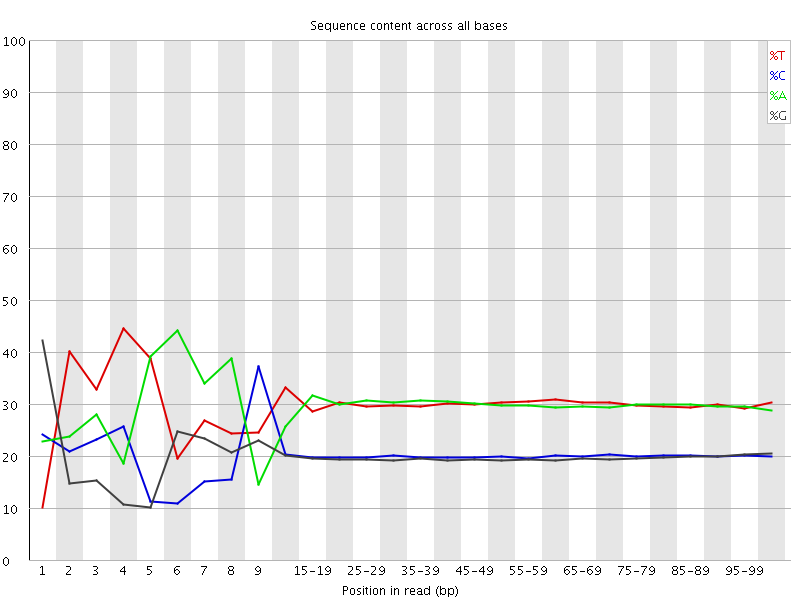
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
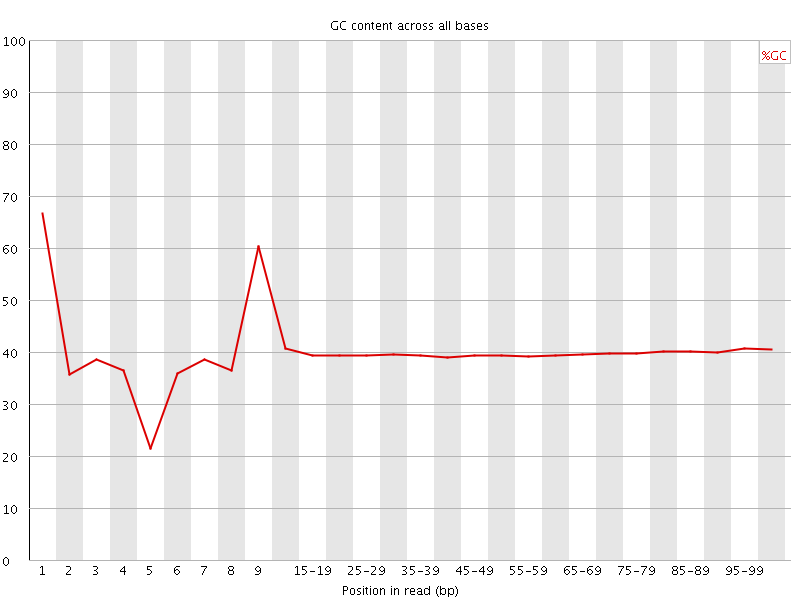
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
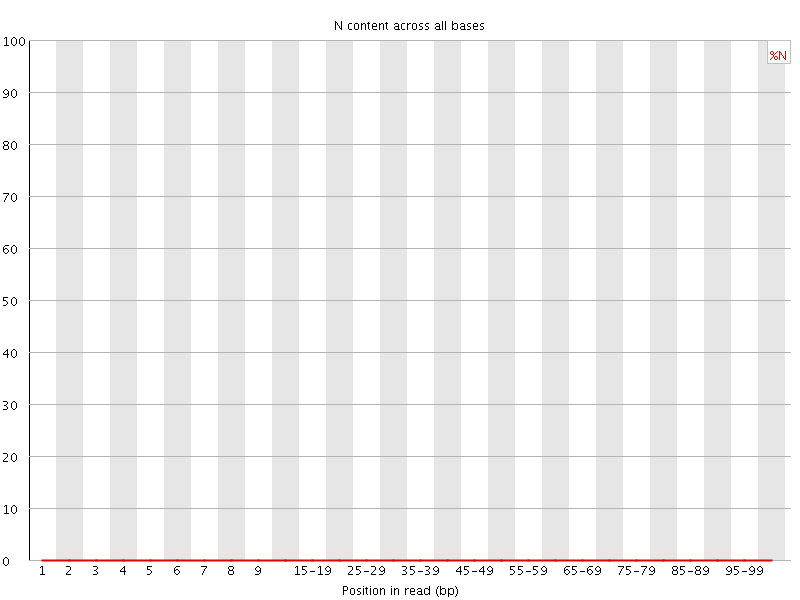
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
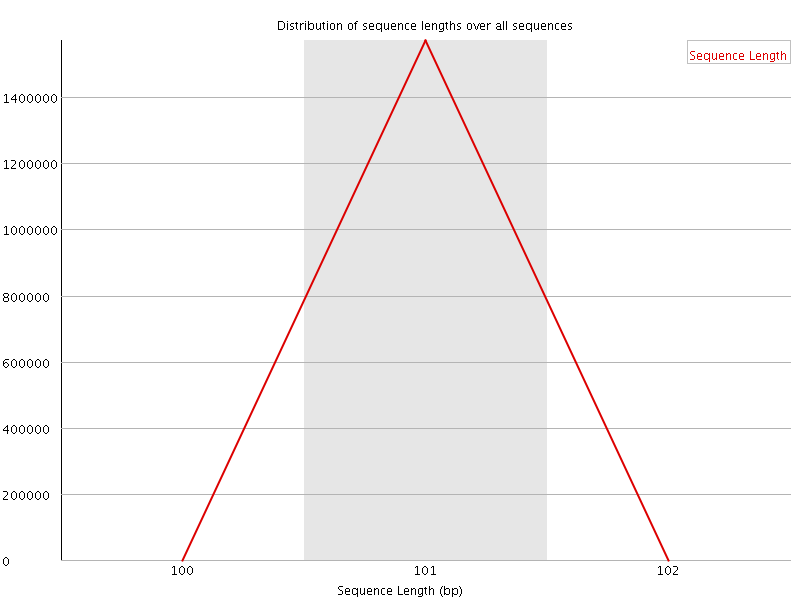
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
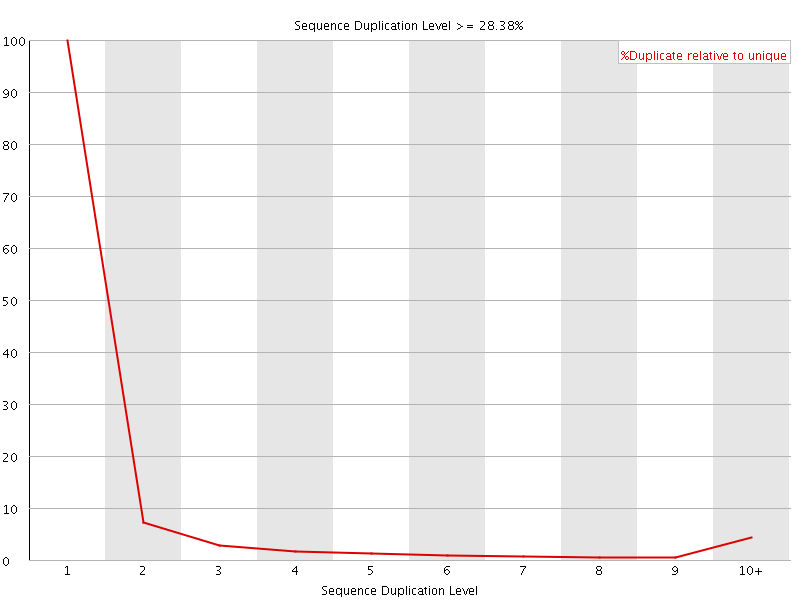
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 1857 | 0.11822182157099421 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
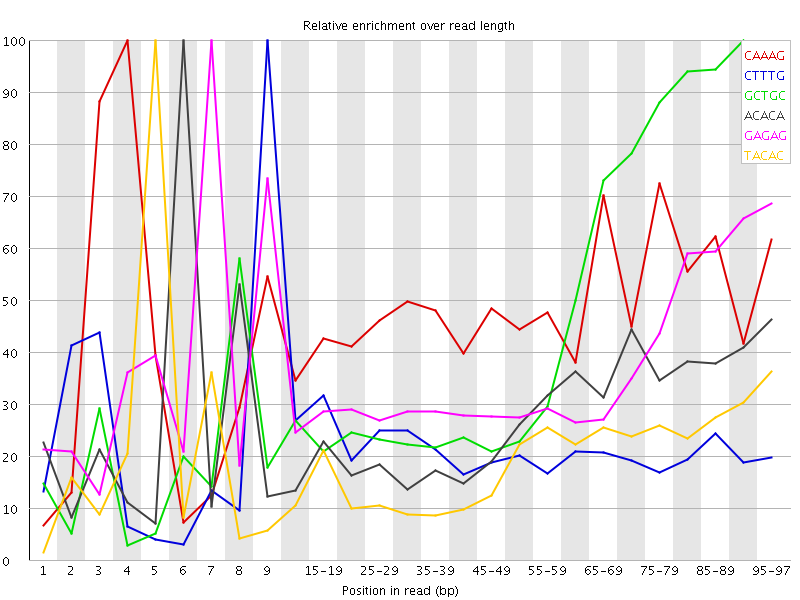
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CAAAG | 406300 | 2.5007725 | 5.177008 | 4 |
| CTTTG | 397305 | 2.3851721 | 10.937837 | 9 |
| GCTGC | 173360 | 2.3731558 | 5.0736556 | 90-94 |
| ACACA | 379415 | 2.2991896 | 8.310715 | 6 |
| GAGAG | 231230 | 2.1817372 | 5.9851036 | 7 |
| TACAC | 334550 | 2.0105307 | 10.185328 | 5 |
| GCTTT | 333785 | 2.0038376 | 5.96383 | 1 |
