![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-114_AGGCAGAA-CTCTCTAT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1570776 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
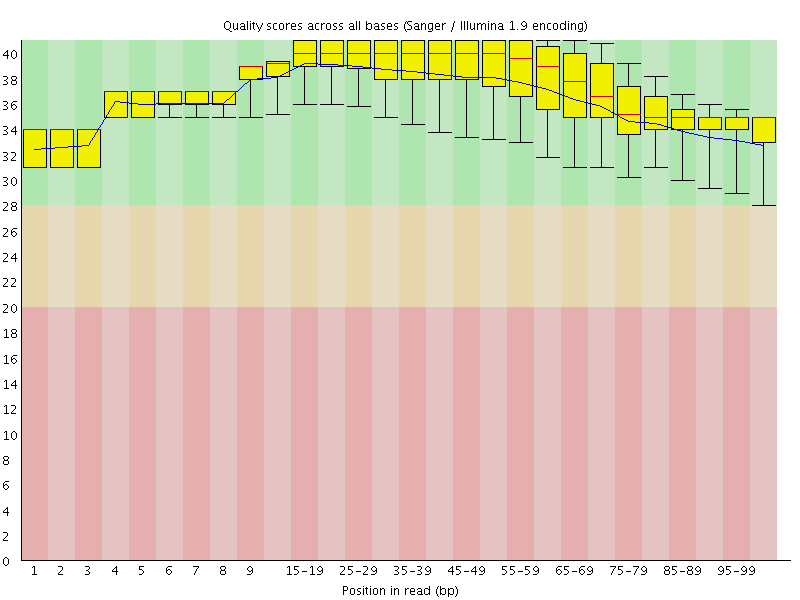
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
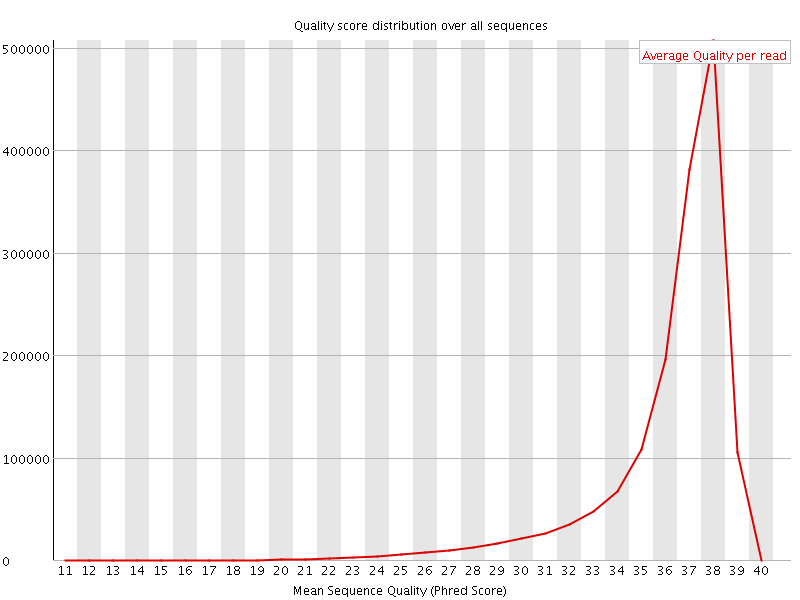
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
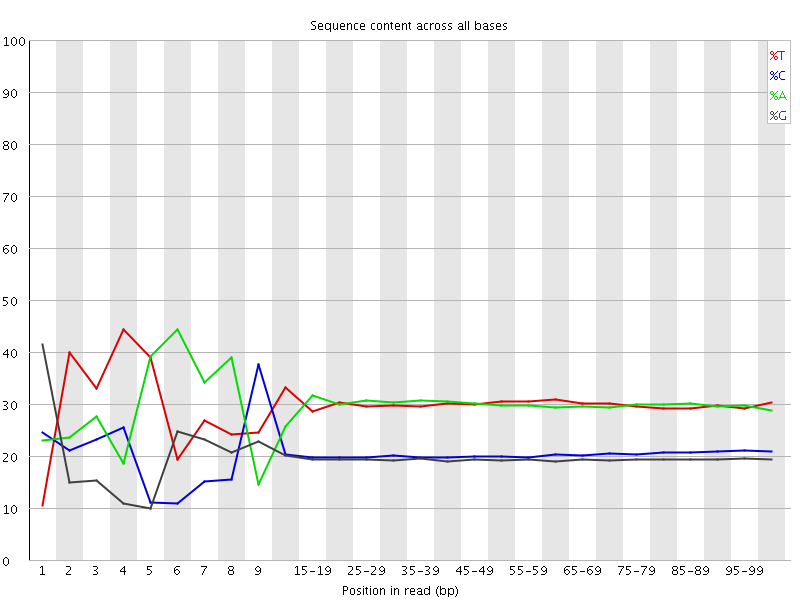
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
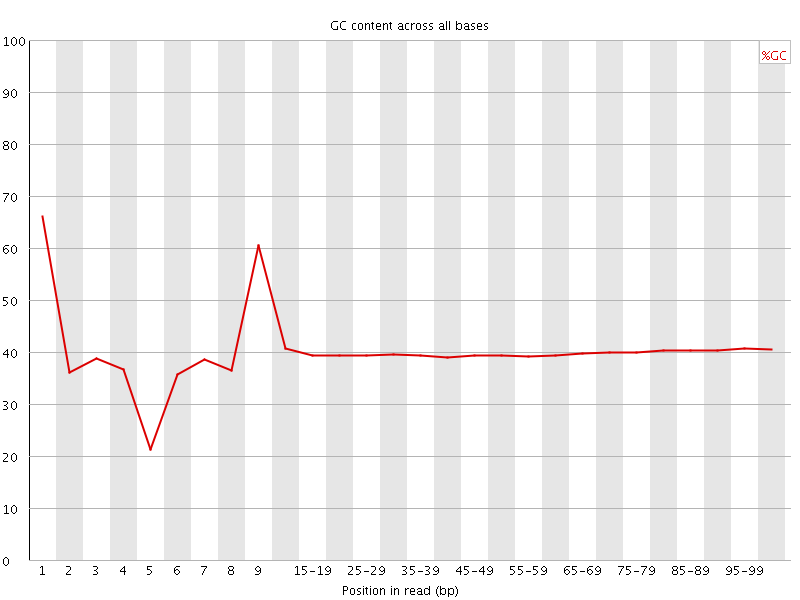
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
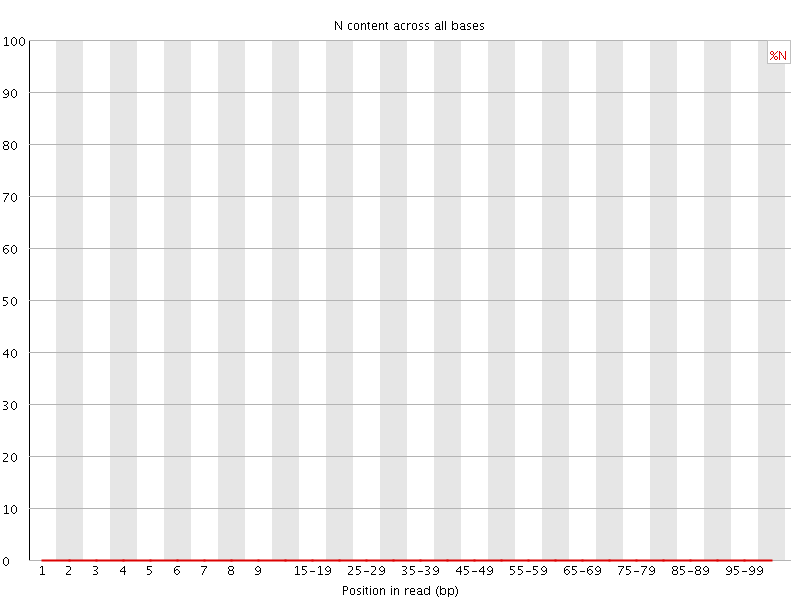
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
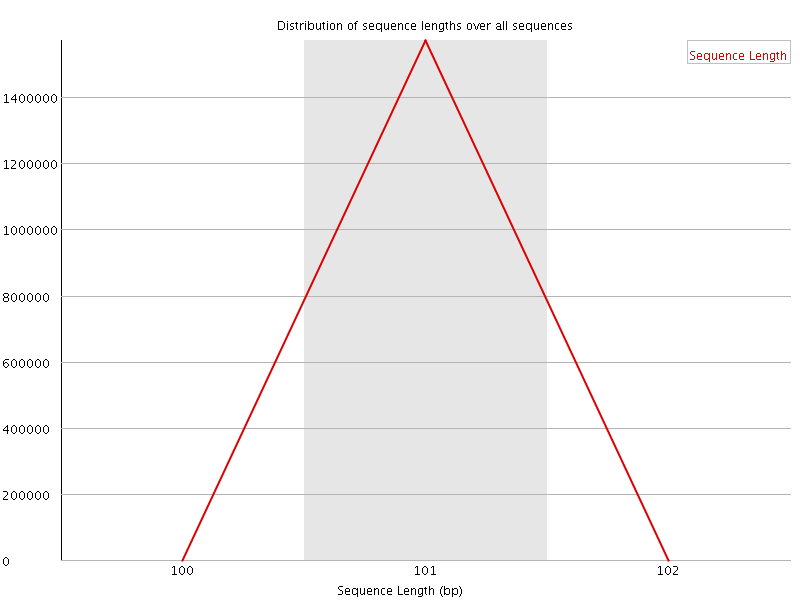
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
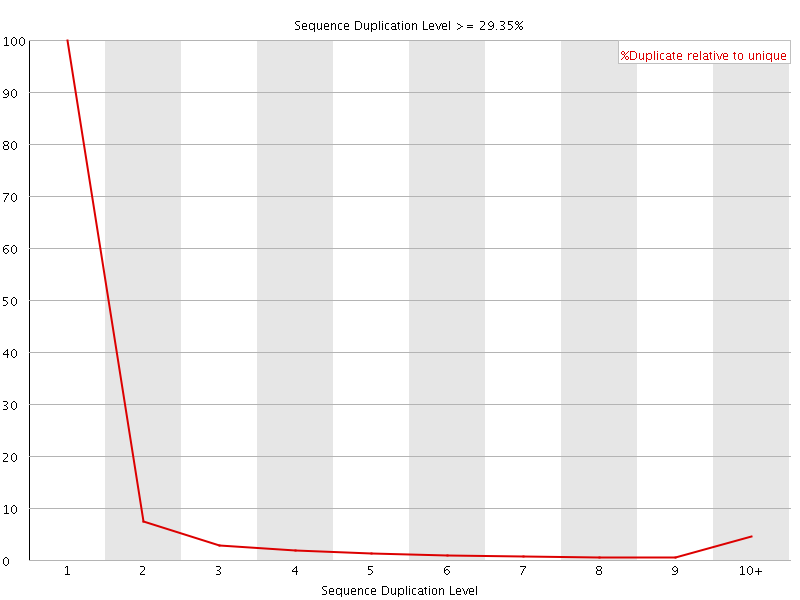
