![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-104_TAAGGCGA-CTAAGCCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 734874 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
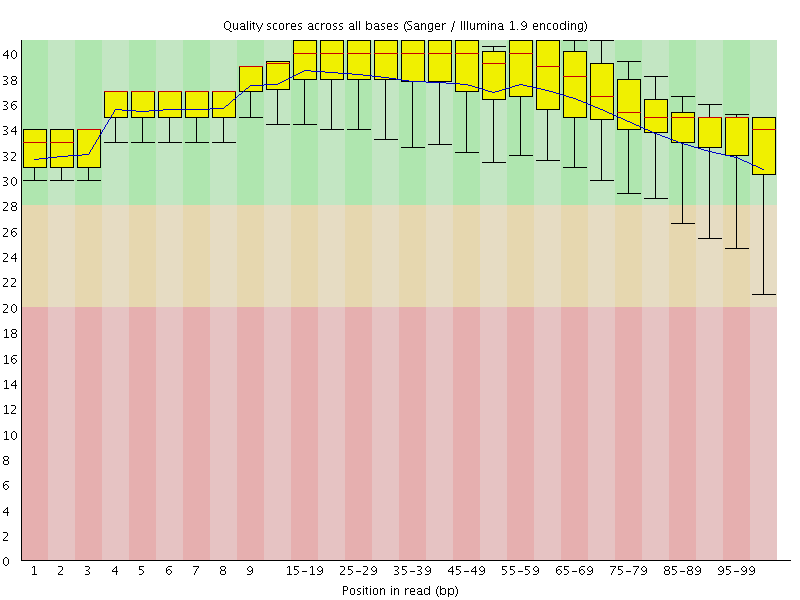
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
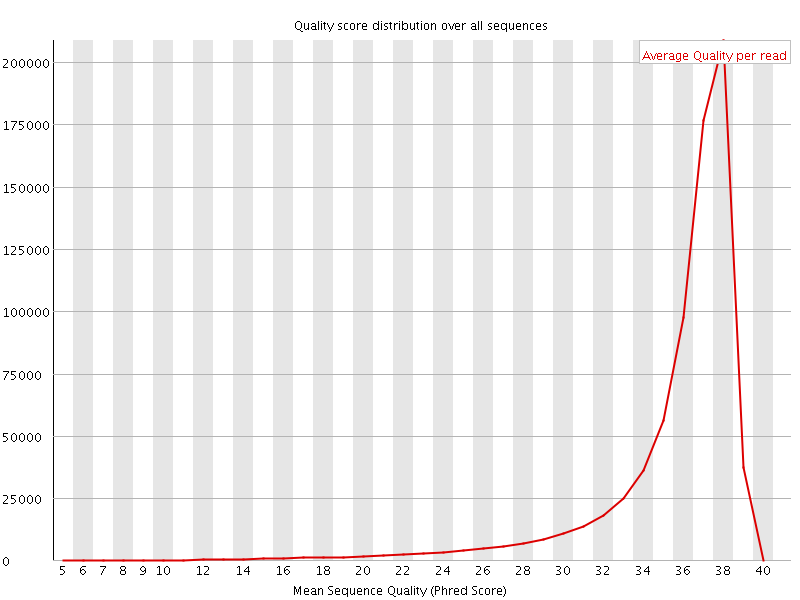
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
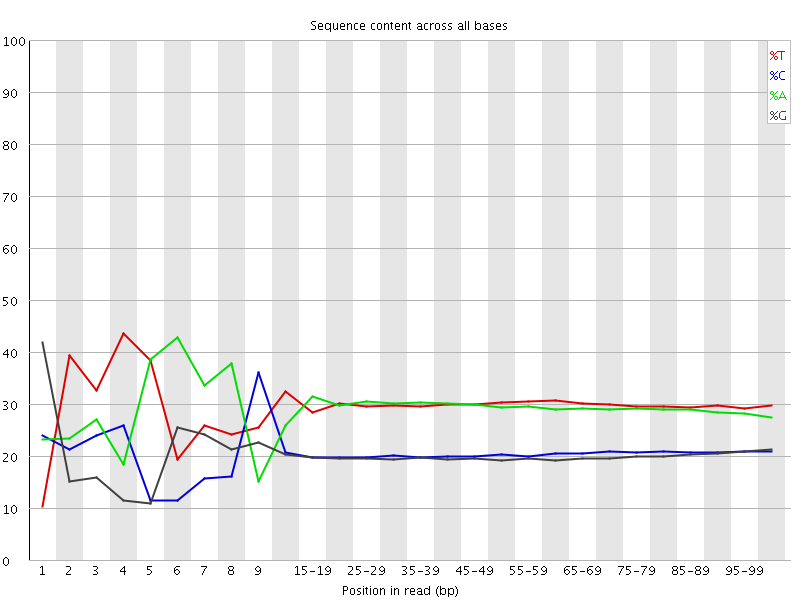
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
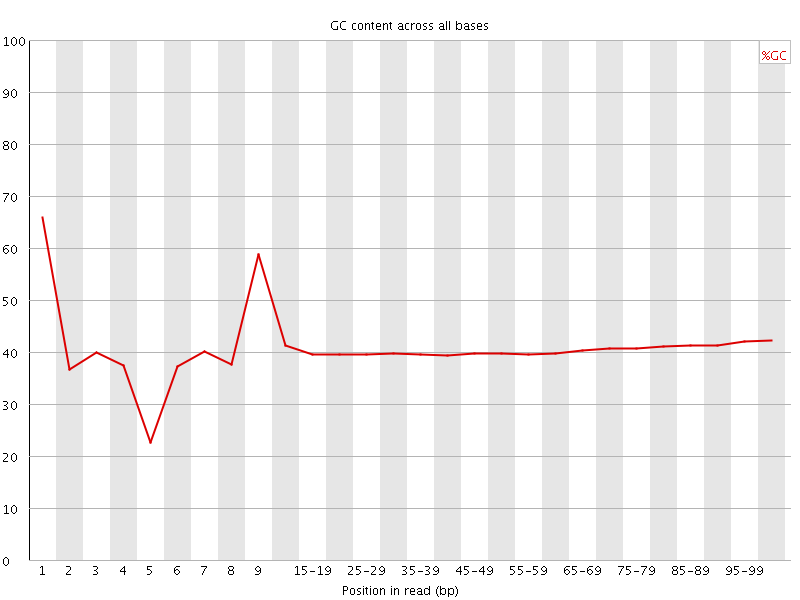
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
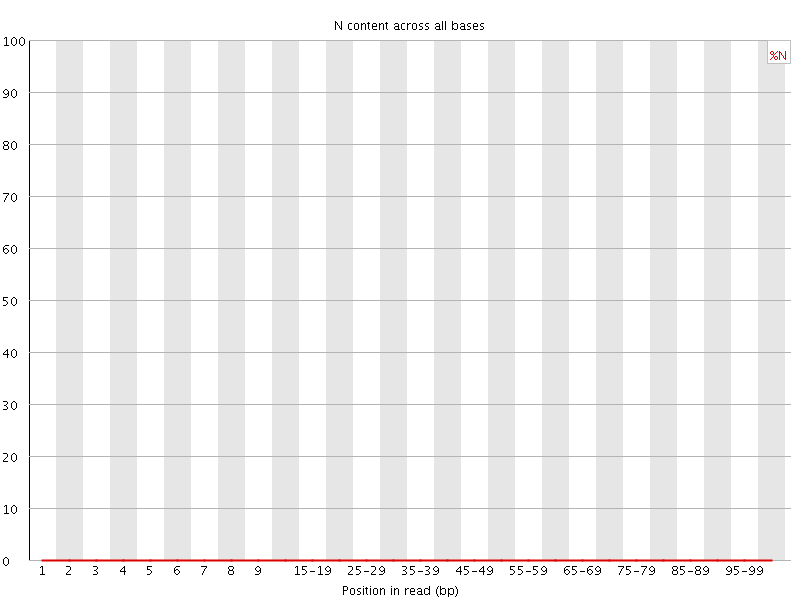
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
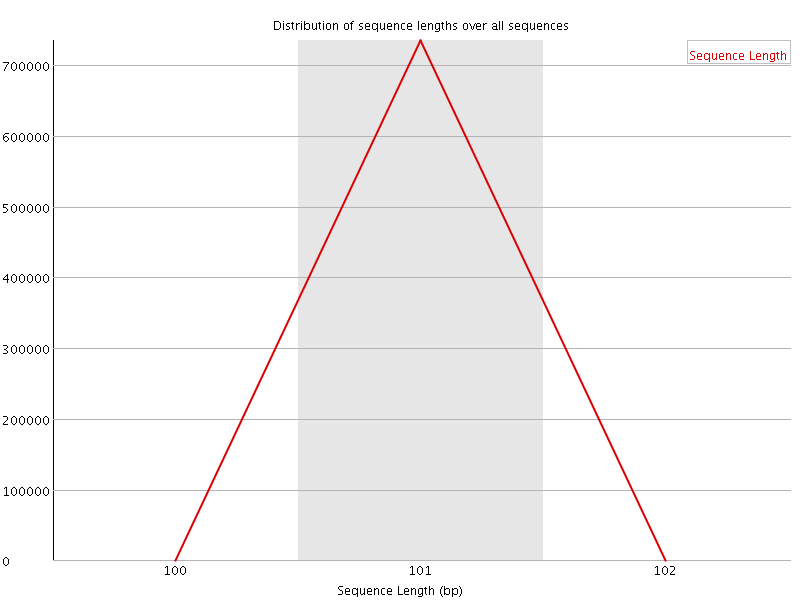
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
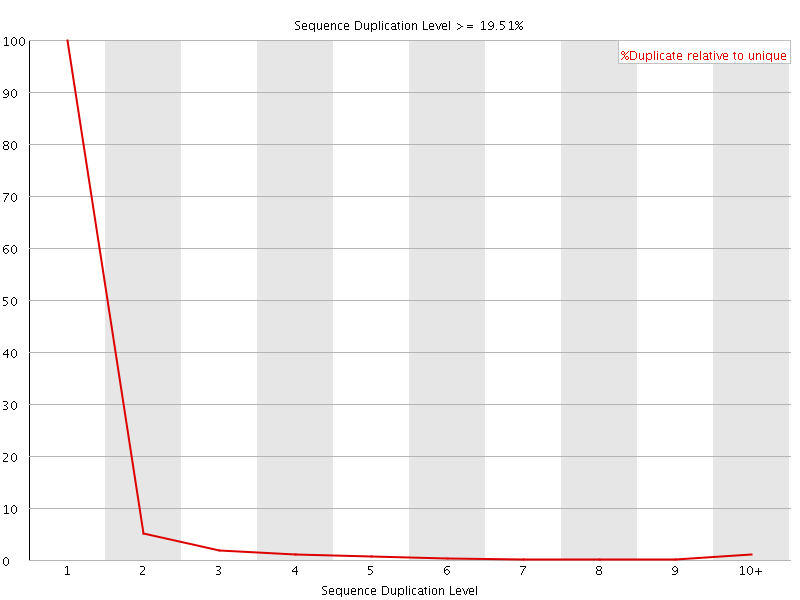
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
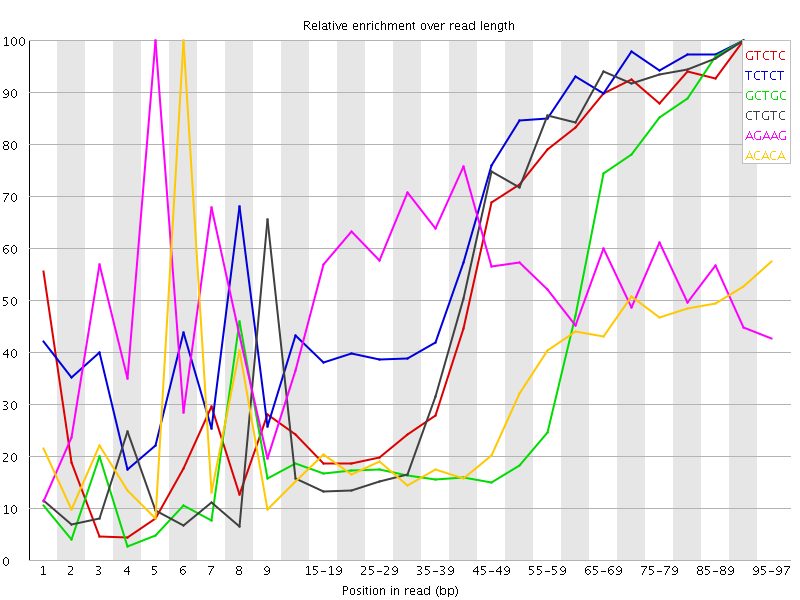