| CGACG | 139700 | 1.9283432 | 5.023659 | 95-97 |
| CACAT | 314160 | 1.8879938 | 6.0474653 | 7 |
| GCCAA | 203230 | 1.8587112 | 5.802718 | 1 |
| TTGGC | 194885 | 1.7805268 | 5.154093 | 5 |
| TGGCT | 194075 | 1.7731262 | 5.7199864 | 6 |
| GACTC | 188000 | 1.7051846 | 6.3428497 | 7 |
| GAGTC | 177880 | 1.6387314 | 9.864481 | 9 |
| TTGAG | 264265 | 1.624849 | 5.9634547 | 9 |
| ATACA | 398420 | 1.6113654 | 5.825163 | 6 |
| AAGAC | 246105 | 1.514774 | 6.4100566 | 5 |
| CATGA | 245090 | 1.4960372 | 5.677537 | 9 |
| CCATG | 164550 | 1.49249 | 5.79727 | 9 |
| ATGGA | 239440 | 1.4845012 | 5.4974647 | 4 |
| GTGTA | 237445 | 1.4599445 | 11.545058 | 1 |
| GTCTT | 241375 | 1.4490653 | 7.24221 | 1 |
| TGTGT | 236560 | 1.4424609 | 5.550542 | 8 |
| GACTT | 237680 | 1.4387947 | 8.263998 | 7 |
| TCCAT | 240415 | 1.4328499 | 5.5071497 | 2 |
| TATGA | 348775 | 1.4208716 | 7.0092754 | 4 |
| TCATA | 349100 | 1.4002069 | 7.075847 | 2 |
| ACTCA | 232245 | 1.395713 | 5.866808 | 8 |
| TGAGT | 225995 | 1.3895435 | 7.311427 | 8 |