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 1935 | 0.12318752005378235 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
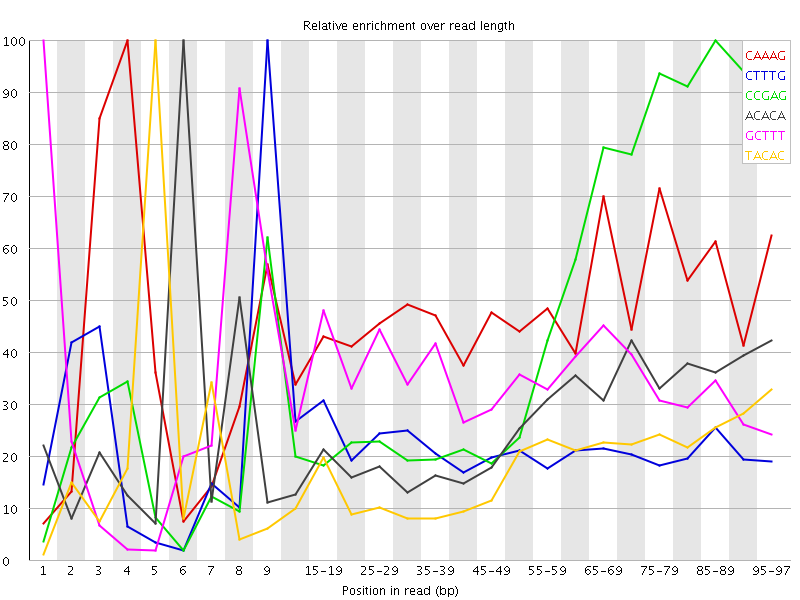
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CAAAG | 409465 | 2.507775 | 5.2380524 | 4 |
| CTTTG | 397925 | 2.4030206 | 10.859569 | 9 |
| CCGAG | 175015 | 2.4011688 | 5.062555 | 85-89 |
| ACACA | 380245 | 2.2364714 | 8.38391 | 6 |
| GCTTT | 330150 | 1.9937353 | 5.7188005 | 1 |
| TACAC | 338930 | 1.9841354 | 10.779924 | 5 |
| CACAT | 316580 | 1.8532959 | 6.161984 | 7 |
| GCCAA | 203270 | 1.8259836 | 5.344593 | 1 |
| TGGCT | 195720 | 1.8136514 | 5.8855815 | 6 |
| TTGGC | 195035 | 1.8073038 | 5.2655735 | 5 |
| GAGAG | 178285 | 1.7365305 | 5.5630426 | 7 |
| GACTC | 189805 | 1.6970427 | 6.185981 | 7 |
| TTGAG | 265220 | 1.6756103 | 6.101766 | 9 |
| GAGTC | 179560 | 1.6717322 | 9.4702635 | 9 |
| CACCA | 190770 | 1.6457418 | 5.102828 | 7 |