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 172770 | 3.1856337 | 5.4636655 | 90-94 |
| TCTCT | 249725 | 3.0734391 | 4.460988 | 90-94 |
| GCTGC | 108870 | 3.0074654 | 7.014673 | 90-94 |
| CTGTC | 158695 | 2.9261107 | 5.011192 | 90-94 |
| AGAAG | 205150 | 2.8171637 | 5.1489515 | 5 |
| ACACA | 208335 | 2.7257824 | 8.3875475 | 6 |
| GACGC | 94595 | 2.6669571 | 6.800858 | 85-89 |
| CGCTG | 93890 | 2.5936522 | 6.594007 | 90-94 |
| GAAGG | 127210 | 2.5643141 | 5.6608276 | 95-97 |
| GCCGA | 90525 | 2.55221 | 6.907591 | 90-94 |
| CTGCC | 94230 | 2.540831 | 6.5802536 | 90-94 |
| CGACG | 89375 | 2.5197875 | 6.9418936 | 95-97 |
| TACAC | 187705 | 2.4062974 | 10.210461 | 5 |
| CAAAG | 177995 | 2.3858466 | 5.1559253 | 4 |
| CCGAC | 86195 | 2.3720517 | 6.5716662 | 90-94 |
| TGCCG | 84060 | 2.3221045 | 6.5832887 | 90-94 |
| CACAT | 179590 | 2.302267 | 5.7442703 | 7 |