| GTGTT | 221985 | 1.3535877 | 6.415374 | 1 |
| AACAC | 221745 | 1.3437365 | 5.5320015 | 5 |
| ATGAG | 214575 | 1.3303412 | 6.0825458 | 7 |
| TATAC | 324965 | 1.3034039 | 6.4903913 | 5 |
| GATTG | 209285 | 1.2868011 | 6.56593 | 7 |
| ACACC | 139540 | 1.2564803 | 5.5763063 | 6 |
| GTTCT | 206925 | 1.242249 | 5.6784515 | 1 |
| TGGAC | 130445 | 1.2017332 | 8.781033 | 5 |
| GTCCA | 129925 | 1.1784366 | 9.058776 | 1 |
| ACATG | 192295 | 1.1737748 | 6.3467293 | 8 |
| ACACT | 191760 | 1.1524118 | 5.5111203 | 6 |
| ATGTG | 186615 | 1.1474133 | 5.549135 | 7 |
| GGACT | 123030 | 1.1334221 | 8.653843 | 6 |
| AGACT | 176000 | 1.0743096 | 6.260742 | 6 |
| ACTTT | 267085 | 1.0623835 | 5.04057 | 8 |
| GTATG | 170295 | 1.0470688 | 6.1018953 | 3 |
| TGTAT | 258165 | 1.0430287 | 5.4009585 | 2 |
| GAGTA | 164350 | 1.0189518 | 5.2568426 | 1 |
| CATAC | 167480 | 1.0064975 | 5.859109 | 3 |
| ATACT | 200930 | 0.8059111 | 5.302332 | 6 |
| GTATA | 196015 | 0.7985438 | 7.6731997 | 1 |