| GGCTT | 176295 | 1.6336483 | 5.140331 | 1 |
| ATACA | 400705 | 1.5993159 | 6.0337186 | 6 |
| ATGGA | 239585 | 1.5207748 | 5.354933 | 4 |
| GTCTT | 250445 | 1.5124068 | 7.062851 | 1 |
| AAGAC | 246595 | 1.5102751 | 6.3634615 | 5 |
| CATGA | 246365 | 1.5018007 | 5.804626 | 9 |
| CCATG | 166680 | 1.4902824 | 5.925883 | 9 |
| TGTGT | 236450 | 1.4868515 | 5.746972 | 8 |
| GACTT | 239100 | 1.4506891 | 8.12784 | 7 |
| TGAGT | 227860 | 1.4395769 | 7.2228737 | 8 |
| TATGA | 347440 | 1.4372183 | 6.909261 | 4 |
| TCCAT | 241815 | 1.4089838 | 5.040389 | 2 |
| TCATA | 350045 | 1.3905762 | 6.9865847 | 2 |
| GTGTT | 221085 | 1.3902328 | 6.320829 | 1 |
| ATGAG | 215240 | 1.366244 | 6.1489396 | 7 |
| ACTCA | 231275 | 1.3539106 | 5.6908236 | 8 |
| GTGTA | 212225 | 1.3407978 | 11.708571 | 1 |
| GGAGT | 136705 | 1.325298 | 5.1092615 | 8 |