| GGCTT | 121060 | 2.2868302 | 5.182571 | 95-97 |
| CTTTG | 177240 | 2.234756 | 8.939394 | 9 |
| GACGA | 102410 | 2.0150533 | 5.2710466 | 95-97 |
| ATACA | 216125 | 1.9336367 | 5.0167956 | 4 |
| CGAAG | 96480 | 1.8983727 | 5.118355 | 95-97 |
| GCTTT | 149045 | 1.8792554 | 5.608749 | 1 |
| GAGAG | 86625 | 1.7461971 | 5.7186728 | 7 |
| GCCAA | 86980 | 1.6705436 | 5.4782777 | 1 |
| TGGCT | 86435 | 1.6327623 | 5.4778366 | 6 |
| TATAC | 184475 | 1.617156 | 5.4300523 | 5 |
| TTGAG | 118715 | 1.5650754 | 5.7731085 | 9 |
| GACTC | 80650 | 1.5177048 | 5.1834707 | 7 |
| GTGTA | 112415 | 1.4820195 | 10.47543 | 1 |
| GAGTC | 76670 | 1.4781353 | 8.031086 | 9 |
| AAGAC | 108520 | 1.4546032 | 5.962149 | 5 |
| CATGA | 108085 | 1.4195304 | 5.6238317 | 9 |
| CCATG | 73675 | 1.3864465 | 5.776689 | 9 |
| GTCTT | 109115 | 1.3757921 | 6.8381476 | 1 |
| TGAGT | 102010 | 1.3448455 | 5.5110226 | 8 |