| GATTG | 207560 | 1.3113252 | 6.757277 | 7 |
| AACAC | 221415 | 1.3022875 | 5.4580965 | 5 |
| TATAC | 327340 | 1.3003792 | 6.446531 | 5 |
| GTTCT | 206840 | 1.2490815 | 5.3674154 | 1 |
| GTATG | 194420 | 1.2283092 | 6.322312 | 3 |
| ACACC | 141000 | 1.216384 | 5.479266 | 6 |
| TGGAC | 129930 | 1.209669 | 8.590043 | 5 |
| GTCCA | 133505 | 1.1936656 | 8.627495 | 1 |
| ATGTG | 187650 | 1.1855376 | 5.666801 | 7 |
| ACATG | 193435 | 1.1791482 | 6.4903054 | 8 |
| GGACT | 122760 | 1.1429151 | 8.743518 | 6 |
| ACACT | 194655 | 1.1395328 | 5.892344 | 6 |
| AGACT | 176995 | 1.0789325 | 6.088355 | 6 |
| TGTAT | 259815 | 1.0697165 | 5.4861493 | 2 |
| ACTTT | 266505 | 1.0537504 | 5.080164 | 8 |
| GAGTA | 165250 | 1.0489306 | 5.5986648 | 1 |
| CATAC | 169275 | 0.9909554 | 6.0598044 | 3 |
| GTATA | 196995 | 0.81488836 | 7.618059 | 1 |
| ATACT | 201100 | 0.7988827 | 5.3101554 | 6 |