| TATGA | 148890 | 1.3371671 | 6.5083294 | 4 |
| GACTT | 103320 | 1.3295609 | 6.234171 | 7 |
| TCATA | 150595 | 1.320155 | 6.905705 | 2 |
| AACAC | 100465 | 1.314449 | 5.1850114 | 5 |
| GTGTT | 101170 | 1.3068507 | 6.247376 | 1 |
| ATGAG | 94755 | 1.2749326 | 5.3961 | 7 |
| GATTG | 94220 | 1.2421463 | 6.4507494 | 7 |
| GTTCT | 96695 | 1.2191929 | 5.394675 | 1 |
| ACACC | 62110 | 1.1643784 | 5.0818167 | 6 |
| ACATG | 88495 | 1.1622459 | 5.929584 | 8 |
| ACACT | 88350 | 1.132609 | 5.6073103 | 6 |
| TGGAC | 58155 | 1.1211811 | 7.1259446 | 5 |
| GTCCA | 59280 | 1.1155553 | 8.0332985 | 1 |
| GGACT | 54400 | 1.0487878 | 6.544272 | 6 |
| AGACT | 78775 | 1.0345886 | 5.1650524 | 6 |
| GAGTA | 75700 | 1.0185468 | 5.2085433 | 1 |
| GTATG | 76565 | 1.0093921 | 6.043064 | 3 |
| CATAC | 75410 | 0.9667238 | 5.714586 | 3 |
| GTATA | 85380 | 0.7667898 | 6.7745247 | 1 |